Research Article - Biomedical Research (2017) Volume 28, Issue 18
Comparison of TNF-α promoter region with TNF receptor 1 and 2 (TNFR1 and TNFR2) in susceptibility to pulmonary tuberculosis; by PCR-RFLP
Shima Saif1, Poopak Farnia1,2*, Ehsan Ghamari1, Jalaledin Ghanavi1, Parissa Farnia1 and Ali Akbar Velayati1
1Mycobacteriology Research Centre (MRC), National Research Institute of Tuberculosis and Lung Disease (NRITLD), Shahid Beheshti University of Medical Sciences, Tehran, Iran
2Department of Biotechnology, School of Advanced Technologies in Medicine, Shahid Beheshti University of Medical Sciences, Tehran, Iran
- *Corresponding Author:
- Poopak Farnia
Department of Biotechnology
School of Advanced Technologies in Medicine
Shahid Beheshti University of Medical Sciences
Tehran, Iran
Accepted on August 28, 2017
Abstract
Pulmonary tuberculosis, is a granulomatous disease of the lungs that caused by Mycobaterium tuberculosis. A number of genes have been identified in studies of diverse origins to be important in tuberculosis. Out of which the tumor necrosis factor α (TNF-α) gene has been identified to plays a major role in the initial and long-term control of tuberculosis. In this study, the TNF-α gene at -238, -308, -857 and -863 position and tumor necrosis factors receptors one and two (TNFR1, TNFR2) at A36G and T587G position were analyzed and compared, respectively. The genotypes was studied in confirmed TB patients (n=151) and control group (n=83) using PCR-RFLP. The results showed a strong correlation between two polymorphisms in different loci of TNF-α gene including TNF-α T-857 C and A238G. Although no association were found in TNF receptors and susceptibility to TB. Additionally, no correlation between TNF receptors and gene polymorphisms were found. In conclusions, among studied cases only single nucleotide polymorphisms (SNPs) at TNF-857 and TNF-238 regions were associated with susceptibility to tuberculosis.
Keywords
Pulmonary tuberculosis, Susceptibility, TNF alpha, TNRR1, TNFR2.
Introduction
The World Health Organization (WHO) has estimated that more than 1.7 billion people are infected with Mycobacterium tuberculosis (MTB), the causative agent of tuberculosis (TB) [1]. Of these infected individual, 9.6 million people were fallen ill (1.1 million HIV-negative and 0.4 million HIV-positive) and 1.5 million expired due to TB related diseases (the death toll compromised 890,000 men, 480,000 women and 140,000 children) [1-3]. Despite several attempts to control the disease, TB is too hard to control mainly due to chronic character, long term treatment and difficult vaccine preparation as well as more coincidence with HIV infection [4-8]. More than 90% of TB-related death occurs in developing countries where 75% of TB patients are among their most economically beneficent age range (15-54 y) [5-8]. The susceptibility to TB depends upon different factors and the risk of developing diseases after infection with M tuberculosis ranges from 5% to 10% [1,9]. This suggests that besides the mycobacterial itself, the host genetic factors may determine the differences in host susceptibility to TB [10-14]. Among the important risk factor, cytokines and specially tumor necrosis factor alpha (TNF-α) genes, are thought to be responsible in regulating the protective immune responses [9,14]. Tumor necrosis factor alpha (TNF-α), gene that encodes the cytokines TNF-α is located within the class III region of the MHC [14,5]. TNF-α works often through related gene transcription in terms of inflammation, cell division, cell differentiation and apoptosis using two types of receptors, TNFR1 and TNFR2 [15,16]. The former exists in almost all cells while the latter sounds more specific and more effective in cell survival. TNFR2 is one of the main receptors on CD4+ and CD8+ lymphocytes. Cranial Neural Crest cells (CNCs) and endothelial cells. CD4+ and CD8+ seem to be important in immune cell replication and differentiation to prevent the growth of intracellular microorganisms like M tuberculosis [17,18]. TNF-α gene and its receptors have significant suppressive effects on bacterial growth into macrophages. TNFR1 is more responsible when apoptosis is needed but TNFR2 is involved in cell survival and TNF-α conducts its replicative effects on immune cells via the latter [19-21]. Till date, several polymorphisms within the promoter region of TNF-α gene have been shown to be associated with susceptibility or resistance to TB in different ethnic groups [22-26]. In contrast, the co-relation of TNF-α gene with their receptors like TNFR2 in susceptibility to TB has not been resolved yet. In this study, we aimed to analysis the SNPs in TNF-α gene at -238, -308, -857 and -863 positions and compare the susceptibility to TB with polymorphisms at TNFR1 (A36G) and TNFR2 (T587G). To our knowledge this is the first report that investigates the TNF-α gene polymorphisms with their two receptors in susceptibility to pulmonary tuberculosis.
Materials and Methods
Study population
Through a cross-sectional study, 151 TB cases and 83 healthy people as controls were recruited and people with history of malignancy and/or autoimmune diseases were excluded. Patients and control subjects were matched for age, sex and nationality (The Institutional Review Board at the NRITLD approved the study and all the patients have signed informed consent). All the TB patients were confirmed by smear microscopy and culture examination.
DNA extraction
Four millilitres of peripheral blood were collected in EDTA containing tube and stored at -4°C until DNA extraction. Genomic DNA was extracted from peripheral blood leukocytes (PBLs), using the standard phenol-chloroform procedure with slight modifications [22,26,27].
TNF genotyping
Polymorphisms in the TNF promoter region, namely TNF single nucleotide polymorphisms (SNP), -238, -308, -857 and -863 were studied using PCR-RFLP.
For TNF -238 polymorphisms, the following primers were used to amplify a 230 bp product: 5'CCT CAA GGA CTC CAA AGC TTT CTG-3'; 5'ACA CTC CCC ATC CTC CCA GATC-3. For TNF-308 polymorphisms, the following primers were used to amplify a 107 bp product: 5' AGC AAT AGG TGG TTT TGA CTCGGGC CCAT-3'; 5'TCC TCC CTG CTC CGA TTC CG-3'.
For -857 polymorphisms, the following primers were used to amplify a 127 bp product: 5' GGC TCT GAG GAA TGG GTT AC-3'; 5'CCT CTA CAT GGC CCT GTC TAC-3'. For -863 the following primers were used to amplify a 126 bp product: A: 5′- GGC TCT GAG GAA TGG GTT AC-3′; B: 5′-CTA CAT GGC CCT GTC TTC GTT ACG-3′.
The amplification was accomplished by an initial denaturation at 940°C for 5 min, and 30 cycles at 940°C for 40 s, at 560°C for 40 s, at 720°C for 1 min, followed by an extension at 720°C for 6 min [7,9,22,24,25].
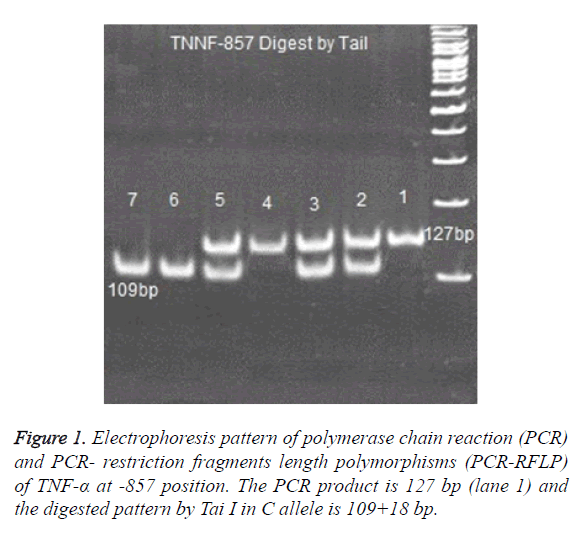
PCR products of, TNF -238, TNF -308 ,TNF -857 and TNF -863 digested with 2 U enzymes of BgI II, Bsaj I, Nco I, Tai I and Tai I, respectively [25,26]. Digested products were run on 8% polyacrylamide gel and were stained with Silver-Nitrate (Figure 1).
TNFR1 genotyping
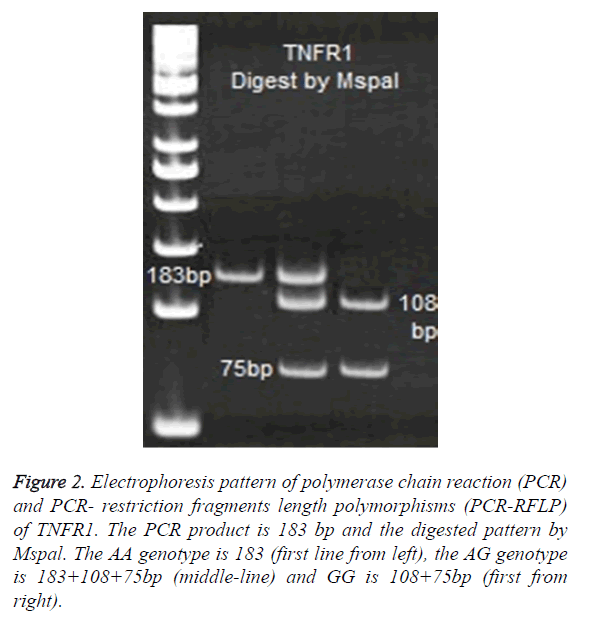
For TNFR1 polymorphisms at 36 positions, the following primers were used to amplify a 183 bp product: A: 5′- GAC CCC AAA TGG GGG AGT GAG AGG-3′ and- B: 5′-ACC AGG CCC GGG CAG GAG AG-3′. PCR products of TNFR1 digested with 2 U enzymes of Mspal. Digested products were run on 8% polyacrylamide gel and were stained with Silver- Nitrate (Figure 2).
Figure 2. Electrophoresis pattern of polymerase chain reaction (PCR) and PCR- restriction fragments length polymorphisms (PCR-RFLP) of TNFR1. The PCR product is 183 bp and the digested pattern by Mspal. The AA genotype is 183 (first line from left), the AG genotype is 183+108+75bp (middle-line) and GG is 108+75bp (first from right).
TNFR2 genotyping
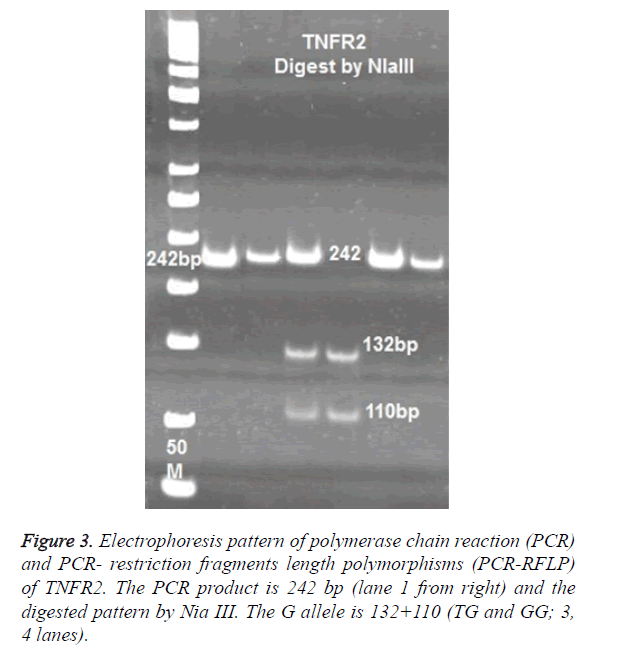
For TNFR2 polymorphisms at 587 positions, the following primers were used to amplify a 242 bp product: 5’-TTCTGGAGTTGGCTGCGTGT-3’ and- ACTCTCCTATCCTGCCTGCT-3’. PCR products of TNFR2 digested with 2 U enzymes of NLA III. Digested products were run on 8% polyacrylamide gel and were stained with Silver- Nitrate (Figure 3).
Statistical analysis
Statistical analysis was performed using chi-square test to determine statistical associations between cases and control. P-value less than 0.05 were considered statistically significant. Data were analyzed using SPSS version 18 Software.
Ethics
All the patients with positive TB smear and culture were explained for the aim and process of the research project before giving their informed consent and the private and personal information were secured by the investigators. There was no limitation to quit the study and patients and controls paid no cost for genetic tests.
Results
Participants and demographics
Totally 151 cases and 83 controls enrolled in this study. The mean age of patient and control was 49.2 ± 21.2 and 42.5 ± 20.16 y, respectively. Majority of patients (87, 57.6%) are over 50 y old, and 12 are more than 65 y old (12, 7.9%). Seventy eight (51.6%) TB cases were male and seventy three (48.3%) were female. 46 patients (30.4%) had previous history of TB and the remaining were new smear positive TB cases (69.5%). Fisher’s exact test showed a correlation between old age and susceptibility to pulmonary TB (P value<0.001).Concerning past medical history, a P value<0.001 was enough to blame previous pulmonary TB involvement as a main factor for current disease among our patients.
TNF-α SNPs at -238, -308, -857 and -863 position
In overall, four types of polymorphisms were observed in TNF-α gene: an A to G substitution at position -238, a G to A substitution at position -308, a C to T substitution at position -857 and C to A substitution at position -863. Among these polymorphisms, C allele of TNF 857 and A allele of TNF 238 were more frequent in TB cases as compared to control group (Table 1). The SNPs at -308, -863 were not statistically significant in studied cases. But the genotypes of TNF at 857 C/C (85, 56.2%) and TNF 238 A/A 127 (84.1%) genotypes were associated with increased risk of acquiring TB.
| Genotypes | Control (83) | TB cases (151) | OR (CI 95%) | P |
|---|---|---|---|---|
| TNF863 Allele C A Genotype CC CA AA |
122 (73.4%) 25 (15.0%) 61 (73.4%) 19 (22.8%) 3 (3.6%) |
255 (84.4%) 47 (15.5%) 106 (70.1%) 43 (28.4%) 2 (1.3%) |
0.6 (0.3-1.2)
0.6 (0.3-1.2) 1.3 (0.5-1.8) 1.5 (1.4-1.7) |
NS*
NS NS NS |
| TNF857 Allele T C Genotype TT TC CC |
56 (33.7) 110 (66.3) 8 (9.6) 40 (48.1) 35 (42.1) |
72 (23.8) 230 (76.21) 6 (3.9) 60 (39.7) 85 (56.21) |
0.6 (0.4-0.9)
0.3 (0.4-1.1) 0.7 (0.4-1.2) 0.5 (0.3-0.9) |
0.02
0.08 NS 0.03 |
| TNF308 Allele G A Genotype GG GA AA |
154 (92.7) 12 (7.3) 71 (85.5) 12 (14.5) 0 (0.0) |
272 (90.0) 30 (10.0) 122 (80.7) 28 (18.5) 1 (0.6) |
0.7 (0.3-1.4)
0.7 (0.3-1.4) 1.3 (0.6-2.8) 1.5 (1.4-1.7) |
NS
NS NS NS |
| TNF238 Allele A G Genotype AA AG GG |
100 (60.2) 66 (39.8) 49 (59.0) 2 (2.4) 32 (38.5) |
272 (89.4) 32 (10.6) 127 (84.1) 16 (10.5) 8 (5.2) |
5.5 (3.4-9.0)
3.6 (1.9-6.8) 4.8 (1.0-2.4) 0.1 (0.03-0.2) |
0
0 0.02 0 |
| TNFR1 (36) Allele A G Genotype AA AG GG |
128 (77.0%) 38 (22.81%) 48 (57.8%) 32 (38.5%) 3 (3.6%) |
230 (76.1%) 72 (23.8%) 79 (52.0%) 72 (47.6%) 0 (0.0) |
0.3 (0.2- 1.4)
1.4 (0.6-2.8) 0.9 (0.3-1.4) 1.5 (1.1-2.8) |
NS
NS NS NS |
| TNFR2 (587) Allele T G Genotype TT TG GG |
77 (46.3%) 89 (53.6%) 8 (9.6%) 61 (73.4%) 14 (16.8%) |
144 (47.6%) 158 (52.3%) 10 (6.6%) 124 (82.1%) 17 (11.2%) |
0.5 (0.2- 1.4)
1.3 (0.6-2.8) 0.7 (0.3-1.4) 1.3 (0.6-2.8) |
NS
NS NS NS |
| NS*: Non-Significant. | ||||
Table 1. The allele and genotyping frequencies of TNF-α gene, TNFR and TNFR1.
TNFR1 SNPs
No significance differences were observed in allele A and G between patients and control cases (Table 1). The frequency of AA genotypes was 57% (48/83) in control and 52% (79/151) in TB patients. Therefore, the most frequent allele in TNFR1 at 36 position was A allele, which observed in 77% of controls and 76% of patients (Figure 2).
TNFR2 SNPs
No significance difference was observed in allele T and G between patients and control cases. The frequency of TT genotypes was 9.6% (8/83) in control and 6.6% (10/151) in TB patients. The most frequent genotypes in TNFR2 at 587 position was TG, which observed in 73.4% of controls and 82.1% of patients (Figure 3).
Correlation of TNF-α and TNFR2 SNPs with TB susceptibility
Combined correlation analysis was then conducted in terms of coincident SNPs in TNF-α (-238, -308, -857and -863), TNFR1 (36) and TNFR2 (587) loci. No significant correlation was found.
Discussion
The current study found a strong correlation between two polymorphisms in different loci of TNF-α gene including 857 C/C (85, 56.2%) and TNF 238 A/A 127 (84.1%). Although, no association between polymorphisms of TNF-α gene and its receptor i.e., TNFR2 (Table 1) was detected. For decades, several reports illustrated different association between genetic polymorphisms and susceptibility to a wide range of infectious diseases such as tuberculosis. Recently Yi et al. performed a meta-analysis on TNF-α polymorphism and its association with pulmonary TB susceptibility [23]. This study evaluated and analyzed 18 previously studies which compromised a total of 2735 cases and 3177 controls. Among various positions in promoter region of TNF-α gene they could outlines the protective roles of TNF-α SNPs at position -857 C>T, -308 G>A, 238 G>A in different ethnical populations e.g., TNF-α SNPs at -308 G>A position is more in Africans origin than Asian [20]. In contrast, the -238 G>A was associated with PTB among Asians. In this regard, Varahram et al. showed the importance of -308 G>A polymorphisms in Iranian PTB patients [26]. But in present study, the frequency of G to A study was 92.7%, 90% and 7.3%, 10.0% in control and patients, respectively. As the number of studied cases in present study was higher, we thereby conclude TNF-α SNPs at -308 G>A position may not be associated with Iranian PTB. More recently, Jafari et al., showed an association between 14 gene polymorphisms and PTB [21]. They found that TLR4 (D299G and T399I) as well as TNF-α (-308 G>A) may influence the risk of PTB [28]. In this regards, Joshi et al., studied synergistic or prognostic role for cytokines like TNF-α t in three groups of individual including pulmonary TB patients, their household contacting people and healthy individuals [29]. They found significantly higher TNF-α serum levels among patients and their caregivers than healthy people. The significant ratio of TNF-α /IL-10 determined Th1 predominance in PTB patients and caregivers as well. Lee et al. released a research report on the role of TNF-α polymorphisms in PTB susceptibility among European, Asian and Middle Eastern population [24]. They introduced TNF-α -857 T as a protective factor for PTB in Asians (OR=0.682, 95% CI= 0.550-0.846) but no relationship was reported for TNF-α -308 A/G and -238 A/G. Our result showed a higher frequency of C allele in TNF-α -857 (76.21%) in patients than control group (66.3%), which is consistent with some early reports [22,26]. During recent years, it become clear that TNF-α exerts its pleiotropic function by activating signaling cascades via binding to two types of receptors i.e., TNFR1 and TNFR2 [3,16,18]. Generally, TNFR1 is associated with inflammation and tissue degeneration [30,31].
TNFR2, encoded by the tumor necrosis factor receptor superfamily member IB gene (TNFRSFIB), can influence the biological activity of TNF-α both in a membrane- bound and a soluble form. In terms of TNFR2 polymorphisms and the risk of PTB, Moller et al., studied a South African population to find a protective impact of T allele of rs3397 alone and/or the 3’untranslated region haplotype GTT of TNF receptor 2 and concluded a role for TNF and TNF receptor mediated immune responses in the pathogenesis of the named disease [32]. In overall, GTT haplotype was found in an intermediate frequency (26%) among different studied population. We found no association between SNPs in TNFR1 and TNFR2 with susceptibility to tuberculosis. Thereby, further studies are needed to elucidate the functionally importance of both TNFR1 and TNFR2 SNPs.
Despite of the big dilemma and controversy for specific human genome diversity to shift individuals toward getting diseases, hereon, pulmonary TB, there are obvious evidence for their impact. Therefore, multidisciplinary studies with multiracial samples are highly advised to confirm the associations which have been raised during recent decades about the risk of involvement, prognosis and response to treatment regarding different demographic and genetic aspects. Concerning our current study, screening assessments for TNF-α -857 and -308 SNPs in Iran would be important in order to make future decisions for preventive treatments before getting the disease among people who are at high risk considering their genotyping.
Conflict of Interest
The author has no conflict of interest.
References
- World Health Organization. Tuberculosis fact sheet no: 104. WHO 2010.
- World Health Organization. Epidemiology, Global tuberculosis control: epidemiology, strategy, financing. WHO 2009; 6-33.
- World Health Organization. The global report on tuberculosis (16th ed). WHO 2011.
- Velayati AA, Bakayev V, Bahadori M. Religious and cultural traits in HIV/AIDS epidemics in sub-Saharan Africa. Archiv Iranian Med 2007; 10: 486-497.
- Gebreegziabiher D, Adane K, Abebe M. A survey on undiagnosed active pulmonary tuberculosis among pregnant mothers in mekelle and surrounding Districts in Tigray, Ethiopia. Int J Mycobacteriol 2017; 6: 43-46.
- Lawn SD, Zumla AI. Tuberculosis. Lancet 2011; 378: 57-72.
- Qi Z, Yang W, Wang YF. Epidemiological analysis of pulmonary tuberculosis in Heilongjiang province China from 2008 to 2015. Int J Mycobacteriol 2017; 6: 264-267
- World Health Organization. The global tuberculosis report (19th ed). WHO 2014.
- Bhatt K, Salgame P. Host innate immune response to Mycobacterium tuberculosis. J Clin Immunol 2007; 27: 347-362.
- Amila A, Acosta A, Sarmiento ME , Suraiya S, Zafarina Z, Panneerchelvam S, Norazmi MN. Sequence comparison of six human microRNAs genes between tuberculosis patients and healthy individuals. Int J Mycobacteriol 2015; 4: 341-346.
- Furci L, Schena E, Miotto P, Cirillo DM. Alteration of human macrophages microRNA expression profile upon infection with Mycobacterium tuberculosis. Int J Mycobacteriol 2013; 2: 128-134.
- Bose M, Farnia P, Sharma S, Chattopadhya D, Saha K. Nitric oxide dependent killing of mycobacterium tuberculosis by human mononuclear phagocytes from patients with active tuberculosis. Int J Immunopathol Pharmacol 1999; 2: 69-79.
- Bakayev VV, Mohammadi F, Bahadori M. Arylamine N-acetyltransferase 2 slow acetylator polymorphisms in unrelated Iranian individuals. Euro J Clin Pharmacol 2004; 60: 467-471.
- Wallis RS. Tumour necrosis factor antagonists: structure, function, and tuberculosis risks. Lancet Infect Dis 2008; 8: 601-611.
- Khalilullah SA, Harapan H, Hasan NA, Winardi W, Ichsan I, Mulyadi M. Host genome polymorphisms and tuberculosis infection: What we have to say? Egypt J Chest Dis Tuberc 2014; 63: 173-185.
- Naudé PJ, den Boer JA, Luiten PG, Eisel UL. Tumor necrosis factor receptor cross-talk. FEBS J 2011; 278: 888-898.
- Faustman D, Davis M .TNF receptor 2 pathway: drug target for autoimmune diseases. Nat Rev Drug Discov 2010; 9: 482-493.
- Faustman D, Davis M. TNF receptor 2 and disease: autoimmunity and regenerative medicine. Front Immunol 2013; 4: 478.
- Al-Ansari AS, Ollier RWE, Villarreal J. Tumor necrosis factor receptor II (TNFRII) exon 6 polymorphism in systemic lupus erythematosus. Tissue Antigens 2000; 55: 97-99.
- Barton A, John SR, Ollier WE, Silman A, Worthington J. Association between rheumatoid arthritis and polymorphism of Tumor Necrosis Factor Receptor II but not Tumor Necrosis Factor Receptor I, in Caucasians. Arthritis Rheumatism 2001; 44: 61-65.
- Komata T, Tsuchiya N, Matsushita M, Hagiwar K, Tokunaga K. Association of tumor necrosis factor receptor 2 (TNFR2) polymorphism with susceptibility to systemic lupus erythematosus. Tissue Antigens 1999; 53: 527-533.
- Anoosheh S, Farnia P, Kargar M. Association between TNF-Alpha (-857) gene polymorphism and susceptibility to tuberculosis. Iranian Red Crescent Med J 2011; 13: 243-248.
- Yi YX, Han JB, Zhao L, Fang Y, Zhang YF, Zhou GY. Tumor necrosis factor alpha gene polymorphism contributes to pulmonary tuberculosis susceptibility: evidence from a meta-analysis. Int J Clin Exp Med 2015; 8: 20690-20700.
- Lee YH, Song GG. Associations between tumor necrosis factor-α polymorphisms and susceptibility to pulmonary tuberculosis: meta-analysis. Genet Mol Res 2015; 14: 8602-8612.
- Mabunda N, Alvarado-Arnez LE, Vubil A. Gene polymorphisms in patients with pulmonary tuberculosis from Mozambique. Mol Biol Rep 2015; 42: 71-76.
- Varahram M, Farnia P, Nasiri MJ. Association of Mycobacterium tuberculosis lineages with IFN-γ and TNF-α gene polymorphisms among pulmonary tuberculosis patient. Mediterr J Hematol Infect Dis 2014; 6: e2014015.
- Joseph S, David WR. Molecular cloning: A laboratory manual. CSHL Press 2001.
- Jafari M, Nasiri MR, Sanaei R. The NRAMP1, VDR, TNF-α, ICAM1, TLR2 and TLR4 gene polymorphisms in Iranian patients with pulmonary tuberculosis: A case-control study. Infect Genet Evol 2016; 39: 92-98.
- Joshi L, Ponnana M, Sivangala R. Evaluation of TNF-α, IL-10 and IL-6 cytokine production and their correlation with genotype variants amongst tuberculosis patients and their household contacts. PLoS One 2015; 10: e0137727.
- Sivangala R, Ponnana M, Thada S. Association of cytokine gene polymorphisms in patients with tuberculosis and their household contacts. Scand J Immunol 2014; 79: 197-205.
- Fischer R, Kontermann RE, Maier O. Targeting sTNF/TNFR1 signaling as a new therapeutic strategy. Antibodies 2015; 4: 48-70.
- Möller M, Flachsbart F, Till A. A functional haplotype in the 3'untranslated region of TNFRSF1B is associated with tuberculosis in two African populations. Am J Respir Crit Care Med 2010; 181: 388-393.