Research Article - Journal of Food Microbiology (2022) Volume 6, Issue 4
Occurrence and antibiotics resistance of Staphylococcus species from â??awaraâ?? sold in wukari old market, Taraba State.
Hammuel C*, Aso RE, Briska J, Daji M, Anguwa ELDepartment of Microbiology, Federal University Wukari, Nigeria
- *Corresponding Author:
- Hammuel C
Department of Microbiology
Federal University Wukari, Nigeria
Mobile: 2347034727440
E-mail: chris2012bansi@gmail.com
Received: 03-Jun-2022, Manuscript No. AAFMY-22-65797; Editor assigned: 06-Jun-2022, PreQC No. AAFMY-22-65797(PQ); Reviewed: 21-Jun-2022, QC No. AAFMY-22-65797; Revised: 27-Jun-2022, Manuscript No. AAFMY-22-65797(R); Published: 04-Jul-2022, DOI:10.35841/aafmy-6.4.117
Citation: Hammuel C, Aso RE, Briska, J, et al. Occurrence and antibiotics resistance of Staphylococcus species from ‘awara’ sold in wukari old market, Taraba State. J Food Microbiol 2022;6(4):117
Abstract
Food poisoning cause by Staphylococcus species is global problem which has caused so many illnesses among the populations of the world and has also resulted to so many deaths. In Nigeria, there are so many health cases which are attributed to consumption of foods that are contaminated by Staphylococcus species. This research was carried out to determine the occurrence of the Staphylococcus species in soya bean cake sold in a market. A total of number twenty (20) samples were collected for this research. Five (5) samples from each of the four locations were collected from Awara vendors/hawkers within the market. Total bacterial count in the food samples ranged from 1.00 x 105 to 1.80 x 106 CFU/ml and the staphylococcal count on mannitol salt agar ranged from 1.2 x 105 to 8.0 x 105 CFU/ml. Two species of Staphylococcus which include Staphylococcus aureus and Staphylococcus epidermidis were identified from the samples. Percentage occurrences of Staphylococcus species showed that 62.50% were Staphylococcus aureus isolates, while 37.50% were other species. The antibiotics susceptibility test showed that Staphylococcus aureus were found to be 100% susceptible to Gentamycin (CN) and where 50, 30 and 10% resistant to Norfloxacin (NB), Rifampicin (RD) and Chloramphenicol (CH) respectively. In conclusion, this research implies that most of the Awara samples collected from the vendors are not fit for consumption. It is important to keep Awara fit for consumption by taking adequate measures to prevent contamination during and after production.
Keywords
Food, Staphylococcus species, Antibiotic resistance, Bacterial count.
Introduction
Diseases caused by food pathogens contaminate food at any stages of food production, delivery and consumption in a food chain. The diseases can result from several forms of environmental contamination including water, soil, or air pollution, as well as unsafe food storage and processing [1]. Diseases transmitted through contaminated food can lead to public health problem [2]. These diseases contribute significantly to the global burden of disease and mortality. Annually, millions of persons become ill from foodborne diseases. In Nigeria and other parts of developing world, many of the foodborne cases ranging from mild to severe cases have always been reported [3].
Food poisoning cause by Staphylococcus species is a global challenge which has cause so many illnesses among the populations of the world and has also resulted to high cost of treatments and so many deaths [4]. In Nigeria, there are so many health cases which are attributed to consumption of foods that are contaminated by Staphylococcus species [5]. Most of the food samples hawked by the road sides in Nigeria are vulnerable to various source of contamination [6].
‘Awara’ is a local fried soya beans cake or cheese, commonly sold in northern Nigeria and is a cheap source of protein which is mostly consumed among middle class consumer and the poor classes of the northerners. It can also be taken at any time of the day as a snack [4-7]. The challenges with the production of ‘Awara’ is that they are usually fried by the road sides which predispose to contaminations by various pathogenic microorganisms most of which may be associated with foodborne poisonings [4] and sometimes the vendors used their hands to serve their customers instead of using spoons or fork where Staphylococcus species may be transmitted to the food and contaminate it [5].
Antibiotic resistance among Staphylococcus species has been a serious public health challenge, especially in the selection of drugs for the treatment of infections cause by this pathogen [8]. Staphylococcus species from foods have developed mechanisms to resist antibiotics; this can pose danger in the treatment of illness cause by these pathogens [9]. Therefore, the purpose of this research is to determine the occurrence and antibiotic resistance of Staphylococcus species in the Awara samples.
Materials and Methods
Study area
The study was conducted in Wukari local government area of Taraba state, which is located in North–Eastern region of Nigeria at southern guinea savanna with coordinate’s latitude 7°, 51’N-7.850°N and longitude 9°, 47’E-9.783°E, annual precipitation of 1205 mm and it has an average temperature of 26.8°. This research took place during the raining season 2021.
Collection of samples
A total of number twelve (12) samples were collected from four (4) locations of the Wukari old market. Five (5) samples from each of the four locations from the vendors/hawkers within the market. The samples were collected in sterile polythene bags directly from the sellers and immediately taken to the laboratory of the Department of Microbiology, Federal University Wukari for microbial analysis.
Total bacteria count
Total bacterial count was carried out using pour plate method [10,11]. Twenty-five gram of the food sample was placed in sterile 225 ml buffer peptone water to obtain a ratio of 1:10 and was thoroughly mixed, from which 1 ml was transferred to the first test tube containing 9 ml of buffer peptone water as diluent. This was repeated in other sets of the tubes containing 9 ml of the buffer peptone water to dilute to 10- 8. The procedure was repeated for each sample. From the last two dilutions, 0. 1 ml each was dispensed to the center of Petri dishes. Prepared and cooled molten nutrient agar and mannitol salt agar were poured into the Petri dishes, gently swirl and allowed to solidified and incubated at 37°C for 24 hours. At the end of incubation, plates with 30-300 colonies were counted using a colony counter and average was taken.
The colony forming unit was calculated as:
CFU/ml = Mean count per plate × dilution factor
Volume of sample plated
Isolation of Staphylococcus species
The medium mannitol salt agar was used for isolation and enumeration of Staphylococcus species. Gram staining technique was carried out to identify the morphology of the pathogens [12]. The biochemical tests carried used to identify Staphylococcus species include catalase, coagulase, citrate utilisation, haemolysin, indole, oxidase and urease tests.
Standardization of inoculum
Dilution from each of the suspension of the test isolates was prepared by picking a 24 h colony of the isolates using sterile wire loop into sterile test tube containing sterile normal saline to form turbidity that match with 0.5 scale of McFarland’s standard (1.5 × 108 cells/ml) [12].
Antibiotic susceptibility test
Staphylococcus isolates were subjected to antibiotics to determine their susceptibility by means of disc diffusion method [13]. Broth culture of the test organisms’ equivalent to 0.5 McFarland standards were used to inoculate plates of Mueller-Hinton agar using sterile swabs. Antibiotic impregnated discs were placed on the plates of the pathogens and incubated at 37°C for 24 hours. Zones of inhibitions were measured and the results were interpreted [14] and all the results were recorded appropriately.
Results
The total bacterial count and Staphylococcus species count in the Awara sampled from four locations of old market Wukari as presented in Table 1 showed that the total bacterial count in the food samples ranged from 1.00 x 105 to 1.80 x 106 CFU/ ml. The staphylococcal count on mannitol salt agar ranged from 1.2 x 105 to 8.0 x 105 CFU/ml.
| Sample | Total Bacteria Count (in CFU) | Staphylococcuscount (in CFU) |
|---|---|---|
| Vendor 1 | ||
| 1 | 8.0x105 | 4.8x105 |
| 2 | 1.00x105 | 2.6x105 |
| 3 | Too numerous | 8.0x105 |
| 4 | 6.4x105 | 6.0x105 |
| 5 | 1.28x105 | 1.2x105 |
| Vendor 2 | ||
| 1 | 1.26x105 | 3.2x105 |
| 2 | 1.80x106 | 6.0x105 |
| 3 | Too numerous | 4.0x105 |
| 4 | No growth | No growth |
| 5 | Too numerous | No growth |
| Vendor 3 | ||
| 1 | 1.40x105 | 5.4x105 |
| 2 | 4.6x105 | 3.2x105 |
| 3 | 1.00 x105 | 6.4x105 |
| 4 | 1.10 x105 | 4.0x105 |
| 5 | 1.6 x105 | 1.2x10-5 |
| Vendor 4 | ||
| 1 | 6.6x105 | 1.6x105 |
| 2 | 1.40x105 | No growth |
| 3 | Too numerous | 6.2x105 |
| 4 | No growth | No growth |
| 5 | 4.0x105 | 3.2x105 |
Table 1. Total bacteria count and Staphylococcus count in the food samples.
The morphological and biochemical characteristics of Staphylococcus species isolated from Awara sold in Wukari old market revealed that two species (Staphylococcus aureus and Staphylococcus epidermidis) were identified as presented in Table 2.
| Morphology | Grams | Biochemical characteristics | Isolates | |||
|---|---|---|---|---|---|---|
| Catalase | Coagulase | Citrates | Urease | |||
| Smooth,creamy, small, flat, opaque, entire | +cocci | + | + | + | + | Staphylococcus aureus |
| Smooth,creamy, small, flat, opaque, entire | +cocci | + | + | - | + | Staphylococcus epidermidis |
Table 2. Biochemical characteristics of all Staphylococcus species isolated from the food sample.
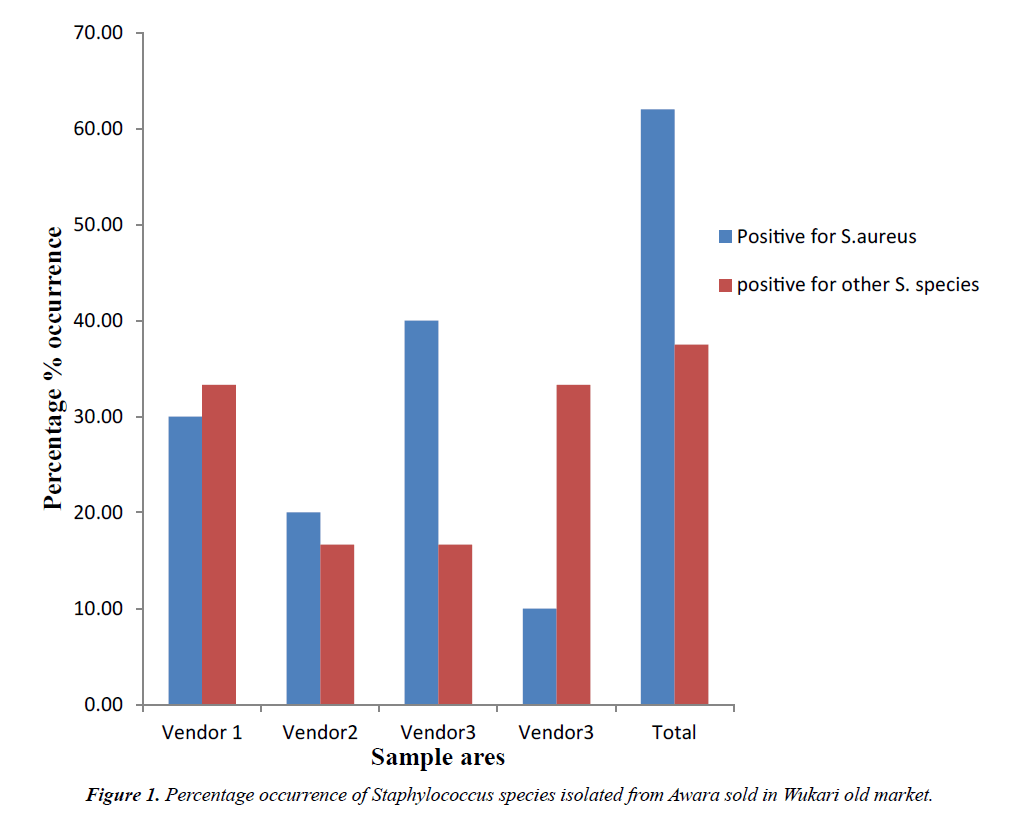
Percentage occurrences of Staphylococcus species isolated from Awara sample sold in Wukari old market is presented in Figure 1. The percentage occurrences of Staphylococcus species isolated from Awara sample sold in Wukari old market shows that 62.5% was Staphylococcus aureus while other Staphylococcus species was 37.5%.
Antibiotics susceptibility of Staphylococcus species isolated from the food sample as presented in Table 3 shows that the isolates were 100% susceptible to Gentamycin (CN), none of the isolates was susceptible to Ciprofloxacin (CPX), Erythromycin (E), Levofloxacin (LEV), Ampiclox (APX) and Amoxil (AML). Fifty (50%), Thirty (30%) and Ten (10%) were resistant to Norfloxacin (NB), Rifampicin (RD) and Chloramphenicol (CH) respectively.
| S/N | Antibiotics | S(%) | I(%) | R(%) |
|---|---|---|---|---|
| 1 | CPX | 0(0.0) | 10(100) | 0(0.0) |
| 2 | E | 0(0.0) | 10(100) | 0(0.0) |
| 3 | LEV | 0(0.0) | 10(100) | 0(0.0) |
| 4 | CN | 10(100) | 0(0) | 0(0.0) |
| 5 | APX | 3(30) | 7(70) | 0(0.0) |
| 6 | RD | 0(0.0) | 7(70) | 3(30) |
| 7 | AML | 5(50) | 5(50) | 0(0.0) |
| 8 | S | 9(90) | 1(10) | 0(0.0) |
| 9 | NB | 2(20) | 3(30) | 5(50) |
| 10 | CH | 2(20) | 7(70) | 1(10) |
Table 3. Antibiotic susceptibility of Staphylococcus species isolated from the food sample.
Key S= Sensitive, I= Intermediates, R= Resistance, CPXCiprofloxacin, NB-Norfloxacin, CN-Gentamycin, AMLAmoxil, S-Streptomycin, RD-Rifampicin, E-Erythromycin (30 mcg), CH-Chloramphenicol, APX-Ampiclox and LEVLevofloxacin.
Antibiotics resistance pattern of Staphylococcus species isolated was obtained from the food sample indicates two (2) isolates were resistant to Norfloxacin (NB), two (2) were also resistant to Rifampicin (RD) and Chloramphenicol (CH). While one of the isolates was resistant to Norfloxacin (NB), Rifampicin (RD) and Chloramphenicol (CH) respectively (Table 4). Key NB=Norfloxacin, RD=Rifampicin, CH=Chloramphenicol. The multiple antibiotics resistance index of Staphylococcus species indicates that the two isolates had MAR index of 0.2 and one of the isolates had MAR index of 0.3 (Table 5).
| S/N | Pattern | Frequency (No. of isolate) |
|---|---|---|
| 1 | NB | 2 |
| 2 | RD, NB | 2 |
| 3 | RD, NB, CH | 1 |
Table 4. Antibiotic resistance patterns of Staphylococcus species isolated from the food sample.
| Number of antibiotic multi-resistant isolates | Antibiotic combination | MAR index |
|---|---|---|
| 2 | 2 | 0.2 |
| 1 | 3 | 0.3 |
Table 5. Multiple antibiotic resistance (MAR) index of Staphylococcus species isolated from the food samples.
Discussion
Staphylococcus aureus is an important food poisoning organism because it’s widely distribution in nature. Staphylococcus aureus could be transmitted to foods when contaminated food products are used as constituents in the processing of the foods. The isolation of S. aureus in Awara samples implies extreme contamination and potential health risk associated with these food samples [15].
The processing of this food normally involves heat treatment; it is obvious that considerable number of bacteria associated with raw materials would have been killed or suppressed during production [16]. From this research result there was high bacterial load on the food sample collected, this could occur as a result of contamination due to exposure and the handling of the food [17]. Therefore, the high microbial load on the foods could be due poor sanitary conditions during preparation, storage and personal hygiene of the food handlers [18].
High occurrence of Staphylococcus species among the samples examined is an indication of frequent handling since the organism is a normal skin flora of humans. The result of this research is in agreement with the results observed by Yusha’u et al. [19], who in his studies confirmed high occurrence Staphylococcus aureus (85%), and [20]. The present research does not agree with the findings of Bristone et al. who isolated Staphylococcus aureus (37.5%) from Awara sold at University of Maiduguri Campus [21], who in their studies confirmed Staphylococcus aureus (30%) from soya beans cake (Awara) vended in Kano metropolis, Northern Nigeria.
The drug resistance among the Staphylococcus species could be due indiscriminate used of the antibiotics in the community where the bacteria were isolated [22]. The Multiple Antibiotic Resistances (MAR) index used in this study was 0.1 MAR index. This gives an indirect suggestion of the probable source of the organism and MAR index greater than 0.2, it indicates that the organism must have originated from an environment where these antibiotics are frequently used [22].
The antimicrobial resistance patterns of Staphylococcus aureus from foods are useful in epidemiological studies, as it provides adequate awareness about the safety of consumable foods contaminated with high level of potential microbes for food borne disease outbreak, resulting to a distribution that cannot be controlled and thus bringing about changes in public health and deterioration thereby increasing susceptibility rate [23].
Conclusion
Total bacterial count in the food samples ranged from 1.00 x 105 to 1.80 x 106 CFU/ml and the staphylococcal count on mannitol salt agar ranged from 1.2 x 105 to 8.0 x 105 CFU/ ml. The percentage occurrence of Staphylococcus species showed that 62.50% were Staphylococcus aureus isolates, while 37.50% were other species. None of the Staphylococcus species were ciprofloxacin, erythromycin, levofloxacin, gentamycin, ampiclox amoxil, streptomycin and some species were resistant to rifampicin (30%), rorfloxacin (50%), and chloramphenicol (10%). Two of the isolates had MAR index of 0.2 and one of the isolates had MAR index of 0.3. The bacterial load and presence of some resistant Staphylococcus species in the food sample can pose threat to human health in the study area. It is therefore, important to keep Awara fit for consumption by taking adequate measures to prevent contamination during and after production.
References
- Van der Goot AJ, Pelgrom PJ, Berghout JA, et al. Concepts for further sustainable production of foods. J Food Eng. 2016;168:42-51.
- Centers for Disease Control and Prevention, CDC. US Foodborne Disease Outbreaks: Available at: http://www.cdc.gov/foodborneoutbreaks/outbreak_data.htm. 2009.
- Odeleye OP, Whong CM, Jatau ED. Characterization and antibiotic susceptibility pattern of Bacillus cereus isolates from fried soyabean cake in Zaria, Nigeria. Sci J Microbiol. 2014;3(4):38-44.
- Bristone C, Mshelia HA, Ogori FA, et al. Microbial Quality Evaluation of Awara (Soybean Cheese) Processed and sold at University of Maiduguri Campus. J Bacteriol Mycol. 2018;6(1):00172.
- Okoli CE, Njoga EO, Enem SI, et al. Prevalence, toxigenic potential and antimicrobial susceptibility profile of Staphylococcus isolated from ready-to-eat meats. Vet World. 2018;11(9):1214.
- Daniyan SY, Abalaka ME, Momoh JA, et al. Microbiological and physiochemical assessment of street vended soyabean cheese sold in Minna, Nigeria.
- Zumbes JH, Dabo AD, Dakul DA, et al. Enteropathogenic bacterial contamination of some ready to eat foods sold in Jos Metropolis, Nigeria. Indian J Appl Res. 2014;4(7):456-8.
- Gandra S, Barter DM, Laxminarayan R. Economic burden of antibiotic resistance: how much do we really know?. Clin Microbiol Infec. 2014;20(10):973-80.
- Tenover FC. Mechanisms of antimicrobial resistance in bacteria. Am J Medi 2006;119(6):S3-10.
- Kawo AH, Dabai YU, Manga SB, et al. Prevalence and public health implications of the bacterial load of environmental surfaces of some Secondary Schools in Sokoto, North-western Nigeria. Int Res J Microbiol. 2012;3(5):186-90.
- Owolabi JB, Ichoku CK. Evaluation of antibiotic resistance pattern of gram-positive bacilli isolated from ready-to-eat vegetables sold in ota metropolis, Nigeria. Cov J Phy Life Sci. 2013:138.
- Cheesbrough M. Medical laboratory manual for tropical countries. M. Cheesbrough, 14 Bevills Close, Doddington, Cambridgeshire, PE15 OTT.; 1981.
- Hudzicki J. Kirby-Bauer Disk Diffusion Susceptibility Test Protocol (online). Available:www.microbelibrary.org/component/resource/laboratorytest/3189/kirbybauer-disk-diffusion-susceptibilitytest-protocol. 2013
- CLSI. Performance standards for Antimicrobial Susceptibility Testing; Twenty-Second Informational Supplement. CLSI document M100-S22. Wayne, PA: Clinical and Laboratory Standards Institute. 2016.
- Gizaw Z. Public health risks related to food safety issues in the food market: a systematic literature review. Environmental health and preventive medicine. 2019;24(1):1-21.
- Biranjia-Hurdoyal S, Latouche MC. Factors affecting microbial load and profile of potential pathogens and food spoilage bacteria from household kitchen tables. Can J Infect Dis Med Microbiol. 2016;2016.
- Bagumire A, Karumuna R. Bacterial contamination of ready-to-eat meats vended in highway markets in Uganda. Afr J Food Sci. 2017;11(6):160-170.
- Makinde OM, Ayeni KI, Sulyok M, et al. Microbiological safety of ready-to-eat foods in low- and middle-income countries: A comprehensive 10-year (2009 to 2018) review. Compr Rev Food Sci Food Saf. 2020; 19(2):703-32
- Yusha’u M, Atabor, Taura DW, et al. Bacteriological quality of soya bean cake (awara) sold at bayero university, Kano-Nigeria. International conference on advances in engineering and technology (RTET-2017), 2017; 173-175.
- Sina H, Baba-Moussa F, Kayodé AP, et al. Characterization of Staphylococcus aureus isolated from street foods: Toxin profile and prevalence of antibiotic resistance. J Applied Biosci. 2011; 46:3133-143
- Idris A, Dabo NT. Microbiological and nutritional analysis of soya beans cake (Awara) and Camel milk cheese (Chukwui) local snacks, vended in Kano, Metropolis-Nigeria. J Mirobiol Res. 2016;1:7-14
- Hammuel C, Jatau ED, Whong CMZ. Prevalence and antibiogram pattern of some nosocomial pathogens isolated from hospital environment in Zaria, Nigeria. Aceh Int J Sci Technol. 3(3):131-139.
- Sahota P, Jairath S, Pandove G, et al. Emerging food borne pathogens-A review. Asian J Microbiol Biotechnol Environ Sci. 2008; 4:921-926.
Indexed at, Google Scholar, Cross ref
Indexed at, Google Scholar, Cross ref
Indexed at, Google Scholar, Cross Ref
Indexed at, Google Scholar, Cross Ref
Indexed at, Google Scholar, Cross Ref
Indexed at, Google Scholar, Cross Ref