Review Article - Journal of Food Microbiology (2017) Journal of Food Microbiology(Special Issue-2017)
The spoilage microorganisms in seafood with the existed quorum sensing phenomenon
Feifei Wang, Linglin Fu, Xingyue Bao and Yanbo Wang*
Key Laboratory for Food Microbial Technology of Zhejiang Province, School of Food Science and Bioengineering, Zhejiang Gongshang University, Hangzhou, China
- *Corresponding Author:
- Yanbo Wang
18 Xuezheng road
Xiasha University Town
Hangzhou, 310018, China
Tel: +86-571-28008963
E-mail: wyb1225@163.com
Accepted on September 27, 2017
Citation: Wang F, Fu L, Bao X, et al. The spoilage microorganisms in seafood with the existed quorum sensing phenomenon. J Food Microbiol 2017;1(1):14-19.
Abstract
Most food products are highly perishable as they constitute a rich nutrient source for microbial development. Seafood is one of the most highly perishable food products due to the chemical effects of atmospheric oxygen and the growth of spoilage microorganisms. Therefore, the spoilage of food depends up on the physiological state of spoilers and on their ability to resist the processing/storage conditions. In addition, spoilage relies on the density of the population and the interactions between the microorganisms composing the ecosystems of seafood involving quorum sensing. This review mainly introduces the SSOs of seafood under different preserve conditions, the spoilage microorganisms employing quorum sensing system, and describes the relationship between quorum sensing and spoilage potential in these microorganisms.
Keywords
Seafood, Spoilage microorganisms, Quorum sensing.
Introduction
Seafood is one of the most highly perishable food products because of the chemical effects of atmospheric oxygen and the growth of spoilage microorganisms [1]. Spoilage of seafood can be caused by enzymes, dehydration, oxidation, contamination and physical damage. Sulphurous, ammoniacal, or fishy odours are some of the main organoleptic changes taking place during spoilage development [2].
The major cause of seafood spoilage is microbial growth and metabolic activity which result in the formation of amines, sulphides, alcohols, aldehydes, ketones, and organic acids with unpleasant and unacceptable off-flavours [3].
However, only a fraction of the initial microbiota of seafood known as specific spoilage organisms (SSOs), which is favoured by storage conditions (e.g., atmosphere, temperature), prevails over the rest of the microbiota, reaching high populations and producing corresponding metabolites (biochemical spoilage indices) [4].
Quorum sensing (QS), which involves the production, release and community-wide detection of extracellular signaling molecules called autoinducers, is a cell-to-cell communication process enabling microorganisms to collectively alter behavior patterns upon changes in cell density and species composition in surrounding community.
When a threshold concentration of the signaling molecule is reached, the group detects and responds to it with a populationwide alteration in gene expression. Therefore, QS-controlled processes, such as bioluminescence, the secretion of virulence factors, biofilm formation and the production of public goods, require the collective action of the group to be effective [5].
The most commonly studied autoinducers of QS signals include N-acyl-L-homoserine lactones (AHLs) in Gram-negative bacteria, oligopeptide in Gram-positive bacteria, and autoinducer-2 (AI-2) used in both Gram-negative and Grampositive bacteria [6].
Beyond these classes, recently, a range of cyclic dipeptides (diketopiperazines, DKPs) produced by multiple Gram-negative bacteria were reported to modulate supposedly AHLs-specific sensor system [7-9].
Therefore, DKPs have been suggested to represent a new class of naturally occurring QS signals, and to potentially play a role in both intra- and interspecies QS regulation [10,11].
The physiological and clinical aspects of QS have attracted considerable attention and been studied at the molecular level. However, there is a lack of knowledge on the role of QS in food spoilage. As cell-to-cell communication occurs in diverse bacterial species, QS likely plays a role in the microbial ecology of foods [3].
Main Spoilage Microorganisms in Different Seafood and Seafood Products
The recent establishment of the SSO concept has contributed significantly to our understanding of seafood spoilage [12].
The growth of different SSOs depends on several parameters: food product, type of preservation, temperature, atmosphere, and salt content, among others. During storage, the microflora changes owing to different abilities of the microorganisms to tolerate the preservation conditions [13].
Here, the storage conditions of seafood were divided into two major categories, and the spoilage microorganisms predominant in both conditions were discussed and listed in Table 1, respectively.
| Spoilage bacteria | Sea foods and seafood products | References |
|---|---|---|
| Shewanella spp. | Gutted sea bass | [4] |
| Iced sea salmon | [15] | |
| Air stored swordfish | [17] | |
| Refrigerated shrimp | [19] | |
| Refrigerated large yellow croaker | [20] | |
| Pseudomonas spp. | Gutted sea bass | [4] |
| Air stored swordfish | [17] | |
| Aeromonas spp. | Iced sea salmon | [15] |
| Photobacterim phosphoreum | MAP/VP stored raw salmon | [16] |
| VP packaged squid mantle | [23] | |
| Enterobacteriaceae | MAP/VP packaged swordfish | [17] |
| VP packaged pressurised squid mantle | [23] | |
| LAB | MAP/VP stored raw salmon | [16] |
| MAP/VP packaged swordfish | [17] | |
| VP packaged pressurised squid mantle | [23] |
Table 1. Main spoilage microorganisms in different seafood and seafood products.
Fresh Seafood Stored in Ice or under MAP/VP
In newly caught marine seafood from temperate waters, microflora is formed mainly by aerobic rods-shapes, anaerobic facultative and psychrotrophic Gram-negative bacteria, whose growth is possible at 0 and optimal at around 25. The majority belongs to the Gammaproteobacteria: Pseudomonas, Shewanella, Acinetobacter, Aeromonas, Vibrio, Moraxella, Psychrobacter, Photobacterium, etc. The same bacterial genus can be found in tropical marine seafood, but Cram-positive bacteria, Enterobacteriaceae and Vibrionaceae are often dominant [14]. Generally, Pseudomonas spp., S. putrefaciens, S. baltica or Aeromonas spp. were common dominant spoilage bacteria in iced sea salmon [15,16]; gutted sea bass [4]; chilled fresh Mediterranean swordfish [17]; tropical prawns [18,19]; large yellow croaker [20]. Pseudoalteromonas and Vibrio were dominant microorganisms in shucked oysters during iced-storage [21], as spoilage proceeded, enterococci, lactobacilli, and yeasts dominated at the later stages [22], the spoilage patterns of Mollusca shellfish differ in most species of seafood as they contain high levels of carbohydrate in the form of glycogen [6]. Modified Atmosphere Packaging (MAP) and Vacuum-Packaging (VP), along with refrigeration, have become increasingly popular preservation techniques. Dominant strains isolated from spoiled squid were identified as Photobacterium phosphoreum [23]. Bacteria grew faster under aerobic conditions, while the increase of CO2 and O2 reduction in MAP inhibited the bacterial growth and changed the microbial spoilage by suppressing mostly the Gram negatives and favouring the Gram positives [4]. P. phosphoreum and L. piscium were identified as the main bacterial groups in MAP/VP raw salmon [16]. The main SSO of modified atmosphere packaged Norway lobster is P. phosphoreum, since P. phosphoreum is known to withstand high CO2 concentrations [24]. Lactic Acid Bacteria (LAB) and Brochothrix thermosphacta were co-dominant with Pseudomonas and H2S producing bacteria in gutted sea bass stored at 2 under MAP [4]. Carnobacterium. maltaromaticum was the organism that showed the highest resistance to CO2 and to the lack of O2 among the organisms responsible for spoilage in mackerel fillets packed under modified atmospheres [25].
Lightly preserved seafood
Lightly preserved seafood are uncooked or mildly cooked products with low level preservatives which can influence their aw, pH, including brined/pickled/marinated seafood, cooked and peeled shrimp and shucked shellfish stored in MAP/VP or in brine, cold-smoked fish, etc. As a result, aerobic Gramnegative bacteria are inhibited, which allows the growth of other organisms more resistant to reduced aw [14].
Psychrobacter spp. and Pseudoalteromonas spp. were the dominant microbiota of cooked brown shrimp and enhanced spoilage by breaking down lipids and hydrolysing amino acids and proteins [26]. The major spoilage bacterial isolates from spoiled cooked and whole tropical shrimp stored under MAP were C. maltaromaticum and S. baltica [27]. LAB and Brochothrix spp. were dominant bacteria in the latter storage period of the VP-packed cold-smoked salmon, whereas Brochothrix spp. rather than LAB were responsible for spoilage [28]. Differently, Joffraud et al. [2006] identified L. sakei and S. liquefaciens-like as the most spoiling bacteria. Besides, psychrotrophic marine vibrio and Photobacterium spp. were reported to be dominant microflora [29]. The different spoilage microorganism’s profiles of cold-smoked salmon may result from the different treatments and environment.
In conclusion, the microflora changes owing to different abilities of the microorganisms to tolerate the storage conditions. Pseudomonas spp. and a few other Gram-negative psychrotrophic organisms will dominate seafoods stored aerobically at chill temperatures. CO2 packing or vacuum packing will inhibit the respiratory pseudomonads and cause a shift in the microflora to P. phosphoreum, LAB, Enterobacteriaceae and sometimes B. thermosphacta. Increasing the preservation by a decrease in pH, an increase in the NaCl concentration and by adding low level preservatives eliminates the Gram-negative microflora, LAB is the remaining organisms in semi-preserved fish products.
QS Regulated Seafood Spoilage
The physiological and clinical aspects of QS have attracted considerable attention and been studied at the molecular level. However, there is a lack of knowledge on the role of QS in food spoilage, especially in seafood. As cell-to-cell communication exists in diverse bacterial species, QS likely plays a role in the microbial ecology of foods [3]. In the past few years, the possible role of QS in food spoilage has been explored, including siderophore synthesis, metabolic activities and biofilm formation, predominantly.
The Siderophores synthesis
All aerobic and facultative anaerobic bacteria require iron for growth and only LAB do not depend on supplementation of this mineral [30]. In fish muscle, the environment is iron-limited despite of the rich nutrient and high affinity chelators, the so-called siderophores are produced to scavenge ironduring bacterial growth. Although fish tissue allowed the siderophore production by most Pseudomonas and S. putrefaciens isolates from fish, S. putrefaciens was inhibited by Pseudomonas sp. particularly when iron was limited [8]. Later, the biosynthesis of siderophore in Pseudomonas aeruginosa was firstly reported to be controlled by QS system, lasR mutants showed a reproducible 2-fold decrease in production of the catecholate-hydroxamate siderophore pyoverdine during grown under iron-limited conditions. Similarly, lasI mutants defective in the biosynthesis of the autoinducer PAI-1 also had a 2-fold decrease in pyoverdine production which could be largely restored upon addition of exogenous PAI-I [31].
It was reported that exogenous AHL was required for the stimulated biosynthesis of heterologous siderophore in marine-isolated bacteria, and stimulated growth by exogenous siderophores and AHLs was also observed in other non-siderophore- producing bacteria [32]. Rasch et al. have reported that bacterial spoilage of bean sprouts was influenced by QS, the AHL-negative mutant of Enterobacteriaceae was impaired in siderophore activities and spoilage potential, for the first time demonstrating that iron chelation in Enterobacteriaceae was regulated by AHL [33]. These reports offer a new perspective for exploring seafood spoilage mediated by intra and inter-species cell-cell communication (Figure 1), although little study has focused on the regulation of QS on the siderophore-associated spoilage in seafood.
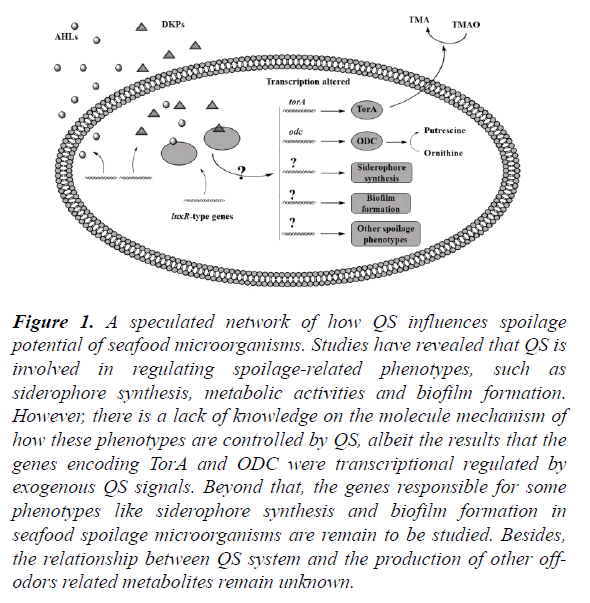
Figure 1. A speculated network of how QS influences spoilage potential of seafood microorganisms. Studies have revealed that QS is involved in regulating spoilage-related phenotypes, such as siderophore synthesis, metabolic activities and biofilm formation. However, there is a lack of knowledge on the molecule mechanism of how these phenotypes are controlled by QS, albeit the results that the genes encoding TorA and ODC were transcriptional regulated by exogenous QS signals. Beyond that, the genes responsible for some phenotypes like siderophore synthesis and biofilm formation in seafood spoilage microorganisms are remain to be studied. Besides, the relationship between QS system and the production of other off-odors related metabolites remain unknown.
Metabolic activities
As described before, spoilage microorganisms are responsible for various sensory deterioration, but these sensory descriptors are not easily associated with enzymatic functions or metabolic pathways. Several studies have reported the detection or measurement of molecules (biogenic amines and volatile compounds) in seafood spoiled by known microorganisms. However, it is difficult to correlate the production of spoilage-related metabolites to the functions of spoilers, and the studies investigating the metabolism and physiology of bacteria responsible for seafood spoilage are much less abundant, not to mention the studies on these spoilage phenomena caused by spoilers employing QS system. The QS system involved in metabolic activity in microorganisms of seafood was listed in Table 2.
| Bacterial group/species | Seafood | Production of | Phenotypes regulated by QS | References | ||
|---|---|---|---|---|---|---|
| AHLs | AI-2 | DKPs | ||||
| Shewanella | Shrimp | - | + | + | TVB-N | [19] |
| volatile organic components | ||||||
| Extracellular proteases | ||||||
| Biofilm formation | ||||||
| Fish | - | + | + | TVB-N | [38] | |
| TMA | ||||||
| Putrescine | ||||||
| Biofilm formation | ||||||
| Pseudomonas | Fish | + | nr | nr | Exoenzyme production | [2] |
| Biofilm formation | [40] | |||||
| Enterobacteriaceae | Fish | + | nr | nr | Exoenzymic activities | [2] |
| [39] | ||||||
| Nr: not reported | ||||||
Table 2. QS System involved in microorganism spoilage in seafood.
Biogenic amines (BAs) are low molecular weight organic bases that possess biological activity. BAs are basic nitrogenous compounds formed mainly by decarboxylation of amino acids or by amination and transamination of aldehydes and ketones [34]. BAs can be divided into three groups according to chemical structure: aliphatic (putrescine, cadaverine, spermine, spermidine); aromatic (tyramine, phenylethylamine); heterocyclic (histamine, tryptamine). In seafood, BAs are formed due to the presence of decarboxylase-positive microorganisms, and conditions that allow bacterial growth, decarboxylase synthesis and decarboxylase activity [34].
Tri Methyl Amines (TMA) is formed from bacterial use of TMAO which is found in most marine fish species [35]. Pseudomonas spp. cannot use TMAO and produce no TMA on spoiling fish. SSOs such as Aeromonas spp., Photobacterium phosphoreum, Shewanella putrefaciens-like organisms, Enterobacteriaceae and Vibrio spp. are all capable of using TMAO as final acceptor of electrons and produce TMA, causing “fishy” odours associated with seafood spoilage [21,36]. TMAO is reduced to TMA by the enzyme TMAO reductase, encoded by the torCAD operon [37]. In Shewanella baltica, the SSO of refrigerated large yellow croaker (Pseudosciaena crocea), the TMA and putrescine were significantly increased in the presence of cyclo-(L-Pro-L-Leu), the transcription levels of torA and ornithine decarboxylase (ODC) were upregulated in accordance with the spoilage phenotypes [20].
As described in the previous section that DKPs were suggested as QS signals, it seems that QS system was involved in the S. baltica spoilage through regulating TMA and putrescine production. Similarly, the production of total volatile base nitrogen (TVB-N) in sterile fish muscle juice inoculated with S. baltica was significantly improved by synthetic DKPs supplementation [38]. AHLs and cyclo-(L-Pro-L-Leu) were reported to promote the extracellular proteolytic activities in SSO of refrigerated shrimp (Litopenaeus vannamei), and to increase the levels of TVB-N and the volatile organic components in the shrimp samples [19]. AHL-modulated exoenzymic activities have been reported in Serratia proteamaculans B5a isolated from cold-smoked salmon, and the lipB-encoded secretion system was identified as one target gene of the QS system.
LipB was required for the production of extracellular lipolytic and proteolytic activities, thus rendering the production of food-deterioration-relevant exoenzymes indirectly under the control of QS [39]. In addition, the C4-HSL signaling molecule produced by the isolate strain Pseudomonas psychrophila PSPF19 played a role in spoilage of freshwater fish stored in refrigerated conditions via inducing exoenzyme production by twofold [40]. However, there is a lack of knowledge on the relationship between QS system and the production of other off-odors related metabolites, it can be the consequence of a complex succession of enzymatic reactions, potentially associated with non-enzymatic reactions, the spoilage can also result from reactions catalyzed by enzymes that are not well defined (Figure 1).
Biofilm formation
Biofilms are formed by bacteria attached to surfaces, which upon their aggregation release extracellular polysaccharides that form a polymeric matrix or glycocalyx. Biofilms are architecturally complex structures made up of microcolonies and characteristic mushroom or pillar-like arrangements that are separated by channels that permit the circulation of water and nutrients [41]. There are many reports on QS regulating biofilm formation in pathogens. Davies et al. have first suggested the control of biofilm differentiation and integrity by las QS in Pseudomonas aeruginosa in vivo and in vitro, which makes an inextricable connection between QS and biofilm formation [42]. In Streptococcus mutans dependent on the ComCDE QS system, biofilms formed by the comC mutant that did not produce CSP had a reduced biomass, and conversely, adding synthetic CSP into the culture restored the wild-type biofilm [43]. The agr QS system was reported to play a role in biofilm development in Staphylococcus aureus [44]. The luxS-controlled quorum-sensing (QS) system was proved to play a major role in the control of Streptococcus pneumoniae biofilm formation [45]. As for microorganism spoilers, the molecule mechanism of QS-regulated biofilm remains to be studied, albeit some experiments were carried out in vitro. AHLs and/or DKPs were reported to promote the biofilm formation in Pseudomonas psychrophila PSPF19 and Shewanella baltica [19,20,40,46], and the QS system involved in biofilm formation in microorganisms of seafood was listed in Table 2. The molecular mechanisms of biofilm formation regulated by QS system in food spoilers needs further study (Figure 1).
Conclusions
Many studies have shown that cell density-dependent signalling systems in bacteria controls a range of phenotypic traits. However, there is little studies focus on the QS system in seafood spoilage, although there are many reports on the major spoilage microorganisms in seafood and the functional properties of these spoilers. Here, we list the spoilage microorganisms employing QS system and the corresponding spoilage phenotypes regulated by QS system in seafood in Table 2. Besides, according to the previous reports related to the role of QS system on group behaviour including cell growth and metabolic activities and biofilm formation, we speculate a molecule network of how QS influences spoilage potential of seafood microorganisms in Figure 1. The QS-involved spoiling mechanisms at molecule level still remain to be studied. It is important to have awareness and an understanding of the mechanisms involved in the bacterial quorum sensing, since preservatives targeting quorum sensing will offer a new means to control the proliferation of undesirable microorganisms in seafood.
Acknowledgment
This study was financially supported by the Zhejiang Provincial Natural Science Foundation of China (LZ15C200001) and National Natural Science Foundation of China (31571913 and 31772050). We thank Dr. Yang Zhu for the recommendation and instruction of the manuscript.
References
- Özogul F, Polat A, Özogul Y. "The effects of modified atmosphere packaging and vacuum packaging on chemical, sensory and microbiological changes of sardines (Sardina pilchardus)". Food Chem. 2004;85:49-57.
- Gram L, Ravn L, Rasch M, et al. "Food spoilage-interactions between food spoilage bacteria", Int J Food Microbiol. 2002;78(1-2):79-97.
- Robson A A, Kelly MS, Latchford JW. "Effect of temperature on the spoilage rate of whole, unprocessed crabs: Carcinus maenas, Necora puber and Cancer pagurus", Food Microbiol. 2007;24(4): 419-24.
- Parlapani FF, Haroutounian SA, Nychas GJ, et al. "Microbiological spoilage and volatiles production of gutted European sea bass stored under air and commercial modified atmosphere package at 2 degrees C", Food Microbiol. 2015;50: 44-53.
- Bassler B LLosick R. "Bacterially speaking", Cell, 2006;125(2): 237-46.
- Fernandez-Piquer J, Bowman JP, Ross T, et al. "Molecular analysis of the bacterial communities in the live Pacific oyster (Crassostrea gigas) and the influence of postharvest temperature on its structure", J Appl Microbiol. 2012;112(6): 1134-43.
- Holden MT, Ram CS, De NR, et al. "Quorum-sensing cross talk: isolation and chemical characterization of cyclic dipeptides from pseudomonas aeruginosa and other gram-negative bacteria", Mol Microbiol. 1999;33(6):1254-66.
- Degrassi G, Aguilar C, Bosco M, et al. "Plant growth-promoting Pseudomonas putida WCS358 produces and secretes four cyclic dipeptides: cross-talk with quorum sensing bacterial sensors", Curr Microbiol. 2002;45(4):250-4.
- Park D K, Lee K E, Baek C H, et al. "Cyclo(Phe-Pro) modulates the expression of ompU in Vibrio spp", J Bacteriol. 2006;188(6):2214-21.
- Shiner E K, Rumbaugh K P, Williams S C. "Inter-kingdom signaling: deciphering the language of acyl homoserine lactones", FEMS Microbiol Rev. 2005;29(5):935-47.
- Klose K E. "Increased chatter: cyclic dipeptides as molecules of chemical communication in Vibrio spp.", J Bacteriol. 2006;188(6):2025-6.
- Dalgaard P. "Qualitative and quantitative characterization of spoilage bacteria from packed fish", Int J Food Microbiol. 1995;26(3):319.
- Bekaert K, Devriese L, Maes S, et al. "Characterization of the dominant bacterial communities during storage of Norway lobster and Norway lobster tails (Nephrops norvegicus) based on 16S rDNA analysis by PCR-DGGE", Food Microbiol. 46:2015;132-8.
- Francoise L. "Occurrence and role of lactic acid bacteria in seafood products", Food Microbiol. 2010;27(6):698-709.
- Hozbor M C, Saiz A I, Yeannes M I, et al. "Microbiological changes and its correlation with quality indices during aerobic iced storage of sea salmon (Pseudopercis semifasciata)", LWT-Food Sci. Technol, 2006;39(2):99-104.
- Mace S, Cornet J, Chevalier F, et al. "Characterisation of the spoilage microbiota in raw salmon (Salmo salar) steaks stored under vacuum or modified atmosphere packaging combining conventional methods and PCR-TTGE", Food Microbiol. 2012;30(1):164-72.
- Pantazi D, Papavergou A, Pournis N, et al. "Shelf-life of chilled fresh Mediterranean swordfish (Xiphias gladius) stored under various packaging conditions:microbiological, biochemical and sensory attributes", Food Microbiol. 2008;25(1):136-43.
- Chinivasagam H N, Bremner H A, Wood A F, et al. "Volatile components associated with bacterial spoilage of tropical prawns", J Food Microbiol. 1998;42(1-2):45-55.
- Zhu S, Wu H, Zeng M, et al. "The involvement of bacterial quorum sensing in the spoilage of refrigerated Litopenaeus vannamei", Int J Food Microbiol. 2015;192:26-33.
- Zhu J, Zhao A, Feng L, et al. "Quorum sensing signals affect spoilage of refrigerated large yellow croaker (Pseudosciaena crocea) by Shewanella baltica", Int J Food Microbiol., 2016;217:146-55.
- Madigan T L, Bott N J, Torok V A, et al. "A microbial spoilage profile of half shell Pacific oysters (Crassostrea gigas) and Sydney rock oysters (Saccostrea glomerata)", Food Microbiol. 2014;38:219-27.
- Cao R, Xue C H, Liu Q. "Changes in microbial flora of Pacific oysters (Crassostrea gigas) during refrigerated storage and its shelf-life extension by chitosan", Int J Food Microbiol. 2009;131(2-3):272-6.
- Paarup T, Sanchez J A, Peláez C, et al. "Sensory, chemical and bacteriological changes in vacuum-packed pressurised squid mantle (Todaropsis eblanae) stored at 4 degrees C", International Journal of Food Microbiology,2002;74(1-2):1.
- Gornik S G, Albalat A, Theethakaew C, et al. "Shelf life extension of whole Norway lobster Nephrops norvegicus using modified atmosphere packaging", Int J Food Microbiol. 2013;167(3):369-77.
- Alfaro B, Hernández I, Le Marc Y, et al. "Modelling the effect of the temperature and carbon dioxide on the growth of spoilage bacteria in packed fish products", Food Control. 2013;29(2):429-37.
- Broekaert K, Noseda B, Heyndrickx M, et al. "Volatile compounds associated with Psychrobacter spp. and Pseudoalteromonas spp., the dominant microbiota of brown shrimp (Crangon crangon) during aerobic storage", Int J Food Microbiol. 2013;166(3):487-93.
- Macé S, Cardinal M, Jaffrès E, et al. "Evaluation of the spoilage potential of bacteria isolated from spoiled cooked whole tropical shrimp (Penaeus vannamei) stored under modified atmosphere packaging", Food Microbiol. 2014;40:9-17.
- Leroi F, Joffraud J J, Chevalier F, et al. "Study of the microbial ecology of cold-smoked salmon during storage at 8 degrees C", Int J Food Microbiol. 1998;39(1-2):111-21.
- Truelstrup Hansen L, Røntved S D, Henrik Huss H. "Microbiological quality and shelf life of cold-smoked salmon from three different processing plants", Food Microbiol. 1998;15(2): 137-50.
- Pandey A, Bringel F, Meyer J M. "Iron requirement and search for siderophores in lactic acid bacteria", Appl. Microbiol. Biotechnol. 1994;40(5):735-39.
- McCarter L L. "Polar flagellar motility of the Vibrionaceae", Microbiol Mol Biol Rev, 65(3):2001;445-62, table of contents.
- Bardocz S, Grant G, Brown D S, et al. "Polyamines in food-implications for growth and health", J Nutr Biochem. 1993;4(2):66-71.
- Rasch M, Andersen J B, Nielsen K F, et al. "Involvement of bacterial quorum-sensing signals in spoilage of bean sprouts", Appl Environ Microbiol. 2005;71(6): 3321-30.
- Ten B B, Damink C, Joosten H M, et al. "Occurrence and formation of biologically active amines in foods", Int J Food Microbiol. 1990;11(1):73.
- Koutsoumanis KNychas G J E. "Chemical and Sensory Changes Associated with Microbial Flora of Mediterranean Boque (Boops boops) Stored Aerobically at 0,3,7, and 10°C", Appl. Environ. Microbiol. 1999;65(2):698.
- Joffraud J J, Leroi F, Roy C, et al. "Characterisation of volatile compounds produced by bacteria isolated from the spoilage flora of cold-smoked salmon", Int J Food Microbiol. 2001;66(3):175-84.
- Méjean V, Iobbinivol C, Lepelletier M, et al. "TMAO anaerobic respiration in Escherichia coli: involvement of the tor operon", Mol. Microbiol. 1994;11(6):1169-79.
- Gu Q, Fu L, Wang Y, et al. "Identification and characterization of extracellular cyclic dipeptides as quorum-sensing signal molecules from Shewanella baltica, the specific spoilage organism of Pseudosciaena crocea during 4 degrees C storage", J Agric Food Chem. 2013;61(47): 11645-52.
- Christensen A B, Riedel K, Eberl L, et al. "Quorum-sensing-directed protein expression in Serratia proteamaculans B5a", Microbiol. 2003;149(Pt 2):471-83.
- Bai A JRai Vittal R. "Quorum Sensing Regulation and Inhibition of Exoenzyme Production and Biofilm Formation in the Food Spoilage BacteriaPseudomonas psychrophilaPSPF19", Food Biotechnol. 2014;28(4):293-308.
- Costerton J W, Stewart P S, Greenberg E P. "Bacterial biofilms: a common cause of persistent infections", Science. 1999;284(5418):1318-22.
- Davies D G, Parsek M R, Pearson J P, et al. "The involvement of cell-to-cell signals in the development of a bacterial biofilm", Science. 1998;280(5361):295-8.
- Li Y H, Tang N, Aspiras M B, et al. "A Quorum-Sensing Signaling System Essential for Genetic Competence in Streptococcus mutans Is Involved in Biofilm Formation", J. Bacteriol. 2002;184(10):2699-2708.
- Yarwood J M, Bartels D J, Volper E M, et al. "Quorum Sensing in Staphylococcus aureus Biofilms", J. Bacteriol. 2004;186(6):1838-50.
- Vidal J E, Ludewick H P, Kunkel R M, et al. "The LuxS-dependent quorum-sensing system regulates early biofilm formation by Streptococcus pneumoniae strain D39", Infect Immun. 2011;79(10):4050-60.
- Françoise L. "Occurrence and role of lactic acid bacteria in seafood products", Food Microbiol. 2010;27(6):698-709.