- Journal of RNA and Genomics (2010) Research Report
Prediction of human targets for viral-encoded microRNAs by thermodynamics and empirical constraints
| Alessandro Laganà1,2,*, Stefano Forte3, Francesco Russo2, Rosalba Giugno1,§, Alfredo Pulvirenti1,§, Alfredo Ferro1,2 1Department of Mathematics and Computer Science, University of Catania, Italy, 2Department of Biomedical Sciences, University of Catania, Italy, 3IOM Ricerca, Viagrande, Italy |
| *Correspondence to: Alessandro Laganà, Email: lagana@dmi.unict.it, Tel: +39 095 7337208, Fax: +39 095 330094 |
| §These authors contributed equally to this work. |
| Received 26 February 2010; Revised 03 May 2010; Accepted 17 May 2010; Published online 24 May 2010 |
| J RNAi Gene Silencing, 2010, 6(1), 379-385 |
| © Copyright The Authors: This is an open access article, published under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/2.0/uk/). This license permits noncommercial use, distribution and reproduction of the article, provided the original work is appropriately acknowledged with correct citation details. |
Abstract
MicroRNAs (miRNAs) are small RNA molecules that modulate gene expression through degradation of specific mRNAs and/or repression of their translation. miRNAs are involved in both physiological and pathological processes, such as apoptosis and cancer. Their presence has been demonstrated in several organisms as well as in viruses. Virus encoded miRNAs can act as viral gene expression regulators, but they may also interfere with the expression of host genes. Viral miRNAs may control host cell proliferation by targeting cell-cycle and apoptosis regulators. Therefore, they could be involved in cancer pathogenesis. Computational prediction of miRNA/target pairs is a fundamental step in these studies. Here, we describe the use of miRiam, a novel program based on both thermodynamics features and empirical constraints, to predict viral miRNAs/human targets interactions. miRiam exploits target mRNA secondary structure accessibility and interaction rules, inferred from validated miRNA/mRNA pairs. A set of genes involved in apoptosis and cell-cycle regulation was identified as target for our studies. This choice was supported by the knowledge that DNA tumor viruses interfere with the above processes in humans. miRNAs were selected from two cancer-related viruses, Epstein-Barr Virus (EBV) and Kaposi-Sarcoma-Associated Herpes Virus (KSHV). Results show that several transcripts possess potential binding sites for these miRNAs. This work has produced a set of plausible hypotheses of involvement of v-miRNAs and human apoptosis genes in cancer development. Our results suggest that during viral infection, besides the protein-based host regulation mechanism, a post-transcriptional level interference may exist. miRiam is freely available for downloading at http://ferrolab.dmi.unict.it/miriam.
Keywords |
| miRNA, virus, cancer, apoptosis, cell cycle, EBV, KSHV |
Introduction |
| MicroRNAs (miRNAs) are small non-coding RNAs involved in post-transcriptional gene silencing (PTGS) that is a highly-conserved mechanism for regulating gene expression. miRNAs inhibit protein synthesis by binding with the 3’UTR or, rarely, the coding sequence of specific mRNAs (Bartel, 2004). |
| miRNAs are known to be involved in several physiological processes, such as apoptosis, development and cell differentiation, and their aberrant expression is observed in several diseases such as cancer, neurodegeneration and cardiac pathologies (Garofalo et al, 2008). |
| miRNAs are also expressed by viruses (v-miRNAs), which may act as self-regulators. It has been shown that the EBV-encoded miRNA, miR-BART2, down-regulates the viral DNA polymerase BALF5, thus inhibiting transition from latent to lytic viral replication (Barth et al, 2008). vmiRNAs can also interference with the host cell transcripts (Cullen, 2006). The cellular protein p53, an up-regulated modulator of apoptosis (PUMA), is regulated by the EBVencoded miR-BART5. The expression of miR-BART5 renders infected miR-BART5-expressing cells less sensitive to pro-apoptotic agents, but apoptosis can be triggered by depleting miR-BART5 or inducing the expression of PUMA. This shows the capability of vmiRNAs to facilitate the establishment of latent infection by promoting host cell survival (Choy et al, 2008). |
| The silencing of some nodes of the apoptotic pathway would give to the virus an important advantage during its lysogenic phase, since the host cells would not die by cell death triggered from infection and the virus could complete its vital cycle. Many v-miRNAs are located in proximity or inside viral latency genes and, presumably, co-transcribed with them (Nair and Zavolan, 2006). On the other hand, in many cases of viral infection the activation of apoptosis is needed during the lytic cycle of the virus allowing the processing of capsides and viral particles dissemination (Mendez et al, 2004). |
| The negative and positive modulation of the apoptotic process is a fundamental activity for beginning and concluding the vital cycle of the virus. The above mentioned cases and other recent findings show that, besides well-known protein mechanisms of apoptosis control by viruses, there are other strategies of apoptosis regulation based on translational repression by miRNAs (Klase et al, 2009). |
| Apoptosis may also represent one of the most important links between viral infections and oncogenesis. All tumor related viruses seem to share the same approach to interfere with apoptosis and cell-cycle genes (Damania, 2007). Thus, they may directly promote neoplastic transformation of infected cells or facilitate cancer insurgence in favourable contexts. For example, the Kaposi Sarcoma Herpes Virus encodes a protein called LANA, which inhibits the tumor suppressor protein p53, thus promoting cell survival, virus latency and oncogenesis (Si and Robertson, 2006). |
| In this article we present computational predictions of human targets for v-miRNAs encoded by two tumor related viruses, Epstein-Barr Virus (EBV) and Kaposi Sarcoma Herpes Virus (KSHV). The predictions are computed by a novel approach, which combines thermodynamics and empirical constraints in order to produce reliable miRNA/target duplexes. |
| Our results demonstrate v-miRNA/human target pairs which possess good binding scores, some of which are also supported by publicly available data on gene expression profiles. These findings are consistent with previously reported regulation mechanisms and offer a set of candidate targets for further biological validation. |
Materials and Methods |
| Computational prediction of v-miRNA/target duplexes |
| v-miRNAs/human target interactions were predicted by miRiam, a novel tool capable of screening large databases of miRNAs in order to identify the most probable binding sites on a given mRNA, in a reasonable time. miRiam is based on a combined approach which makes use of thermodynamics and pairing constraints inferred from known miRNA/target pairs. |
| The importance of target secondary structure in miRNAbased silencing has been previously demonstrated (Robins et al, 2005). Target binding sites must be accessible and interactions with miRNAs have to be energetically favourable (Kertesz et al, 2007). Moreover, to predict duplexes reliably several features inferred from known miRNA/mRNA pairs need to be taken into account. For example, miRNA 5’end contains the seed region, which is important in their binding with the target using near perfect complementarity. Nevertheless, the seed region may sometimes contain G:U wobbles, and the number of G:U wobbles in this region appears to be inversely related to silencing efficacy of the miRNA. Almost all known miRNA binding sites are located in the 3’ UTRs of target mRNAs, and multiple binding sites have been observed in several cases, potentially increasing the efficacy of silencing (Grimson et al, 2007). |
| Given a target mRNA sequence and a database of miRNAs, which can be retrieved from miRBase, miRiam performs the following steps: |
| i) Target accessibility evaluation: Base pairing probabilities of the target are computed by Vienna RNA package functions (Hofacker et al, 1994) and used to identify accessible regions, i.e. less paired sub-sequences. |
| ii) Alignment of miRNA/binding sites: The complementary base-pair alignment of accessible sub-sequences with miRNAs is then computed. A variant of the classical Needleman-Wunsch pairwise sequence alignment in which only matches of complementary base pairs are allowed is used. |
| iii) Post alignment filtering: All duplexes which do not satisfy previously described empirical sequence constraints are discarded. The remaining pairs are presented to the user ordered by their binding energies. |
| miRNAs and targets selection |
| EBV and KSHV miRNAs were obtained from miRBase (Griffiths-Jones et al, 2008). Target mRNA transcripts were downloaded from GenBank, chosen based on their Gene Ontology annotations. Genes annotated with the following terms were selected: Pro-apoptosis, cell cycle arrest, cell cycle checkpoint, negative regulation of cell cycle, negative regulation of cyclin-dependent kinase activity, and negative regulation of mitotic cell cycle. |
| Expression data |
| The expression profiles of the predicted targets were retrieved from the NCBI GEO Profiles database. Among the available datasets, NCBI GDS940 and GDS989 were selected as they were obtained from suitable biological specimens (infected cells and corresponding uninfected controls). For each predicted target, experiments performed on EBV and KSHV infected cells were selected and the ratios between the mean gene expression values in the infected and in the control samples were computed. The evaluation of fold change significance using formal statistical methods was not possible due to the small number of available samples. For this reason the observed ratios should be considered as qualitative descriptors supporting or confuting the proposed hypotheses. The data on the v-miRNA expressing tissues were obtained from the literature (Pfeffer et al, 2004; Pfeffer et al, 2005; Cai and Cullen, 2006; Cai et al, 2006; Kim et al, 2007; Marshall et al, 2007; Zhu et al, 2009) and from the miR-Z database (Hausser et al, 2009). |
Results |
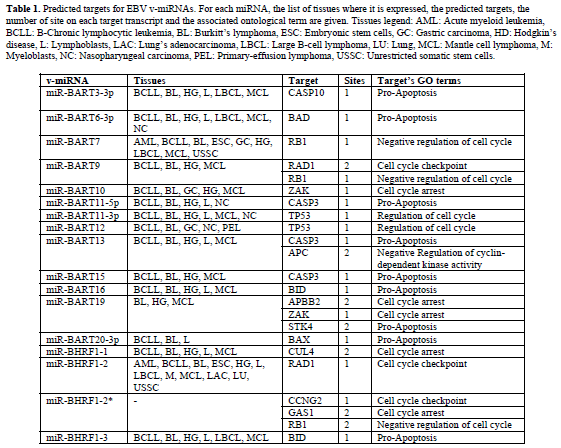
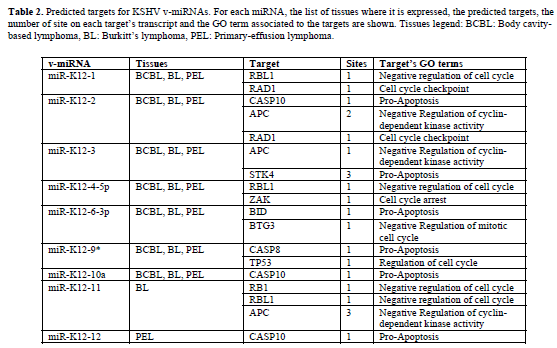
| Representative results of our predictions are shown in Tables 1 and 2. Table 1 contains the EBV miRNAs predicted targets while Table 2 shows the predictions for KSHV miRNAs. For each v-miRNA, the tissues where it is expressed, the predicted human targets and. Only the transcripts with perfect matches with the seed of vmiRNAs in structurally-accessible regions were selected (maximum one G:U wobble allowed in the seed region). These results indicate the putative capability of EBV and KSHV to interfere with the expression of host genes involved in apoptosis and cell cycle regulation. Moreover, some of the predictions are supported by gene expression profiles in EBV and KSHV infected cells. |
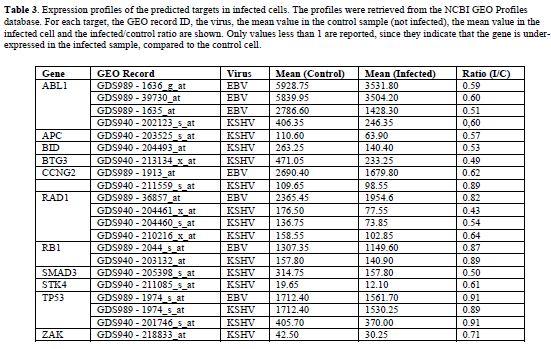
| Table 3 shows, for each predicted target, the ratios between the mean expression values in the infected and control cells. A value less than 1 indicates that the gene is under-expressed in the infected samples, compared to the control cells. Only these values are reported, since they are the significant ones concerning our analysis. The complete data are given in supplementary tables (Tables S1-S3). |
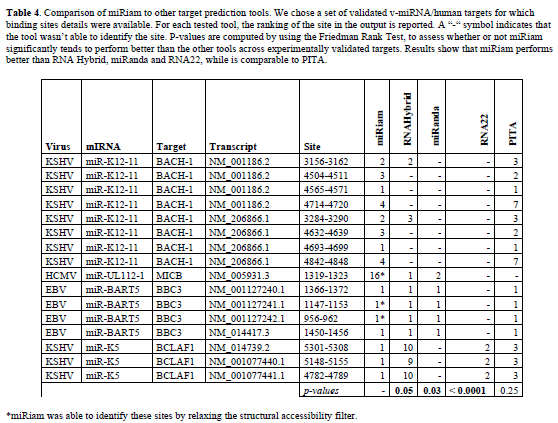
| Finally, in order to assess the reliability of the proposed method, we performed a comparison between miRiam and some of the most popular available tools for miRNA target prediction: RNA Hybrid, miRanda, RNA22 and PITA (Rehmsmeier et al, 2004; John et al, 2005; Huynh et al, 2006). For this purpose, we chose a set of validated vmiRNA/ human targets for which binding sites details were available. Results are reported on Table 4. They show the capability of miRiam to identify all the validated binding sites, in most cases as top ranking. We evaluated the statistical significance of our results by the Friedman rank test. While miRiam is comparable to PITA, in terms of number of correct sites identified, it outperforms the other tools in terms of the number of correct binding sites and their ranking. |
 |
 |
 |
 |
Discussion |
| v-miRNAs vs human targets |
| Epstein Barr herpesvirus (EBV) is associated with several tumor types, such as Burkitt’s lymphoma, nasopharyngeal and gastric carcinoma (Damania, 2007). EBV expresses two families of miRNAs, which are located in the BART and BHRF1 genes (Pfeffer et al, 2004; Cai et al, 2006). The BHRF1 miRNAs are expressed in infected primary lymphoma, Burkitt’s lymphoma and lymphoblast cell lines during late latency and lytic stages, while BART members are always expressed in lymphoblasts in all latent stages. miR-BART12 has also been found in infected nasopharyngeal and gastric carcinoma cell lines (Kim et al, 2007). |
| Kaposi’s sarcoma-associated herpesvirus (KSHV) is associated with several forms of cancer like Kaposi’s sarcoma and primary effusion lymphomas (Damania, 2007). KSHV expresses an array of 13 miRNAs in infected primary effusion lymphoma and endothelial cell lines (Pfeffer et al, 2005; Marshall et al, 2007). miRBHRF1- 3, miR-BART16 in EBV and miR-K12-6-3p in KSHV show good matches to BID transcript. Moreover, BID mRNA levels decrease after viral infection. The BID silencing could cause the partial block of the connection between the extrinsic apoptotic pathway and the mitochondrial one. BID is a target of proteolytic activity of CASP8, which is activated by cell surface death receptors. Truncated BID translocates in the mitochondria engaging BAX to trigger its pro-apoptotic activity, starting the release of the cytochrome c. This is a key event in the formation of the apoptosoma complex (cytochrome c, APAF1 and CASP9) for the beginning of the intrinsic death pathway. Consequently, BID inhibition may lower cell sensitivity to external signals and promote resistance to apoptosis. Moreover, it has been shown that BHRF, the host gene of these miRNAs, encodes a protein which acts downstream of BID cleavage and upstream of mitochondrial damage, resulting in inhibition of TRAIL-induced apoptosis (Kawanishi et al, 2002). This makes plausible a function of these miRNAs as repressors of apoptosis. |
| TP53 is a predicted target for EBV miR-BART11-3p, miR-BART12 and KSHV miR-K12-9*. In particular two different favorable binding sites for both EBVmiRBART12 and KSHV-miR-K12-9* have been identified in TP53 transcript. We also observed a significant decrease in TP53 transcript levels following EBV and KSHV infections (see Table 3). The presence of multiple binding sites for the same miRNA has been observed in several experimentally validated pairs. This potentially increases the degree of translational suppression. TP53 is an important regulator of cell cycle arrest, apoptosis, and cellular senescence and it may be activated by various forms of cellular stress, as viral infection. Several reports provide evidence of how TP53 is involved in antiviral activity and how viruses in their infection strategy lead to protein inactivation of TP53 (Si and Robertson, 2006; Pampin et al, 2006). That being so, a further repression device of TP53 based on RNA Interference from viruses looks plausible. Moreover, some data suggest that functional TP53 can promote the lytic cycle of some viruses. Consequently a repression strategy controlled by viral microRNAs could be an efficient and inexpensive switch on / off system of TP53 activities. |
| Our data suggest that Cyclin G2 (CCNG2) could be target of miRNAs from both EBV and KSHV. This pro-apoptotic protein has been found to be downregulated in many different tumours (Choi et al, 2009), and a decrease in its activity has been associated with improved proliferation. Its key role in cell-cycle and cell plasticity in cancer is underlined by the observation that the therapeutic effect of rapamicyn is also exerted through the enhancement of its transcription rate (Kasukabe et al, 2008). |
| The ability to interfere with tp53 pathway through the post transcriptional regulation of some of its fundamental components (tp53, BID, cyclin g2) may constitute a mechanism to tightly and efficiently control the host survival rate improving replication and diffusion of viral progeny. Similarly, other genes involved in apoptosis and in the regulation of cell cycle, BAD, BTG3, BAX, RAD1, ZAK and RB1, among others, are predicted to be targets of EBV and KSHV miRNAs. The repression of these genes may help the virus to gain control of the host cells, thus favoring the accomplishment of the viral life cycle. |
| v-miRNAs involvement in cancer |
| Viruses are often involved in cancer development. They may directly promote cell malignancy or facilitate the progression of already established tumors (Damania, 2007). While protein based mechanisms of interaction between virus and hosts have been widely described, a direct involvement of v-miRNAs in cancer is not yet demonstrated. However, since experimental evidences show the capability of v-miRNAs to perturb host genes expression (Barth et al, 2008; Klase et al, 2009), their contribution to tumor insurgence is plausible. Moreover, a miRNA based host regulation system may represent a faster way for viruses to counteract cellular response to infection promoting their latency. According to Pfeffer and Voinnet (Pfeffer and Voinnet, 2006), v-miRNAs could act as direct oncogenes or indirectly promote cancer development and diffusion, by supporting external etiologic agents. |
| The predictions presented in this study are consistent with these scenarios. In particular the inhibition of central nodes of the apoptotic pathways may lead to loss of cell cycle control by the cell itself. |
Conclusions |
| The aim of this work was to produce a set of plausible hypotheses of involvement of v-miRNAs and human apoptosis genes in cancer development. In the study, we have presented computational predictions of v-miRNAs and human targets interactions, and show miRNA/target duplexes which were energetically favourable and consistent with other experimentally validated pairs. Moreover, the target binding sites were characterized by high structural-accessibility for interactions with miRNAs. Future works will focus on providing experimental validations of such estimated interactions. Extension of predictions to other reasonable human transcripts, such as additional nodes of apoptosis pathway, cell-cycle regulators and genes involved in immune response, will be performed. |
Acknowledgements |
| The authors would like to thank Michael Levitt and Dennis Shasha for their helpful suggestions and comments, and the authors of the Vienna RNA Package for providing their software. This work was supported in part by Sicily Region Grants PROGETTO POR 3.14: “Ricerca e Sviluppo suite di programmi per l’analisi biologica, denominata: BIOWARE”. |
Compteting Interests |
| None declared |
List of Abbreviations |
| miRNA: microRNA |
| v-miRNA: viral microRNA |
| EBV: Epstein-Barr herpesvirus |
| KSHV: Kaposi’s Sarcoma associated herpesvirus |
| SV40: Simian Virus 40 |
| UTR: Un-translated region |
References
|