Review Article - Journal of RNA and Genomics (2018) Volume 14, Issue 1
Polycomb at the Crossroads: An Integrated Model with LncRNA- and Silencermediated Repression
Nadine Obier1 and Pierre Cauchy1,2*1Institute of Cancer and Genomic Sciences, University of Birmingham, Vincent Drive, B15 2TT Birmingham, UK
2Max Planck Institute for Immunobiology and Epigenetics, Stübeweg 51, 79108 Freiburg im Breisgau, Germany
Received: 31 July 2017; Revised: 18 December 2017; Accepted: 27 December 2017; Published: 03 January 2018
© Copyright The Author(s). First Published by Allied Academies. This is an open access article, published under the terms of theCreative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0). This license permitsnon-commercial use, distribution and reproduction of the article, provided the original work is appropriately acknowledged with correct citation details.
Abstract
Keywords
Genetics, gene silencing, gene expression, lncRNA-, Polycomb
Introduction
Gene expression is achieved by the stepwise recruitment of general transcription factors (GTFs) and RNA Polymerase II (Pol II) to promoter regions. This process results in proximal pausing of Pol II, and is followed by p-TEFb-mediated elongation across the gene body (Marshall and Price 1995; Peng et al, 1998; Lee and Young 2000; Koch et al, 2008). However, this process must be tightly developmentally and spatially regulated in order to achieve correct gene expression across cell types. A first level of control entails the chromatin superstructure: transcription is associated with the 10 nm euchromatin, while the 30nm heterochromatin remains mostly inaccessible to the transcriptional machinery (Razin and Gavrilov 2014). Heterochromatin is preferentially marked by the histone tail modification H3K9me3 (Nakayama et al, 2001). Facultative heterochromain, as opposed to constitutive heterochromatin, is induced by the recruitment of histone deacetylases (HDACs) (Thiel et al, 2004; Vaquero et al, 2004). Facultative heterochromatin however allows returning to a transcriptionally active state. One of the hallmarks for facultative heterochromatin is the H3K9me2 mark (Trojer and Reinberg 2007; Sugeno et al, 2016). Well described examples of repressors known to induce facultative heterochromatin formation include REST/NRSF, which recruits Sin3A as well as HDACs (Naruse et al, 1999; Pennock et al, 2013). One other particular state of facultative heterochromatin is marked by H3K27me3, which is decorated and read by Polycomb group proteins (PcG) (Cao et al, 2002). Polycomb-mediated gene repression is a widespread mechanism involved in the silencing of pluripotency and lineage-specific genes during development and differentiation (Mohn et al, 2008; Kashyap et al, 2009). The Hoxa-d gene clusters, for example, are well-characterized targets of PcG-mediated silencing (Satijn and Otte 1999). PcG complexes consist of two mutually exclusive complexes, PRC1 and PRC2. While PRC2 can both read and decorate H3K27me3, PRC1 recognizes H3K27me3 but deposits H2AK119ub (Kuzmichev et al, 2002). Thus, both PcG- and HDAC-mediated processes of gene silencing constitute epigenetic repression mechanisms acting in parallel. Both mechanisms further entail sequence-specific binding, being recruited to silencer elements and Polycomb responsive elements (PREs), respectively.
Euchromatin on the other hand is marked by H4K16ac as well as by other activation marks, depending on the state of gene activation (Lu et al, 2015) (Barski et al, 2007; Bannister and Kouzarides 2011). These are conditioned by further fine-tuning of transcription regulation which involves two main types of cisregulatory elements (CREs): enhancers and classical silencers, which have opposite effects on gene regulation (Ogbourne and Antalis 1998; Riethoven 2010). Ultimately, tallying up enhancer and silencer function dictates the epigenetic state of a gene locus, i.e. whether permissive (H3K9/K14ac, H3K27ac, H3K4me3) or non-permissive histone tail modification marks (H3K27me3) decorate the promoter of a given gene. Classical enhancer function involves recruitment of p300/CBP (which deposits the mark H3K27ac) by transcription factors (TFs) and looping to the promoter region, ultimately recruiting the preinitiation complex to initiate transcription (Buratowski et al, 1989; Tie et al, 2009). These interactions have been described to occur within transcriptional units, known as topological association domains (TADs), reflecting the tridimensional organization of the genome (Razin et al, 2016). At the nuclear level, transcription is thus proposed to occur in a coordinated manner via transcription factories, purportedly supported by the nuclear matrix (Rieder et al, 2012). Importantly, one of the modes to repress enhancer function is the establishment of new CTCF-sites which create insulators and physically disturb enhancer-promoter interactions (Xie et al, 2007). Other modes of gene silencing involve the recently identified long noncoding RNA-mediated repression, which has changed perceptions and research orientations in the field of transcription regulation (Royo and Cavaille 2008; Kung et al, 2013). LncRNA activation can repress transcription of genes in cis or in trans at almost every level of gene regulation (Kung et al, 2013) (Goodrich and Kugel 2006; Latos et al, 2012; Di Ruscio et al, 2013) (Yoon et al, 2013). Importantly, PcG- and lncRNA- mediated repression have been characterized to sharing a direct link, whereby PcG proteins reportedly co-localize and are co-transported with lncRNAs to their targets (Kaneko et al, 2014; Chu et al, 2015). However, whether those interactions are specific is still a hotly debated topic. Further, the distinction between PREs and classical silencers merits to be investigated. In this review, we describe and discuss the mechanisms underlying PolycomblncRNA-, classical silencer- mediated repression, as well as their possible relationship and cross-talk mechanisms.
PRC1
The core PRC1 complex contains Ring1a/b and one or two of the 6 polycomb group ring finger proteins (PCGF1-6) (Simon and Kingston 2009; Aranda et al, 2015). The presence of PCGF2- 4 typically indicates the canonical PRC1 with CBX2,4,6-8 (PRC1.2 or PRC1.4). Other combinations correspond to noncanonical PRC1 complexes (PRC1.1, PRC1.3, PRC1,5 and PRC1.6). CBX2 is the only CBX protein to exhibit a functional DNA AT-hook binding domain, allowing it to efficiently bind DNA (Figure 1), perhaps at specific nucleosome dyads (Kaustov et al, 2011). Crucially, the phenotype for CBX2 knockout mice is male-to-female sex reversal due to aberrant repression of Sry and Sox9 (Katoh-Fukui et al, 1998; Katoh-Fukui et al, 2012). There is evidence of that the AT-hook nucleotide binding motif adjacent to the chromodomain is conserved among CBX proteins, even if the only functional one is found in CBX2 (Senthilkumar and Mishra 2009). The transcription factor E2F6, part of the cell-cycle related E2F family but lacking the transactivation and pocket protein binding domains, is a facultative member of the PRC1 complex (Trimarchi et al, 1998; Trimarchi et al, 2001; Attwooll et al, 2005) (Figure 1). The transcriptional repressor Bcl6 was also shown to recruit the co-repressor B-COR to form an alternative PRC1 complex, PRC1.1 (Huynh et al, 2000). TFs such as GATA3 were also shown to interact with PCGF4 (Bmi1) in Th2 cells, suggesting that PRC1 can occasionally be recruited by activating TFs (Hosokawa et al, 2006; Armstrong 2012). It is also generally believed that that CBX proteins can also be recruited by Phc1-3, and that Ring1a/b by ncRNAs (Armstrong 2012). Importantly, H3K27 methylation is recognized by PRC1 (Cao et al, 2002). Canonical PRC1 binds H3K27me3 and H3K9me3, via the chromodomains of CBX2 binding H3K27me3, and those of CBX4-7 showing higher affinity towards H3K9me3 (Kaustov et al, 2011; Aranda et al, 2015). However, Gao et al, showed that genome-wide, CBX2-PRC1 clearly occurs with H3K27me3 genome-wide, while most core PRC1 complexes (Ring1b and PCGF1-6) do not co-localize with H3K27me3 (Gao et al, 2012). In fact, ncPRC1.1 targets are independent of H3K27me3, which confers it an important role in leukemogenesis, since PRC1.1 components are overexpressed in leukemic stem cells (van den Boom et al, 2016). Importantly, Ring1B also catalyzes ubiquitination of H2AK119 (Kuzmichev et al, 2002), triggering chromatin compaction and Pol II pausing, notably on the inactive X chromosome (de Napoles and Francis, 2004; Cao et al, 2005; Margueron et al, 2008).
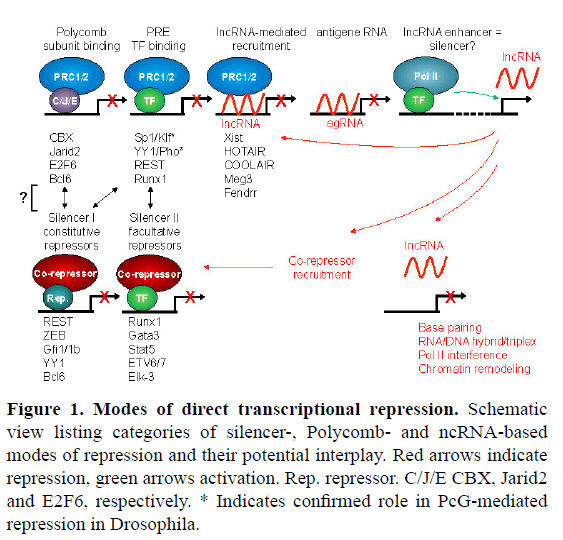
Figure 1: Modes of direct transcriptional repression. Schematic view listing categories of silencer-, Polycomb- and ncRNA-based modes of repression and their potential interplay. Red arrows indicate repression, green arrows activation. Rep. repressor. C/J/E CBX, Jarid2 and E2F6, respectively. * Indicates confirmed role in PcG-mediated repression in Drosophila.
PRC2
In vertebrates, the core PRC2 complex is made up of three essential subunits: Suppressor of Zeste 12 (Suz12), one of the Embryonic ectoderm development (Eed)1-4 proteins (extra sex combs in Drosophila) and one of the mutually exclusive Enhancer of zeste (EZ) histone K27 methyltransferase homologs Ezh1 or Ezh2 (Kuzmichev et al, 2002). The nucleosome remodeling factor RbAp46-48/Nurf55 is also co-purified along with Eed/ Ezh2 (Tie et al, 2001; Kuzmichev and Muller, 2002). Nurf55 is believed to stabilize the PRC2-nucleosome interaction (Liu and Zhu 2017). Contrary to Ezh2, Ezh1 displays low levels of histone methyltransferase activity and appears to efficiently repress transcription and to induce chromatin compaction (Margueron et al, 2008). Even the expression of Ezh1 and 2 was originally described as almost mutually exclusive (Laible et al, 1997), although Ezh1 expression was actually shown to be ubiquitous while Ezh2 expression is more associated with proliferating cells (Margueron et al, 2008). An elegant model would entail that Ezh1 is required for the maintenance of PRC2 in resting cells (Son et al, 2013). Interestingly, Ezh1 was also reported to promote elongation (Mousavi et al, 2012). The Eed subunit antagonizes H3K27ac/CBP, as it interacts with HDAC2 (van der Vlag and Otte 1999). Eed possesses a WD40 aromatic cage domain, capable of binding H3K27me3, and causes PRC2 to methylate further away from PRC2 nucleation sites (Margueron et al, 2009). This domain is also responsible for the interaction with Ezh2 (Denisenko et al, 1998). Importantly, Y365A Eed mutants are unable to bind H3K27me3 (Lewis et al, 2013; Barnash and et al, 2017). Suz12 contains a zincfinger domain and participates in the anchoring of Ezh2 to the chromatin (Nekrasov et al, 2005). Facultative PRC2 members are numerous, but one of the most prominent ones is the JmjC demethylase family member Jumonji AT-rich domain 2 (Jarid 2). Members of the JmjC family of demethylases are normally capable of demethylating H3K4me2 and 3, however no demethylase activity was observed for Jarid2 (Klose et al, 2006). Jarid2 possesses an AT-rich DNA interaction domain (ARID) (Agulnik et al, 1994; Kortschak et al, 2000) and can recruit PRC2 to the chromatin (Pasini et al, 2010) (Figure 1). Importantly, Jarid2 is not ubiquitously expressed (Son et al, 2013), thus making recruitment of PRC2 facultativelydependent on Jarid2 and likely involving other actors. On the other hand, Suz12 recruitment to the chromatin is thought to be mediated by Mtf2 and Phf1, while in the case of Eed this process is likely TF-dependent (Armstrong el al, 2012).
PRC1 or PRC2, who tracks whom?
Because PRC1 can read but is not able to induce H3K27me3 decoration, a simple model would entail that PRC1 tracks PRC2. It indeed seems that bivalent genes that are PRC1- positive also seem to maintain higher levels of H3K27me3, as opposed to PRC2-only genes (Ku et al, 2008). However, as PRC1 ablation results in the loss of PRC2 recruitment, while the converse is not true, it is generally believed that PRC2 tracks PRC1 (Kahn et al, 2016). Crucially, Jarid2 can recognize and bind H2AK119ub, deposited by PRC1, which also argues in this direction (Cooper et al, 2016). Further, in X-inactivation, PRC1 efficiently deposits H2AK119ub in the absence of H3K27me3 in Eed-/- ESC (Schoeftner et al, 2006). However, canonical PRC1 also seems to be partly involved in the maintenance of H3K27me3 as it was recently shown that Drosophila Polycomb and its mammalian CBX orthologs are able to prevent CBPmediated histone acetylation by directly targeting its catalytic domain (Tie et al, 2016). Interestingly however, PRC1 and PRC2 can act redundantly to repress genomic repeats, although in the absence of both, stem cell differentiation is abrogated (Leeb et al, 2010). The order in which PRC1 and PRC2 can track each other sequentially is depicted in Figure 2. Further, neither is required for targeting of H2A.Z to developmental genes in embryonic stem cells (Illingworth et al, 2012). Overall, these various mechanisms suggest that PRC1 and PRC2 can track each other in both directions, and that PRC2 can track itself as well.
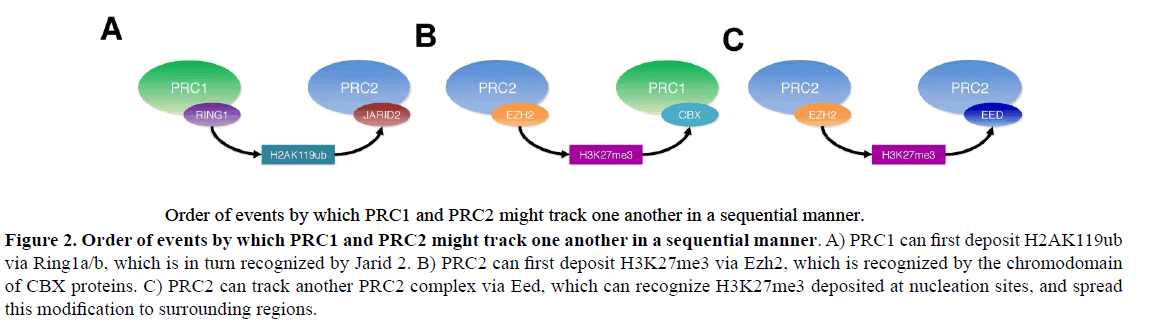
Figure 2: Order of events by which PRC1 and PRC2 might track one another in a sequential manner.
Order of events by which PRC1 and PRC2 might track one another in a sequential manner. A) PRC1 can first deposit H2AK119ub
via Ring1a/b, which is in turn recognized by Jarid 2. B) PRC2 can first deposit H3K27me3 via Ezh2, which is recognized by the chromodomain
of CBX proteins. C) PRC2 can track another PRC2 complex via Eed, which can recognize H3K27me3 deposited at nucleation sites, and spread
this modification to surrounding regions.
The balance between polycomb repression and gene activation
Polycomb activity is generally considered to confer a repressed or poised chromatin state. However, it must compete with activating transcriptional mechanisms. The main modes PcGmediated promoter H3K27me3 decoration have been proposed to entail i) spreading from nucleation sites, ii) PRE-promoter looping mechanisms or iii) a translocation mechanism (Bauer et al, 2016). Changes to promoter regions can also entail H1 recruitment, as PcG groups were shown to interact with the HDAC SIRT1 (Furuyama et al, 2004). SIRT1 is known to specifically mediate facultative heterochromatin formation to proximal and not distal regions (Vaquero et al, 2004). Polycomb-dependent repression and classical CBP- or Setmediated gene activation are essentially antagonistic: as a result, the presence of H3K27ac, H3K4me3 or H3K36me2/3 is mutually exclusive with Polycomb activity (Tie et al, 2009; Schmitges et al, 2011). Specifically, a landmark study using a thermosensitive EZ mutant, also showed widespread decoration of H3K27ac instead of H3K27me3 in the absence of EZ function in Drosophila (Zenk et al, 2017). This work however showed maintained PRC2 occupancy in those sites, suggesting a loss of H3K27ac antagonization. In development, H3K27me3 was found to specifically mark oocytes and thus to constitute a major form of imprinting up until the early embryo stages (Pathak and Feil 2017). Importantly, in differentiating Eed-/- embryonic stem cells, the expression of H3K27me3-marked pluripotency genes is aberrantly maintained (Obier et al, 2015). In these cells, although H3K27me3 signal is absent, gene upregulation is not coupled with the loss of H3K27me3, but rather with increases in H3K27ac (Ai et al, 2017). Interestingly, Eed-/- and WT – specific H3K27ac peaks are mostly distal elements (Ferrari et al, 2014), suggesting a promoter-independent regulation of the balance between H3K27me3 and H3K27ac for Eed. In fact, PRC2 nucleation sites were shown correspond to long-range interactions (van Heeringen et al, 2014; Schoenfelder et al, 2015). H3K27me3 spreading can be observed during development, from domains, in a segmental manner as for Hox genes or from nucleation sites as seen in differentiating T-cells (Zhang et al, 2012; Bowman et al, 2014). Further, while expression levels of genes with H3K27me3-marked promoters can be undetectably low, their promoters are usually still DNaseI hypersensitive and thus responsive to fast regulation (Goode et al, 2016). Typically, H3K4me3- and H3K27me3- marked bivalent chromatin and promoters are associated with developmental genes in pluripotency cells (Bernstein et al, 2006a; Sanz et al, 2008; Young et al, 2011) and with cancer-upregulated genes (Rodriguez et al, 2008). Finally, nascent RNA and eRNA were shown recruit PRC2, which could suggest a targeted mode of repression as CBP binding to eRNA stimulates histone acetylation (Vos et al, 2016; Bose et al, 2017). Ezh2 was also found to to be involved in RNA-RNA pairing between small RNA and nascent RNA transcripts, suggesting its recruitment at areas of transcription initiation (Turner and Morris 2010). Conversely, in addition to corresponding to DNase hypersensitive sites, TFs and Pol II were unexpectedly found in promoters of genes repressed by polycomb complexes (Breiling et al, 2001; Boyer et al, 2006), suggesting a balance of polycomb-mediated repression and coactivators in genes poised for later upregulation.
Cross-talk with H3K9me2/3?
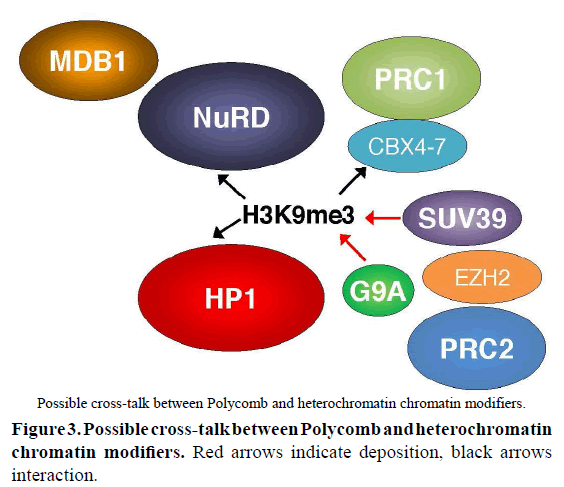
Long-standing questions in the field of transcriptional regulation are whether H3K27me3- and H3K9me2/3-mediated gene repression are entirely independent or interchangeable to some degree, or even whether they can occur successively. Although CBX4-7 can bind H3K9me3, evidence of targeting of Polycomb complexes to heterochromatin remains elusive, as well as that of cross-talk between H3K9me2/3 and H3K27me3 decoration. However, it was shown that Jarid2 can initiate decoration by H3K9 methylation (Shirato et al, 2009), consistent with the fact that HP1 can bind EZ, in addition to SU(var) 3-9 and trithorax (Bannister et al, 2001; Boyer et al, 2006). Further, PRC1 colocalizes with paternal pericentric heterochromatin (Tardat et al, 2015). It is however still unclear whether H3K9 methylation can compensate for the loss of H3K27me3 in PRC2 subunit knockouts, even if there is evidence for cross-talk between H3K9me3 and H3K27me3 decoration in X-inactivation (Escamilla-Del-Arenal et al, 2013). In fact, binding to H3K9me3 by HP1 is increased by the presence of H3K27me3 (Boros et al, 2014), indicating that PRC2 components can be co-recruited to bind H3K9me3 at least indirectly. This is further supported by findings that showed that G9a could recruit PRC2 (Mozzetta et al, 2014), suggesting some compensation mechanism. The chromatin remodeler NuRD, which also shares Nurf55, facilitates PRC2 recruitment by mediating deacetylation of H3K27 (Reynolds et al, 2012). NuRD is also able to bind to H3K9me3 in pericentric chromatin (Helbling Chadwick et al, 2009; Sims and Wade 2011), in addition to H3K4me3 and H3K27me3 (Reynolds et al, 2012). The H3K9me3-related proteins with which Polycomb proteins can interact is shown in Figure 3. However, while HDAC inhibition prevents binding of PRC2 to NuRD, deletion of PRC2 components does not seem to affect targeting of NuRD, suggesting that in this case, both mechanisms could be independent (Reynolds et al, 2012). The difference may lie in cell-cycle progression, or in the dynamics of DNA methylation, as both HP-1 and NuRD can share a link to methylated DNA (Fujita et al, 2003; Clouaire and Stancheva 2008) (Jiang et al, 2004).
Polycomb response elements (PREs)
PREs were shown to coincide with CpG islands (Boyer et al, 2006; Tanay et al, 2007; Ku et al, 2008; Mendenhall et al, 2010), likely constituting their main mode of silencing as CpG islands are usually not subject to DNA methylation (Mendenhall et al, 2010). Generally, PREs correspond to areas of increased density of H3K27me3 around CpG islands compared to non-CpG sites (Fenouil et al, 2012). GC-rich sequences in activating, motifdepleted CpG islands were also shown to recruit PRC2 in mammalian cells, hinting at a specific mode of recruitment for subset of genes (Mendenhall et al, 2010). Further, PRC2 was shown to be recruited at G quadruplexes (Wang et al, 2017). YY1/ Pho motifs are commonly found in PREs of Drosophila (Mihaly et al, 1998; Ringrose and Paro 2007). In mammals, Ezh2, YY1, and HDAC1 co-localize at muscle-specific regulatory elements, resulting in their specific silencing (Caretti et al, 2004). Mechanistically, YY1 was shown to interact with EED via the REPO domain of YY1 (Satijn et al, 2001; Wilkinson et al, 2006). The role of YY1 in mammalian PcG function is however disputed (Bauer et al, 2016). Drosophila PREs are further enriched in GAGA motifs, bound by the TF GAF (Strutt et al, 1997; Ringrose and Paro 2007). Interestingly, GAGA motifs are also found in target regions of the NURF chromatin remodeler (Kwon et al, 2016), thus providing another base for sequencespecific PRC2 recruitment and nucleosome remodeling. So far, a functional role for GAF-mediated PcG recruitment was only confirmed in Drosophila (Bauer et al, 2016). Other factors that have been characterized as having prominent other roles in gene regulation are also found in mammalian PREs, such as Runx1 and REST (Dietrich et al, 2012; Yu et al, 2012), reviewed in (Bauer et al, 2016). Drosophila PREs are further characterized by the presence of Zeste, Grh and Sp1 motifs (Ringrose et al, 2003; Ringrose and Paro 2007). In mammals, Oct4, Nanog and Sox2 co-localize with PREs at significant subsets of key developmental genes (Lee et al, 2006). Regulatory regions of Eed-dependent pluripotency genes are also enriched in Oct4 binding sites (Obier et al, 2015). However, their functional role in PRC1/2 recruitment remains to be demonstrated. TFs that have been shown to be involved in Polycomb recruitment are shown in Figure 1. Functionally, at least certain PREs can act as general transcriptional silencers by default (Sengupta et al, 2004). However, PREs are usually sites of extensive noncoding transcription (Natoli and Andrau 2012; Barlow and Bartolomei 2014).
lncRNAs
The field of transcriptional regulation has recently been focusing on long non-coding RNAs (lncRNAs), whose activation represses transcription of genes or gene clusters in cis or in trans (Royo and Cavaille 2008; Kung et al, 2013). Repression is achieved via several distinct mechanisms at almost all levels of gene regulation. Classical modes of action entail recruitment of the transcriptional machinery and the mere act of transcription in the case of antisense lncRNAs (Goodrich and Kugel 2006; Latos et al, 2012; Di Ruscio et al, 2013). There is also evidence of that a number of lncRNAs share promoter elements with protein coding genes, in which case they would hijack preinititation complex in their favor (Khachane and Harrison 2010). Interference can also occur at the posttranscriptional level via base-pairing in cis or in trans (Gong and Maquat 2011; Yoon et al, 2013) (Figure 1). LncRNAs can also induce dynamic changes to genes in cis-, notably by influencing nucleosome positioning and chromatin remodeling (Bohmdorfer and Wierzbicki 2015). Importantly, a viral ncRNA was shown to recruit TFs by binding nascent RNA to drive Pax5 in to the EBV genome (Lee et al, 2015), reviewed in (Lee and Steitz 2015). This alternative mode of action remained largely uninvestigated and could constitute an important mechanism in lncRNA function. Other modes of action include lncRNA interaction with nascent the RNA processing and matchmaking machinery (Meredith et al, 2016). Direct RNA-DNA hybrid or triplex structures have also been identified, such as in the case of Meg3, which regulates TGF beta signaling, or Fendrr (Grote et al, 2013; Mondal et al, 2015). Triplex interactions with lncRNA were also shown to co-occur with PRC2 (Li et al, 2016).
Paradoxically, lncRNAs were also shown to induce transcription in some cases. Notably, the simple act of lncRNA transcription can also induce TF recruitment to target sites (Clark and Blackshaw 2014; Takemata et al, 2016). A number of lncRNAs originate from enhancer regions, such as in the case of the Lockd lncRNA, whose transcript is dispensable for Cdkn1b expression while its actual locus is not (Paralkar et al, 2016). However in this case, this could suggest that a number of lncRNAs may merely correspond to elongation originating from enhancer regions.
A non-exhaustive list of notable lncRNAs includes Xist, which plays a paramount role in X-inactivation by coating the inactive X chromosome (Chu et al, 2015), as well as being one of the best characterized lncRNAs to date (Augui et al, 2011); HOTAIR, an lncRNA located in the Hoxc cluster but that represses transcription of the HoxD cluster by binding between Hoxd3 and Hoxd4 (Rinn et al, 2007). The Airn lncRNA, located downstrean of Igfr2 but on the opposite strand, achieves Igfr2 silencing by promoting elongation on its opposite strand, thereby interfering with Pol II recruitment at the Igfr2 promoter and gene body on the sense strand (Latos et al, 2012). The Haunt lncRNA locus itself also appears to contain enhancer regions for neighboring Hoxa genes, its transcription thus preventing activation of those enhancers and the lncRNA itself repressing those genes (Yin et al, 2015). Other examples of lncRNAs include the antisense Spi1-AS (Ebralidze et al, 2008) and Cebpa-AS (Di Ruscio et al, 2013) lncRNAs, the latter interacting with DNMT1 to block DNA methylation in Cebpa.
Polycomb and lncRNAs
Several reports have shown that both PRC1 and PRC2 can bind RNA (Geisler and Coller 2013). For PRC1, in vitro, CBX7 can bind RNA and its interaction with H3K27me3 is RNA dependent (Bernstein et al, 2006b). Most prominently, PRC1 was pulled down as one of the interacting complexes in a chromatin isolation by RNA purification assay followed by mass spectrometry (ChIRP-MS) targeting Xist (Chu et al, 2015), whereby Ring1b is recruited by Xist (Schoeftner et al, 2006) (Figure 1). Further, PCGF3/5-PRC1 was shown to initiate Polycomb recruitment to the inactivated X chromosome (Almeida et al, 2017). ANRIL, a lncRNA that is involved in silencing the INK4a/b locus, as well as both Cat7 and catl7 zebrafish lncRNAs were also shown to interact with PRC1 (Yap et al, 2010; Ray et al, 2016).
In the case of PRC2, co-localization with RNA occurs genomewide, demonstrating its capacity to bind RNA generally (Hendrickson et al, 2016). Suz12 and Ezh2 further exhibit cryptic RNA binding properties (Betancur and Tomari 2015). Direct PRC2 recruitment was actually shown to be an lncRNAmediated mechanism via Jarid2 binding (Kaneko et al, 2014). An Ezh2 CLlP-Seq revealed PRC2-RNA interactions in HCT116 cells, with a lncRNA located in intronic regions of Smyd3, an H3K4me3 methyltransferase, whereby overexpression of this lncRNA reduces expression levels of Smyd3 (Guil et al, 2012). Other cases include the Kcnq1ot1 lncRNA which recruits PRC2 to synergize with G9a to silences Kcnq1 (Pandey et al, 2008), representing another example of cross-talk with H3K9me3. This finding was confirmed by a study via Ezh2 CLIP-Seq that mapped to the Kcnq1ot1 as well as Meg3 lncRNAs (Kaneko et al, 2014). Notably, a direct relationship was established relationship between X-inactivation and PRC2, as the inactive X chromosome is decorated by high levels of H3K27me3 (Chadwick and Willard 2004; Chaumeil et al, 2006). In fact, PRC2 is bound by the short repeat RepA, located in the 5’ end of Xist, redirecting PRC2 to the targeted X chromosome (Wutz et al, 2002; Zhao et al, 2008). This was confirmed in at least cross-linking conditions, whereby Xist was pulled down by RIP targeting core elements of the PRC2 complex (Brockdorff 2013) (Figure 1). Ezh2 was shown to be indeed bound by the RepA repeat. Functional assay also showed a loss of Xist coating in Eed-/- cells (Maclary et al, 2017). Mechanistically, Xist recruits PRC2 distally but non-specifically to accessible regions of the targeted X chromosome, and spreads along transcribed regions (Engreitz et al, 2013). However, while Stochastic Optical Reconstruction Microscopy experiments (STORM) did reveal co-localization of Xist and PRC2, this interaction was characterized by a low stoichiometry, indicating a possible hit and run mechanism (Sunwoo et al, 2015). Further, while PRC2 represses transcribed genes on the imprinted inactive X chromosome in mice, few X-linked genes are derepressed in Eed-/- cells (Maclary et al, 2017). These regions show active hallmarks including H3K27ac but not H3K4me2. Therefore, it was suggested that PRC2 acts as a brake to prevent induction of transcribed genes on the inactive X chromosome, whereby it is acting in cis to Xi.
Another classical example of PRC2 binding to lncRNAs, but in trans, is HOTAIR. In fact, PRC2 binds with high affinity to HOTAIR (Rinn et al, 2007). Suz12 interacts with HOTAIR via its zinc finger domain (Rinn et al, 2007). Surprisingly, however, PRC2 was shown to be dispensable for HOTAIR-mediated repression (Portoso et al, 2017), reviewed in (Blanco and Guttman 2017), although possibly indicating a compensation mechanism. This finding was consistent with results from another study using ChIRP-Seq and ChIP-chip that showed that HOTAIR occupancy seems independent of Ezh2 (Chu et al, 2011), even though the HOTAIR binding site co-localizes with high Suz12, Ezh2 and H3K27me3 levels. Interestingly, this study also revealed that HOTAIR binds to a GA-rich homopurine motif at enhancer elements (Chu et al, 2011). that the fact that certain lncRNAs seem to exhibit protein sequenceand structure- specific binding to PRC2 actually gave rise to algorithms predicting association of lncRNAs with PRC2 based on human training models (Glazko et al, 2012). Such predictions could also be extended to binding sites of lncRNAs, as for classical motif discovery for TFs, given that at least HOTAIR seems to exhibit sequence-specific DNA binding. Other notable examples of PRC2 recruitment to lncRNAs include the lncRNAs COOLAIR (Swiezewski et al, 2009), ANRIL (Kotake et al, 2011) and ROR1-AS1 (Hu et al, 2017), Fendrr, which can also target MLL (Grote et al, 2013), as well as HoxA11-AS, which also targets LSD1 and DNMT1 (Liu et al, 2017). Overall, this supports important scaffolding roles for lnRNAs that direct key chromatin remodelers to their targets. The mechanisms by which this targeting is achieved however remain unclear.
Consistent with the fact that PRC2 nucleation sites correspond to long range interactions (Schoenfelder et al, 2015), it was suggested that PcG group bodies could constitute silencing factories as a parallel mechanism to transcription factories (Hodgson and Brock 2011). One could further speculate there might exist lncRNA factories whereby one lncRNA locus interacts with several others at the same time, targeted by lncRNAs directing PRC2 bodies (Figure 4). An indication for this might be that Ezh2 generally seems to bind the 5’ region of nascent RNA (Wutz et al, 2002; Zhao et al, 2008; Kaneko et al, 2014), perhaps reflecting a close and rapid interaction.
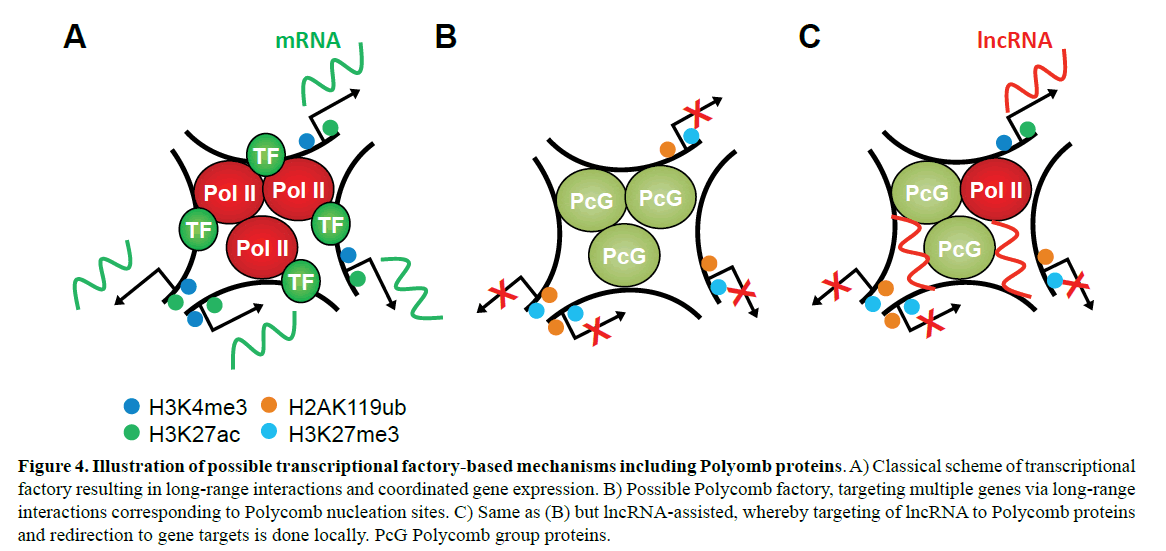
Figure 4: Illustration of possible transcriptional factory-based mechanisms including Polyomb proteins. A) Classical scheme of transcriptional factory resulting in long-range interactions and coordinated gene expression. B) Possible Polycomb factory, targeting multiple genes via long-range interactions corresponding to Polycomb nucleation sites. C) Same as (B) but lncRNA-assisted, whereby targeting of lncRNA to Polycomb proteins and redirection to gene targets is done locally. PcG Polycomb group proteins.
In terms of dynamics, even if those as well as the recruitment of PRC2 have been extensively described (Mohn et al, 2008; Pasini et al, 2010; Son et al, 2013; Narendra et al, 2015), signals that trigger PRC2 recruitment remain largely uncharacterized, even though there exists evidence that lncRNAs can be activated by signaling (Gao et al, 2016). However, at least RAS signaling is known to lift PRC2 binding via down-regulation of antisense lncRNAs (Kotake et al, 2011), perhaps hinting that conformational changes that lead to the recruitment of PRC2 by lncRNAs can be signaling-dependent. Finally, as a model, a recent study proposed a strand-specific mechanism for mutually exclusive bidirectional lncRNA at PREs/TREs, whereby the forward strand lncRNA recruits PRC2, while the reverse strand inhibits it (Herzog et al, 2014), potentially reflecting “switch” lncRNA loci leading to increased or decreased PRC2-mediated inhibition genome-wide.
Classical silencer elements
Silencers have long been recognized as regulatory regions bound by repressor elements or complexes leading to the decrease of interactions between the transcriptional machinery and promoter regions (Ogbourne and Antalis 1998). While enhancer elements and the general process of transcriptional activation have represented the bulk of the focus of recent research, notably using large-scale screens such as e.g. the ENCODE project (Neph et al, 2012), silencer elements generally remain less characterized (Ogbourne and Antalis 1998; Kolovos et al, 2012). Silencers are currently divided in two main classes: i) classic, position-independent silencers, and ii) passive silencers or position-dependent silencers, that flank promoters or enhancers, (Ogbourne and Antalis 1998) and that can be bound by repressors such as YY1 (Liu et al, 1994). It was proposed that binding of one or more repressive TFs to silencer elements establishes a repressive transcriptional context, overriding the activating nature of bound TFs (Ogbourne and Antalis 1998). A general description of silencers would be that they typically function by inhibiting recruitment of TFIID to promoter regions, although the mechanism by which this is achieved varies greatly (Ogbourne and Antalis 1998; Lee and Young 2000). This can be via direct interaction, or via recruitment of co-repressors and/or via recruitment of chromatin remodelers. Active silencers are typically located in open chromatin regions, as determined by locus-specific or genome-wide DNAseI hypersensitivity assays (Wu et al, 1979; Gross and Garrard 1988; Crawford et al, 2006; Boyle et al, 2008). An extensive list of silencers was originally established from manually curated sites (Ogbourne and Antalis 1998), followed by the ORegAnno database that contains known enhancer and silencer elements in several species (Griffith et al, 2008). Importantly, certain silencer elements can exhibit contrasting reversibility in time, such as the CD4 silencer, with two distinct modes of function during thymopoiesis, by which gene silencing is permanent in CD8 cells via recruitment of chromatin remodelers (Taniuchi et al, 2002b).
A first class of silencers consists of silencer elements that are bound by bona fide constitutive repressors and co-repressors (Figure 1). Several TFs with transcriptionally inhibiting activity have already been described. The transcriptional repressor ZEB1, for example, is co-recruited with CtBP2 and HDAC1 and silences IL-2 expression in T cells (Wang et al, 2009). Several members of the Ets family of TFs also display prominent repressive activity, notably NERF (Gaspar et al, 2002) which lacks a transactivation domain (Cho et al, 2004), and Tel/Etv6 (De et al, 2014), Tel2/Etv7 (Gu et al, 2001), Elk-3 (Chung et al, 2005) that are also transcriptional repressors (Mavrothalassitis and Ghysdael 2000) due to structural differences (Poon and Kim 2017). REST/NRSF is a key neuronal repressor that binds the RE1 motif (Schoenherr and Anderson 1995). REST cobinds with LSD1 on the NRSE/RE1 sequence and also recruits Sin3a (Kuwahara et al, 2001; Rossbach 2011). The zinc-finger transcriptional transcriptional repressor Gfi1 was shown to bind intronic silencer elements (Zweidler-Mckay et al, 1996; Park et al, 2004). Gfi1 associates with ETO to repress transcription via HDAC recruitment (McGhee et al, 2003).
Importantly, another class of silencers entails recruitment of normally transactivating TFs (Figure 1). In fact, a number of transcription factors that normally recruit co-activators can also display repressor activity. A non-exhaustive list includes TAL1-mediated recruitment of Eto2 (Schuh et al, 2005), Runx1 and Groucho (Levanon et al, 1998) as well as Sin3A (Lutterbach et al, 2000), SIRT1 with LSD1 by NF-κB (Liu et al, 2011), However, in these cases, other activating TFs can also concurrently bind to the silencer (Steinke et al, 2014). Stat5 can also behave as a repressor (Luo and Yu-Lee 1997), whereby it is known to abolish Bcl6 expression by binding to an intronic silencer region located in this locus (Walker et al, 2007). Gata factors can have repressive activity too depending on the presence of Friend of Gata (FOG/FPM1) (Holmes et al, 1999). In some cases, specific residue phosphorylation can trigger a switch between activator and repressor activity, as for Runx1 Runx1, as well as changes in interacting partners (Elagib and Goldfarb 2007). Runx1 plays a major role in silencing CD4 expression by binding to the intronic silencer element (Sun et al, 1995; Hayashi et al, 2001; Taniuchi et al, 2002a). Other TFs with transactivating properties, such as Ets family members that are generally activators, can also recruit co-repressors depending on post-translational modifications (Cowley and Graves 2000; Tootle and Rebay 2005). One such example is the case of Erg-mediated repression (Yuan et al, 2009). Altogether, possible modes of silencer-mediated direct repression can entail i) binding of transcriptional repressors that directly interfere and compete with activators (in this case one CRE would function as an enhancer or silencer depending on the factor binding), ii) binding of transcriptional repressors to de novo DHSs which interact with the respective promoter and overwrite the activation caused by elsewhere located enhancers.
The relationship between lncRNAs, classical gene activation/ silencing and PcG-mediated repression?
However, lncRNAs also seem to be involved in classical silencer function. For example, the 4 kb Gal10 lncRNA was shown to be able to recruit the Rpd3S HDAC complex (Ma et al, 2013). Another example is the Ctbp1-AS lncRNA, which recruits Sin3A to repress transcription of Ctbp1 (Takayama et al, 2013). More recently, it was further shown that antigene RNAs (agRNAs) can inhibit gene transcription by targeting argonaute-dependent promoters (Janowski et al, 2005; Li et al, 2006; Janowski and Corey 2010; Younger and Corey 2011), thus potentially constituting another novel class of silencer elements (Figure 1). Altogether, this constitutes a parallel mode of co-repressor recruitment, which could turn out to be as widespread as lncRNA-mediatedPcG recruitment. In terms of gene regulation, it is generally understood that at least for antisense lncRNA, their activation prevents activation of neighboring genes by affecting the recruitment of the transcriptional machinery, while in a number of cases, the transcription of lncRNAs can induce local chromatin remodeling preventing the expression of neighboring genes. This latter model is however difficult to reconcile with the fact that linked lncRNAs and coding genes tend to co-occur within a same TAD, which usually results in synchronized expression patterns (Franke et al, 2016). However, characterization of the transcriptional regulation of lncRNAs remains largely unexplored, even if lncRNAs seem to follow classical gene activation rules (Mikkelsen et al, 2007). Current knowledge shows that lncRNA promoters tend to be non-CGI promoters (Chakraborty et al, 2014), and are thus likely to follow classical rules of gene activation. For example, the LET lncRNA was shown to be repressed by HDAC3 recruitment (Prabhakar et al, 2017). However, another report showed a class of lncRNAs loci that tends to be enriched in Polycomb binding (Young et al, 2012), while Xist itself appears to be regulated by DNA methylation (Goto and Monk 1998). A number of lncRNAs could however exhibit classical transcriptional activation rules, harboring enhancers, which may also be the case for those that are Polycomb regulated. The Cd8 locus, for example, a known Polycomb target (Harker et al, 2011), is also controlled by several enhancers (Sakaguchi et al, 2015). An example further arguing in this favor is the DHRS4-AS1 lncRNA, which is enhancer controlled (Yang et al, 2016). Above, we have described how a number of silencers can be bound by normally activating TFs. There might however be cases where activating TFs bind silencers, resulting not in repressive chromatin. These cases may correspond to activation of antisense lncRNA (Figure 1). How often are silencers found near lncRNAs? Are they two completely distinct modes of action? A core, legitimate question could thus be: are certain previously-described silencer elements actually enhancers or promoters of lncRNAs? While in many cases silencers are either bound by repressors bringing in co-repressors, there may exist a number of previously annotated silencers that do not exhibit such marks. At any rate, this question could be answered by revisiting ENCODE and HiC RNA-Seq datasets at distinct cell stages, to identify differential interactions in genes exhibiting a proximal lncRNA.
Another obvious question arises from the fact that a number of TFs seem to be shared by both PcG- and silencer-mediated repression. These TFs, described above, include Runx1, Gata3, and YY1 (Figure 1). Further examples notably include Bcl6, also shown to be involved in classical silencing (Deweindt et al, 1995; Lemercier et al, 2002; Takeda et al, 2003), as well as E2F6 (Trimarchi et al, 1998). This list may not be exhaustive. Further, it was also shown that H3K27me3 is also found at distal elements, including to describe a state of enhancer poising (Zentner et al, 2011; Calo and Wysocka 2013; King et al, 2016). Occasional spatial overlapping between PcG- and classical gene regulation is further substantiated by H3K27ac invasion at PRC2 targets in EZ-deficient conditions (Zenk et al, 2017). The Hoxa-d clusters are also prime examples of overlap of classical activation and PcG-mediated repression (Herz et al, 2012). Overlap of these functions was further shown to be dictated by higher-order chromatin organization (Narendra et al, 2015). Further, the balance between enhancer function and PcG-mediated repression may entail Ezh1 recruitment (Mousavi et al, 2012). Thus, a number of previously annotated silencers and enhancers may very well correspond to PREs depending in diverging cell type and activation states. Overlap between these modes of gene regulation may however be limited. Strong silencers were for example shown to be rich in CT motifs, while PREs are generally rich in CpG dinucleotides (Petrykowska et al, 2008). In any case, revisiting the role previously identified silencers may very well reveal that they are more than what they appear to be.
References
- Agulnik AI, Mitchell MJ, Mattei MG, et al. 1994. A novel X gene with a widely transcribed Y-linked homologue escapes X-inactivation in mouse and human. Hum Mol Genet, 3, 879-884
- Ai S, Peng Y, Li C, et al. 2017. EED orchestration of heart maturation through interaction with HDACs is H3K27me3-independent. Elife, 6
- Almeida M, Pintacuda G, Masui O, et al. 2017. PCGF3/5-PRC1 initiates Polycomb recruitment in X chromosome inactivation. Science, 356, 1081-1084
- Aranda S, Mas G, Di Croce L, et al. 2015. Regulation of gene transcription by Polycomb proteins. Sci Adv, 1, e1500737
- Armstrong L. 2012. Epigenetic control of embryonic stem cell differentiation. Stem Cell Rev, 8, 67-77
- Attwooll C, Oddi S, Cartwright P, et al. 2005. A novel repressive E2F6 complex containing the polycomb group protein, EPC1, that interacts with EZH2 in a proliferation-specific manner. J Biol Chem, 280, 1199-1208
- Augui S, Nora EP, Heard E, et al. 2011. Regulation of X-chromosome inactivation by the X-inactivation centre. Nat Rev Genet, 12, 429-442
- Bannister AJ and Kouzarides T. 2011. Regulation of chromatin by histone modifications. Cell Res, 21, 381-395
- Bannister AJ, Zegerman P, Partridge JF, et al. 2001. Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature, 410, 120-124
- Barlow DP and Bartolomei MS. 2014. Genomic imprinting in mammals. Cold Spring Harb Perspect Biol, 6
- Barnash KD, The J, Norris-Drouin JL, et al.2017. Discovery of peptidomimetic ligands of EED as allosteric inhibitors of PRC2. ACS Comb Sci, 19, 161-172
- Barski A, Cuddapah S, Cui K, et al. 2007. High-resolution profiling of histone methylations in the human genome. Cell, 129, 823-837
- Bauer M, Trupke J, Ringrose L, et al. 2016. The quest for mammalian Polycomb response elements: are we there yet? Chromosoma, 125, 471-496
- Bernstein BE, Mikkelsen TS, Xie X, et al. 2006a. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell, 125, 315-326
- Bernstein E, Duncan EM, Masui O, et al. 2006b. Mouse polycomb proteins bind differentially to methylated histone H3 and RNA and are enriched in facultative heterochromatin. Mol Cell Biol, 26, 2560-2569
- Betancur JG and Tomari Y. 2015. Cryptic RNA-binding by PRC2 components EZH2 and SUZ12. RNA Biol, 12, 959-965
- Blanco MR and Guttman M. 2017. Re-evaluating the foundations of lncRNA-Polycomb function. EMBO J, 36, 964-966
- Bohmdorfer G and Wierzbicki AT. 2015. Control of chromatin structure by long noncoding RNA. Trends Cell Biol, 25, 623-632
- Boros J, Arnoult N, Stroobant V, et al. 2014. Polycomb repressive complex 2 and H3K27me3 cooperate with H3K9 methylation to maintain heterochromatin protein 1alpha at chromatin. Mol Cell Biol, 34, 3662-3674
- Bose DA, Donahue G, Reinberg D, et al. 2017. RNA binding to CBP stimulates histone acetylation and Transcription. Cell, 168, 135-149
- Bowman SK, Deaton AM, Domingues H, et al. 2014. H3K27 modifications define segmental regulatory domains in the Drosophila bithorax complex. Elife, 3, e02833
- Boyer LA, Plath K, Zeitlinger J, et al. 2006. Polycomb complexes repress developmental regulators in murine embryonic stem cells. Nature, 441, 349-353
- Boyle AP, Davis S, Shulha HP,et al. 2008. High-resolution mapping and characterization of open chromatin across the genome. Cell, 132, 311-322
- Breiling A, Turner BM, Bianchi ME, et al. 2001. General transcription factors bind promoters repressed by Polycomb group proteins. Nature, 412, 651-655
- Brockdorff N. 2013. Noncoding RNA and Polycomb recruitment. RNA., 19, 429-442
- Buratowski S, Hahn S, Guarente L, et al. 1989. Five intermediate complexes in transcription initiation by RNA polymerase II. Cell, 56, 549-561
- Calo E and Wysocka J. 2013. Modification of enhancer chromatin: what, how, and why? Mol Cell, 49, 825-837
- Cao R, Tsukada Y, Zhang Y, et al. 2005. Role of Bmi-1 and Ring1A in H2A ubiquitylation and Hox gene silencing. Mol Cell, 20, 845-854
- Cao R, Wang L, Wang H, et al. 2002. Role of histone H3 lysine 27 methylation in Polycomb-group silencing. Science, 298, 1039-1043
- Caretti G, Di Padova M, Micales B, et al. 2004. The Polycomb Ezh2 methyl transferase regulates muscle gene expression and skeletal muscle differentiation. Genes Dev, 18, 2627-2638
- Chadwick BP and Willard HF. 2004. Multiple spatially distinct types of facultative heterochromatin on the human inactive X chromosome. Proc Natl Acad Sci USA,101, 17450-17455
- Chakraborty S, Deb A, Maji RK, et al. 2014. LncRBase: an enriched resource for lncRNA information. PloS one, 9, e108010
- Chaumeil J, Le Baccon P, Wutz A, et al. 2006. A novel role for Xist RNA in the formation of a repressive nuclear compartment into which genes are recruited when silenced. Genes Dev, 20, 2223-2237
- Cho JY, Akbarali Y, Zerbini LF, et al. 2004. Isoforms of the Ets transcription factor NERF/ELF-2 physically interact with AML1 and mediate opposing effects on AML1-mediated transcription of the B cell-specific blk gene. J Biol Chem, 279, 19512-19522
- Chu C, Qu K, Zhong FL, et al. 2011. Genomic maps of long noncoding RNA occupancy reveal principles of RNA-chromatin interactions. Mol Cell, 44, 667-678
- Chu C, Zhang QC, da Rocha ST, et al. 2015. Systematic discovery of Xist RNA binding proteins. Cell, 161, 404-416
- Chung SW, Chen YH, Perrella MA, et al. 2005. Role of Ets-2 in the regulation of heme oxygenase-1 by endotoxin. J Biol Chem, 280, 4578-4584
- Clark BS and Blackshaw S. 2014. Long non-coding RNA-dependent transcriptional regulation in neuronal development and disease. Front Genet, 5, 164
- Clouaire T and Stancheva I. 2008. Methyl-CpG binding proteins: specialized transcriptional repressors or structural components of chromatin? Cell Mol Life Sci, 65, 1509-1522
- Cooper S, Grijzenhout A, Underwood E, et al. 2016. Jarid2 binds mono-ubiquitylated H2A lysine 119 to mediate crosstalk between Polycomb complexes PRC1 and PRC2. Nat Commun, 7, 13661
- Cowley DO and Graves BJ. 2000. Phosphorylation represses Ets-1 DNA binding by reinforcing autoinhibition. Genes Dev,14, 366-376
- Crawford GE, Holt IE, Whittle J, et al. 2006. Genome-wide mapping of DNase hypersensitive sites using massively parallel signature sequencing (MPSS). Genome Res,16, 123-131
- de Napoles M, Mermoud JE, Wakao R, et al. 2004. Polycomb group proteins Ring1A/B link ubiquitylation of histone H2A to heritable gene silencing and X inactivation. Dev Cell, 7, 663-676
- De S, Chan AC, Coyne HJ, et al. 2014. Steric mechanism of auto-inhibitory regulation of specific and non-specific DNA binding by the ETS transcriptional repressor ETV6. J Mol Biol, 426, 1390-1406
- Denisenko O, Shnyreva M, Suzuki H, et al.1998. Point mutations in the WD40 domain of Eed block its interaction with Ezh2. Mol Cell Biol,18, 5634-5642
- Deweindt C, Albagli O, Bernardin F, et al.1995. The LAZ3/BCL6 oncogene encodes a sequence-specific transcriptional inhibitor: a novel function for the BTB/POZ domain as an autonomous repressing domain. Cell Growth Differ, 6, 1495-1503
- Di Ruscio A, Ebralidze AK, Benoukraf T, et al. 2013. DNMT1-interacting RNAs block gene-specific DNA methylation. Nature, 503, 371-376
- Dietrich N, Lerdrup M, Landt E, et al. 2012. REST-mediated recruitment of polycomb repressor complexes in mammalian cells. PLoS Genet,8, e1002494
- Ebralidze AK, Guibal FC, Steidl U, et al. 2008. PU.1 expression is modulated by the balance of functional sense and antisense RNAs regulated by a shared cis-regulatory element. Genes Dev, 22, 2085-2092
- Elagib KE and Goldfarb AN. 2007. Regulation of RUNX1 transcriptional function by GATA-1. Crit Rev Eukaryot Gene Expr,17, 271-280
- Engreitz JM, Pandya-Jones A, McDonel P, et al.2013. The Xist lncRNA exploits three-dimensional genome architecture to spread across the X chromosome. Science,341, 1237973
- Escamilla-Del-Arenal M, da Rocha ST, Spruijt CG, et al.2013. Cdyl, a new partner of the inactive X chromosome and potential reader of H3K27me3 and H3K9me2. Mol Cell Biol,33, 5005-5020
- Fenouil R, Cauchy P, Koch F, et al.2012. CpG islands and GC content dictate nucleosome depletion in a transcription-independent manner at mammalian promoters. Genome Res, 22, 2399-2408
- Ferrari KJ, Scelfo A, Jammula S,et al. 2014. Polycomb-dependent H3K27me1 and H3K27me2 regulate active transcription and enhancer fidelity. Mol Cell, 53, 49-62
- Francis NJ, Kingston RE, Woodcock CL, et al. 2004. Chromatin compaction by a polycomb group protein complex. Science, 306, 1574-1577
- Franke M, Ibrahim DM, Andrey G, et al. 2016. Formation of new chromatin domains determines pathogenicity of genomic duplications. Nature, 538, 265-269
- Fujita N, Watanabe S, Ichimura T, et al. 2003. Methyl-CpG binding domain 1 (MBD1) interacts with the Suv39h1-HP1 heterochromatic complex for DNA methylation-based transcriptional repression. J Biol Chem, 278, 24132-24138
- Furuyama T, Banerjee R, Breen TR, et al. 2004. SIR2 is required for polycomb silencing and is associated with an E(Z) histone methyltransferase complex. Curr Biol, 14, 1812-1821
- Gao JZ, Li J, Du JL, et al. 2016. Long non-coding RNA HOTAIR is a marker for hepatocellular carcinoma progression and tumor recurrence. Oncol Lett, 11, 1791-1798
- Gao Z, Zhang J, Bonasio R, et al. 2012. PCGF homologs, CBX proteins, and RYBP define functionally distinct PRC1 family complexes. Mol Cell, 45, 344-356
- Gaspar J, Thai S, Voland C, et al. 2002. Opposing functions of the Ets factors NERF and ELF-1 during chicken blood vessel development. Arterioscler Thromb Vasc Biol, 22, 1106-1112
- Geisler S and Coller J. 2013. RNA in unexpected places: long non-coding RNA functions in diverse cellular contexts. Nat Rev Mol Cell Biol, 14, 699-712
- Glazko GV, Zybailov BL, Rogozin IB, et al. 2012. Computational prediction of polycomb-associated long non-coding RNAs. PloS one, 7, e44878
- Gong C and Maquat LE. 2011. lncRNAs transactivate STAU1-mediated mRNA decay by duplexing with 3' UTRs via Alu elements. Nature, 470, 284-288
- Goode DK, Obier N, Vijayabaskar MS, et al.2016. Dynamic Gene Regulatory Networks Drive Hematopoietic Specification and Differentiation. Dev Cell, 36, 572-587
- Goodrich JA and Kugel JF. 2006. Non-coding-RNA regulators of RNA polymerase II transcription. Nat Rev Mol Cell Biol, 7, 612-616
- Goto T and Monk M. 1998. Regulation of X-chromosome inactivation in development in mice and humans. Microbiol Mol Biol Rev, 62, 362-378
- Griffith OL, Montgomery SB, Bernier B, et al. 2008. ORegAnno: an open-access community-driven resource for regulatory annotation. Nucleic Acids Res, 36, D107-113
- Gross DS and Garrard WT. 1988. Nuclease hypersensitive sites in chromatin. Annu Rev Biochem, 57, 159-197
- Grote P, Wittler L, Hendrix D, et al. 2013. The tissue-specific lncRNA Fendrr is an essential regulator of heart and body wall development in the mouse. Dev Cell, 24, 206-214
- Gu X, Shin BH, Akbarali Y, et al. 2001. Tel-2 is a novel transcriptional repressor related to the Ets factor Tel/ETV-6. J Biol Chem, 276, 9421-9436
- Guil S, Soler M, Portela A, et al. 2012. Intronic RNAs mediate EZH2 regulation of epigenetic targets. Nat Struct Mol Biol, 19, 664-670
- Harker N, Garefalaki A, Menzel U, et al. 2011. Pre-TCR signaling and CD8 gene bivalent chromatin resolution during thymocyte development. J Immunol, 186, 6368-6377
- Hayashi K, Abe N, Watanabe T, et al. 2001. Overexpression of AML1 transcription factor drives thymocytes into the CD8 single-positive lineage. J Immunol, 167, 4957-4965
- He Y, Selvaraju S, Curtin ML, et al. 2017. The EED protein-protein interaction inhibitor A-395 inactivates the PRC2 complex. Nat Chem Biol, 13, 389-395
- Chadwick LH, Chadwick BP, Jaye DL, et al. 2009. The Mi-2/NuRD complex associates with pericentromeric heterochromatin during S phase in rapidly proliferating lymphoid cells. Chromosoma, 118, 445-457
- Hendrickson DG, Kelley DR, Tenen D, et al. 2016. Widespread RNA binding by chromatin-associated proteins. Genome Biol, 17, 28
- Herz HM, Mohan M, Garruss AS, et al. 2012. Enhancer-associated H3K4 monomethylation by Trithorax-related, the Drosophila homolog of mammalian Mll3/Mll4. Genes Dev, 26, 2604-2620
- Herzog VA, Lempradl A, Trupke J, et al. 2014. A strand-specific switch in noncoding transcription switches the function of a Polycomb/Trithorax response element. Nat Genet, 46, 973-981
- Hodgson JW and Brock HW. 2011. Are polycomb group bodies gene silencing factories? Cell,144, 170-171
- Holmes M, Turner J, Fox A, et al. 1999. hFOG-2, a novel zinc finger protein, binds the co-repressor mCtBP2 and modulates GATA-mediated activation. J Biol Chem, 274, 23491-23498
- Hosokawa H, Kimura MY, Shinnakasu R, et al. 2006. Regulation of Th2 cell development by Polycomb group gene bmi-1 through the stabilization of GATA3. J Immunol, 177, 7656-7664
- Hu G, Gupta SK, Troska TP, et al. 2017. Long non-coding RNA profile in mantle cell lymphoma identifies a functional lncRNA ROR1-AS1 associated with EZH2/PRC2 complex. Oncotarget, 8, 802-823
- Huynh KD, Fischle W, Verdin E, et al. 2000. BCoR, a novel corepressor involved in BCL-6 repression. Genes Dev, 14, 1810-1823
- Illingworth RS, Botting CH, Grimes GR, et al. 2012. PRC1 and PRC2 are not required for targeting of H2A.Z to developmental genes in embryonic stem cells. PloS one, 7, e34848
- Janowski BA and Corey DR. 2010. Minireview: Switching on progesterone receptor expression with duplex RNA. Mol Endocrinol, 24, 2243-2252
- Janowski BA, Huffman KE, Schwartz JC, et al. 2005. Inhibiting gene expression at transcription start sites in chromosomal DNA with antigene RNAs. Nat Chem Biol, 1, 216-222
- Jiang CL, Jin SG, Pfeifer GP, et al. 2004. MBD3L1 is a transcriptional repressor that interacts with methyl-CpG-binding protein 2 (MBD2) and components of the NuRD complex. J Biol Chem, 279, 52456-52464
- Kahn TG, Dorafshan E, Schultheis D, et al. 2016. Interdependence of PRC1 and PRC2 for recruitment to Polycomb Response Elements. Nucleic Acids Res, 44, 10132-10149
- Kaneko S, Bonasio R, Saldana-Meyer R, et al. 2014. Interactions between JARID2 and noncoding RNAs regulate PRC2 recruitment to chromatin. Mol Cell, 53, 290-300
- Kashyap V, Rezende NC, Scotland KB, et al. 2009. Regulation of stem cell pluripotency and differentiation involves a mutual regulatory circuit of the NANOG, OCT4, and SOX2 pluripotency transcription factors with polycomb repressive complexes and stem cell microRNAs. Stem Cells Dev, 18, 1093-1108
- Katoh-Fukui Y, Miyabayashi K, Komatsu T, et al. 2012. Cbx2, a polycomb group gene, is required for Sry gene expression in mice. Endocrinology, 153, 913-924
- Katoh-Fukui Y, Tsuchiya R, Shiroishi T, et al. 1998. Male-to-female sex reversal in M33 mutant mice. Nature, 393, 688-692
- Kaustov L, Ouyang H, Amaya M, et al. 2011. Recognition and specificity determinants of the human cbx chromodomains. J Biol Chem, 286, 521-529
- Khachane AN and Harrison PM. 2010. Mining mammalian transcript data for functional long non-coding RNAs. PLoS One, 5, e10316
- King AD, Huang K, Rubbi L, et al. 2016. Reversible regulation of promoter and enhancer histone landscape by DNA methylation in mouse embryonic stem cells. Cell Rep, 17, 289-302
- Klose RJ, Kallin EM, Zhang Y, et al. 2006. JmjC-domain-containing proteins and histone demethylation. Nat Rev Genet, 7, 715-727
- Koch F, Jourquin F, Ferrier P, et al. 2008. Genome-wide RNA polymerase II: not genes only! Trends Biochem Sci, 33, 265-273
- Kolovos P, Knoch TA, Grosveld FG, et al. 2012. Enhancers and silencers: an integrated and simple model for their function. Epigenetics Chromatin, 5, 1
- Kortschak RD, Tucker PW, Saint R, et al. 2000. ARID proteins come in from the desert. Trends Biochem Sci, 25, 294-299
- Kotake Y, Nakagawa T, Kitagawa K, et al. 2011. Long non-coding RNA ANRIL is required for the PRC2 recruitment to and silencing of p15(INK4B) tumor suppressor gene. Oncogene, 30, 1956-1962
- Ku M, Koche RP, Rheinbay E, et al. 2008. Genomewide analysis of PRC1 and PRC2 occupancy identifies two classes of bivalent domains. PLoS Genet, 4, e1000242
- Kung JT, Colognori D, Lee JT, et al. 2013. Long noncoding RNAs: past, present, and future. Genetics, 193, 651-669
- Kuwahara K, Saito Y, Ogawa E, Takahash