Research Article - Biomedical Research (2017) Volume 28, Issue 11
A validation study for the use VE1 immunohistochemical staining in screening for BRAF mutation in cutaneous malignant melanoma
Weiming Zhang1, Guoxin Song1, Xue Han1, Xiao Li1 and Dan Luo2*
1Department of Pathology, the First Affiliated Hospital of Nanjing Medical University, Nanjing, Jiangsu, PR China
2Department of Dermatology, the First Affiliated Hospital of Nanjing Medical University, Nanjing, Jiangsu, PR China
- *Corresponding Author:
- Dan Luo
Department of Dermatology
The First Affiliated Hospital of Nanjing Medical University, PR China
Accepted on March 27, 2017
Abstract
BRAF V600E mutation analysis has been an important companion diagnostic assay to evaluate prognosis of malignant melanoma and to guide target therapy with BRAF inhibitor. The aim of this study was to assess the sensitivity and specificity of Immunohistochemistry (IHC) using VE1 mutation specific antibody on screening for BRAF V600E mutation patients with cutaneous malignant melanoma. We detected BRAF V600E mutation on a total of 90 formalin-fixed paraffin-embedded melanoma tissue samples by ARMS-PCR and immunohistochemistry simultaneously. The results obtained from ARMS were set as a diagnostic gold standard; IHC staining with BRAF V600E mutation-specific antibody displayed 100% sensitivity and 96.8% specificity. Further statistical analysis showed that results from the two methods were consistent (kappa=0.948 p<0.001), Positive Predictive Value (PPV) and Negative Predictive Value (NPV) were 93.1%and 100%. Therefore, as a simple, fast and low cost method, IHC can be used as a routine screening method for BRAF V600E mutation analysis in cutaneous malignant melanoma.
Keywords
Braf, Mutation, Melanoma immunohistochemistry, ARMS
Introduction
Cutaneous Malignant Melanoma (CMM) is a type of highly aggressive skin tumor originated from uncontrolled proliferation of melanocytes. It accounts for 80% of deaths arising from skin cancer and its incidence rate is on the rise over the past decades [1,2]. V-Raf murine sarcoma viral oncogene homolog B (BRAF) belongs to the Raf family of serine/threonine-specific protein kinases, which activate the RAF-MEK-ERK (MAPK) pathway and play crucial role in the process of cell growth, differentiation, and survival [3,4]. BRAF somatic mutations are the most prevalent mutation in human malignant melanoma, it was reported that BRAF gene was mutated in nearly 40-60% melanoma patients. BRAF V600E mutation, which involved in an exon 15 missense mutation (T1796A), that substitute glutamic acid (E) for Valine (V) at amino acid 600 (BRAF V600E), account for 70% to 90% of all BRAF mutations in melanoma, followed by mutations V600K (10%-30%) and V600D/R (3%) [5,6]. Recently, the role of BRAF (V600E) mutation in predicting clinical outcome has been extensively verification. Melanoma patients with BRAF V600E mutation have shown a significantly worse overall survival than those patients without BRAF V600E mutation [7,8]. Therefore, the targeted therapy against on BRAF has attracted more attentions recently [9]. Since BRAF inhibitors are only effective in patients with BARF mutation, BRAF mutation analysis has been important companion diagnostic test to guide the treatment of melanoma, it is necessary for doctors to find out the mutation status of BRAF gene prior to treatment [10,11]. Currently, methods for BRAF V600E mutation detection are mainly DNA-based techniques including Sanger sequencing, real-time PCR and Amplification Refractory Mutation System (ARMS)-PCR [12-14]. Among these methods, ARMS-PCR is the most widely used method due to its high sensitivity and specificity. However, this method also has its limitation. it involve in specialized equipment and laboratory techniques, the whole detection process require tight quality control program, long turnaround time and high detection costs, while most hospitals cannot meet this demand and have to send samples to other eligible laboratories to complete this test [15,16]. Furthermore, the final detection results are easily affected by insufficient tissue fixation or low tumor cell content [17,18]. As the purified BRAF V600E-specific antibody (clone: VE1) was commercially available in 2011, several published researches have reported that IHC was a sensitive and specific screening methods for BRAF V600E mutation [19,20]. Compared to the ARMS-PCR, immunohistochemistry is a simple, fast and practicable assay in common diagnostic pathology laboratory [21,22]. In this study, we detected BRAF V600E mutation using immunohistochemistry and ARMS-PCR in a retrospective diagnostic setting. The data was collected to analyse the consistency of the two methods and to evaluate the role of immunohistochemistry in screening BRAF V600E mutation.
Materials and Methods
Tissue specimen preparations
A total of 90 cases of cutaneous malignant melanoma archival Formalin-Fixed Paraffin Embedded (FFPE) tissue blocks were from the first affiliated hospital of Nanjing medical university, China, between 2012 and 2015. All tissues were fixed in 10% neutral buffer formalin after resection and processed by standard tissue dehydration program. Two pathologists who are blind to this research reviewed all tissue sections respectively and marked the tumor tissue based on H and E staining. Each block was sectioned at 4 μm for immunohistochemical staining and 6 μm for DNA extraction. Patients’ name and other involved privacy were hidden during the whole study process. The ethics committee of the hospital approved our research design, including the use of all tissue blocks.
Immunohistochemistry detection
Immunohistochemical stain was performed according to the routine operation procedure in department of pathology. Mouse anti-human BRAF V600E Primary antibody (clone VE1, ready to use) was purchased from ZSGB-BIO Company, Beijing. Detail stain protocol is briefly as below: 4 μm thick paraffin sections from each block were firstly deparaffinized in xylene and dehydrated, then immersed in pH 9.0 boiling EDTA buffer in a pressure cooker for 10min to retrieve the antigen. After inactivation endogenous peroxidase activity with 3% H2O2, slides were incubated with primary antibody overnight at 4°C in a humidified chamber. Slides were subsequently incubated with HRP conjugated universal secondary antibody (K5007, purchased from Dako Company) for 15 min at room temperature, then DAB/AEC staining for 5-8 min and counterstaining with Hematoxylin. Between each incubation step, slides were washed with pH 7.2-7.4 0.1 mol/L PBST buffer at least 3 times. IHC analysis was performed based on the percent of positively stained cells and the staining intensity. Briefly, the percentage of positive staining was scored as negative (no staining), 1+ (0%-10% cell shows positive staining), 2+ (10%-30%) and 3+ (over 30%).
ARMS PCR analysis and direct sequencing
Pathologists firstly reviewed the H and E slides and circled the interested tumor area to guide macro-dissection of tumor tissues, 5-10 μm thick sections were scraped in a polypropylene micro-centrifuge tube, QIAamp DNA FFPE Tissue Kit (56404) was employed to isolate genomic DNA. BRAF V600E mutation analysis was performed by using AmoyDx™ BRAF V600E Mutation Detection kit (Amoy Diagnostics, Xiamen, China) according to the manufacturer's instructions. When IHC and ARMS results were discrepant, direct sequencing method was performed to verify the real status of BRAF V600E mutation in undetermined cases. Designed sequence of the forward primer was 5- GCTTGCTCTGATAGGAAAATGAG-3’ and the reverse primer was 5-GTAACTCAGCAGCATCTCAGG-3. PCR amplification conditions: 94°C for 10 min, followed by 40 cycles of 94°C for 45 s, 61°C for 45 s, and 72°C for 45 s, with a final extension of 7 min at 72°C. Purified PCR products were then run on an ABI3130 Genetic Analyzer and analysed using software supplied by the manufacturer.
Statistical analyses
Statistical analysis was performed using SPSS (SPSS19.0 Inc., Chicago, Illinois, USA), Kappa test was used to compare the consistency of IHC and ARMS PCR. Kappa value ≥ 0.75, two methods have better consistency, while Kappa value<0.4 indicates poor consistency of two methods. Sensitivity,specificity,Positive Predictive Value (PPV) and Negative Predictive Value (NPV) of IHC were evaluated by using diagnostic test.
Sensitivity of IHC is the proportion of true positives identified as positive (True positives/True positives+False negatives); Specificity is calculated as the proportion of true negatives (True negatives/True negatives+False positives); PPV is the proportion of true positives among test results (True positives/ True positives+False positives); NPV is the proportion of true negatives among test results (True negatives/True negatives +False negatives).
Results
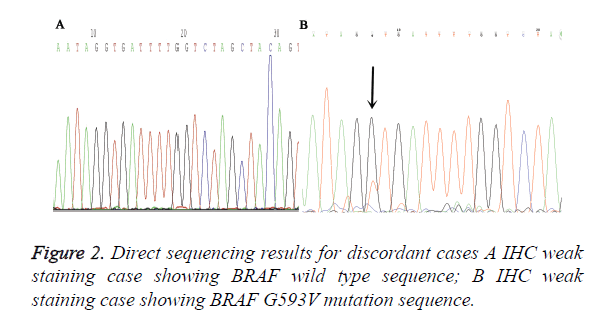
A total of 90 melanoma tissues were tested for BRAF V600E mutation status by ARMS-PCR and IHC. Both methods were performed successfully in all cases and the results from two methods were listed in Table 1. VE1 immunohistochemical positive staining was homogenously located in the cytoplasm of tumor cells. Representative photos are shown in Figure 1. Of the 90 cases, 27 (30%) cases were found to be BRAF V600E mutation by ARMS-PCR, and of 27 cases, 16 cases were strongly positive (3+) staining, 11 cases were moderately positive (2+) staining. Among 63 BRAF wild type cases determined by ARMS-PCR, 2 cases showed IHC weak positive staining. Discordant cases were then re-tested by further direct sequencing method, one case was BARF wild type and the other case was BRAF G593V mutation (Figure 2). To validate the accuracy of VE1 mutant specific antibody, we then compared the results from two methods. ARMS-PCR was set as a gold standard method, the sensitivity and specificity of IHC was 100% and 96.8% respectively, PPV and NPV were 93.1%and 100%. Two methods in our investigation have good consistency with Kappa=0.948 (p<0.01).
| IHC | ARMS | Total | |
|---|---|---|---|
| Mutation | Wild type | ||
| + | 27 | 2 | 29 |
| - | 0 | 61 | 61 |
| Total | 27 | 63 | 90 |
Table 1. Comparison of BRAF V600E mutation by ARMS-PCR and VE1 staining by IHC.
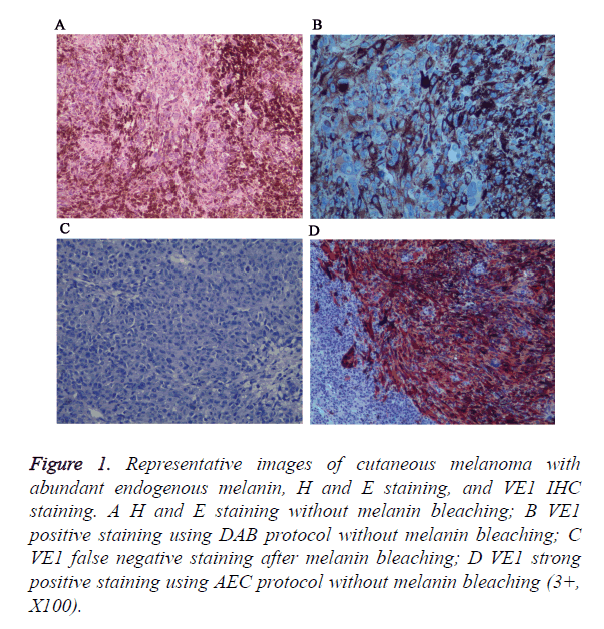
Figure 1. Representative images of cutaneous melanoma with abundant endogenous melanin, H and E staining, and VE1 IHC staining. A H and E staining without melanin bleaching; B VE1 positive staining using DAB protocol without melanin bleaching; C VE1 false negative staining after melanin bleaching; D VE1 strong positive staining using AEC protocol without melanin bleaching (3+, X100).
In addition, insoluble melanins will interfere normal interpretation of positive results, so we try to bleach melanins with oxidizing agents before immunostain. However, we found that strong oxidizing agents will destroy antigen and lead to false negative staining. For that, we use AEC chromogen instead of DAB to avoid interference of endogenous melanin.
Discussion
BRAF (v-raf murine sarcoma viral oncogene homolog B1) is a RAS-regulated serine-threonine kinase and activator of the MAPK signaling cascade [23], which plays a vital role in regulating cellular responses to extracellular signals. In melanoma, it is reported that 40%-60% cases harbored BRAF mutation. Among those mutation cases, over 90% mutation type was a substitution of Valine (V) by glutamate (E) at condon600, referred to as BRAF V600E. Since the importance of BRAF V600E mutations in melanoma was reported in 2002 and most importantly, BRAF V600E-specific inhibitor Vemurafenib (PLX4032) was approved to treat BRAF V600E mutated metastasis melanoma [24], BRAF mutation then thought as a promising diagnostic and prognostic marker, increasing number of research focused on seeking effective, reliable, fast methods to detect BRAF mutation.
In this study, we detected BRAF V600E mutation in formalin-fixed paraffin-embedded cutaneous melanoma tissues by immunohistochemistry with mutation-specific antibody VE1 and compared IHC results with ARMS-PCR. Of 90 cases, ARMS-PCR displayed 27 (27/90) mutation cases, our IHC showed 100% sensitivity and 96.8% specificity. Besides, two methods have better consistency with Kappa=0.948), PPV and NPV were 93.1%and 100% respectively. Our results were basically consistent with previous studies [25,26]. Zhou et al. [27] reported the high concordance between IHC with VE1 and Sanger sequencing/ARMS was 93.2% in primary and metastatic papillary thyroid carcinoma, while in melanoma, Hui et al. [28] reported a 93.8% cases were coherent for VE1 staining. Compared with the reported data, our results showed a more high specificity and sensitivity. This may be partly due to the standard tissue dehydration program, which have a great effect on tissues antigen conservation. In addition, using stringent immunohistochemistry scoring criteria for IHC results interpretation also made an improvement in the sensitivity and specificity of IHC.
In our study, total 2 IHC staining cases were proved as false positive. 1 case showed weak staining in tumor cells. Meanwhile, non-specific background staining was also presented in the stroma of tumors. We speculate that it is such nonspecific background staining interfere cause the misinterpretation made by Pathologists. Currently, IHC has been a reliable routine diagnostic method for pathology in clinics. However, when used as a method for guiding targeted therapy, it still face some knotty problems, For example, the lack of standard interpretation methods and different antigen retrieval procedure of IHC, which are the main reason for the difference between each pathology laboratory. But these drawbacks can be overcome by setting up stringent scoring criteria for result interpretation and standard operation procedure during IHC detection, especially when fully automated IHC detection platform was introduced. The other false positive case was corroborated as BRAF G593V mutation by further direct sequencing method. Although VE1 is a reported specific antibody against on V600E, whether it having cross reaction with other BRAF mutation type, such as BRAF V600K, V600R, still remains unclear. Heinzerling et al. [29] and Ihle et al. [30] found one melanoma case that showed moderate intensity by IHC and cross-reactivity with the V600R mutation and the p.V600K mutation respectively. While in papillary thyroid carcinoma, Zhou et al. considered that cross-reactivity for BRAF mutation may occur.
In conclusion, IHC with VE1 antibody was a sensitive and reliable method for the detection of BRAF V600E mutation in cutaneous malignant melanoma. It can be used as a preliminary screening method. Normally, we still recommend ARMS PCR as the conventional method for BRAF V600E detection in clinical practice. However, under certain circumstances, such as low tumor cell content or poor processed samples, IHC can be a compromised method for BRAF V600E mutation analysis. Besides, IHC screening can effectively reduce the patient's economic burden, for those VE1 negative staining cases, it is no use further to perform expensive molecular detection.
Conflict of Interest
The authors declare no conflicts of interest.
References
- Pakneshan S, Salajegheh A, Smith RA, Lam AK. Clinicopathological relevance of BRAF mutations in human cancer. Pathology 2013; 45: 346-356.
- Jain T, Bryce A. Intermittent BRAF Inhibition Can Achieve Prolonged Disease Control in BRAF Mutant Melanoma. Cureus 2015; 7: 410.
- Vultur A, Villanueva J, Herlyn M. Targeting BRAF in advanced melanoma: a first step toward manageable disease. Clin Cancer Res 2011; 17: 1658-1663.
- Tetzlaff MT, Pattanaprichakul P, Wargo J. Utility of BRAF V600E immunohistochemistry expression pattern as a surrogate of braf mutation status in 154 patients with advanced melanoma. Hum Pathol 2015; 46: 1101-1110.
- Jung YY, Yoo JH, Park ES. Clinicopathologic correlations of the BRAF V600E mutation, BRAF V600E immunohistochemistry,and BRAF RNA in situ hybridization in papillary thyroid carcinoma. Pathol Res Pract 2015; 211: 162-170.
- Boursault L, Haddad V, Vergier B. Tumor homogeneity between primary and metastatic sites for BRAF status in metastatic melanoma determined by immunohistochemical and molecular testing. PLoS One 2013; 8: 70826.
- Saroufim M, Habib RH, Gerges R. Comparing BRAF mutation status in matched primary and metastatic cutaneous melanomas: Implications on optimized targeted therapy. Exp Mol Pathol 2014; 97: 315-320.
- Qiu T, Lu H, Guo L. Detection of BRAF mutation in Chinese tumor patients using a highly sensitive antibody immunohistochemistry assay. Sci Rep 2015; 5: 921-925.
- Dvorak K, Aggeler B, Palting J. Immunohistochemistry with the anti-BRAF V600E (VE1) antibody: impact of pre-analytical conditions and concordance with DNA sequencing in colorectal and papillary thyroid carcinoma. Pathology 2014; 46: 509-517.
- Long GV, Wilmott JS, Capper D, Preusser M, Zhang YE. Immunohistochemistry is highly sensitive and specific for the detection of V600E BRAF mutation in melanoma. Am J Surg Pathol 2013; 37: 61-65.
- Scholtens A, Geukes Foppen MH, Blank CU. Vemurafenib for BRAF V600 mutated advanced melanoma: Results of treatment beyond progression. Eur J Cancer 2015; 51: 642-652.
- Estrella JS, Tetzlaff MT, Bassett RL, Patel KP, Williams MD. Assessment of BRAF V600E status in colorectal carcinoma: tissue-specific discordances between immunohistochemistry and sequencing. Mol Cancer Ther 2015; 14: 2887-2895.
- Zhu X, Luo Y, Bai Q, Lu Y. Specific immunohistochemical detection of the BRAF V600E mutation in primary and metastatic papillary thyroid carcinoma. Exp Mol Pathol 2016; 100: 236-241.
- Chen Q, Xia C, Deng Y, Wang M, Luo P. Immunohistochemistry as a quick screening method for clinical detection of BRAF(V600E) mutation in melanoma patients. Tumour Biol 2014; 35: 5727-5733.
- Capper D, Preusser M, Habel A, Sahm F, Ackermann U. Assessment of BRAF V600E mutation status by immunohistochemistry with a mutation-specific monoclonal antibody. Acta Neuropathol 2011; 122: 11-19.
- Ritterhouse LL, Barletta JA. BRAF V600E mutation-specific antibody: A review. Semin Diagn Pathol 2015; 32: 400-408.
- Colomba E, Helias-Rodzewicz Z, Von Deimling A, Marin C, Terrones N. Detection of BRAF p.V600E mutations in melanomas: comparison of four methods argues for sequential use of immunohistochemistry and pyrosequencing. J Mol Diagn 2013; 15: 94-100.
- Routhier CA, Mochel MC, Lynch K, Dias-Santagata D, Louis DN. Comparison of 2 monoclonal antibodies for immunohistochemical detection of BRAF V600E mutation in malignant melanoma, pulmonary carcinoma, gastrointestinal carcinoma, thyroid carcinoma, and gliomas. Hum Pathol 2013; 44: 2563-2570.
- Sasaki H, Shimizu S, Tani Y, Shitara M, Okuda K. Usefulness of immunohistochemistry for the detection of the BRAF V600E mutation in Japanese lung adenocarcinoma. Lung Cancer 2013; 82: 51-54.
- Busam KJ, Hedvat C, Pulitzer M. Immunohistochemical analysis of BRAF(V600E) expression of primary and metastatic melanoma and comparison with mutation status and melanocyte differentiation antigens of metastatic lesions. Am J Surg Pathol 2013; 37: 413-420.
- Feller JK, Yang S, Mahalingam M. Immunohistochemistry with a mutation-specific monoclonal antibody as a screening tool for the BRAF V600E mutational status in primary cutaneous malignant melanoma. Mod Pathol 2013; 26: 414-420.
- Skorokhod A, Capper D, von Deimling A, Enk A, Helmbold P. Detection of BRAF V600E mutations in skin metastases of malignant melanoma by monoclonal antibody VE1. J Am Acad Dermatol 2012; 67: 488-491.
- Wan PT, Garnett MJ, Roe SM, Lee S, Niculescu-Duvaz D. Mechanism of activation of the RAF-ERK signaling pathway by oncogenic mutations of B-RAF. Cell 2004; 116: 855-867.
- Flaherty KT, Puzanov I, Kim KB, Ribas A, McArthur GA. Inhibition of mutated, activated BRAF in metastatic melanoma. N Engl J Med 2010; 363: 809-819.
- Uguen A, Gueguen P, Legoupil D. Dual NRASQ61R and BRAFV600E mutation-specific immunohistochemistry completes molecular screening in melanoma samples in a routine practice. Human Pathology 2015; 46: 1582-1591.
- Shih-Fan Kuan, Sarah Navina, Kristi L. Immunohistochemical detection of BRAF V600E mutant protein using the VE1 antibody in colorectal carcinoma is highly concordant with molecular testing but requires rigorous antibody optimization. Human Pathol 2014; 45: 464-472.
- Xiaoli Z, Yanli L, Xiaoyan Z. Specific immunohistochemical detection of the BRAF V600E mutation in primary and metastatic papillary thyroid carcinoma. Exp Mol Pathol 2016; 100: 236-241.
- Hui L, Zhongwu L, Weicheng X. Immunohistochemical detection of the BRAF V600E mutation in melanoma patients with monoclonal antibody VE1. Pathology Int 2014; 64: 601-606.
- Heinzerling L, Kuhnapfel S, Meckbach D, Baiter M, Kaempgen E. Rare BRAF mutations in melanoma patients: implications for molecular testing in clinical practice. Br J Cancer 2013; 108: 2164-2171.
- Ihle MA, Fassunke J, Konig K. Comparison of high resolution melting analysis, pyrosequencing, next generation sequencing and immunohistochemistry to conventional Sanger sequencing for the detection of p.V600E and non-p.V600E BRAF mutations. BMC Cancer 2014; 14.