Review Article - Journal of RNA and Genomics (2017) Volume 13, Issue 1
Intramyocardial Gene Silencing by Interfering RNA
María Soledad Brea, Patricio Eduardo Morgan and Néstor Gustavo Pérez*
Centro de Investigaciones Cardiovasculares, Facultad de Ciencias Médicas, Universidad Nacional de La Plata, Argentina
Received date: 13 May 2017, Revised date: 02 June 2017, Accepted date: 05 June 2017, Published date: 12 June 2017
© Copyright: The Author(s). First Published by Allied Academies. This is an open access article, published under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0). This license permits non-commercial use, distribution and reproduction of the article, provided the original work is appropriately acknowledged with correct citation details
Abstract
RNAi is a widely used methodology for gene silencing. The action mechanism of siRNA molecules has been well studied in recent years, and the technique has been optimized in terms of safety and effectiveness. Cardiovascular diseases have a high incidence in the current population, and despite of the extensive research, safe and efficient therapeutics have not yet been found, which is reflected by 17.1 million people who die each year for this cause. In this context, siRNAs are being considered a therapeutic tool to regulate the expression of genes involved in the generation of these pathologies. The efficacy of siRNAs entry to cardiomyocytes, the safety of the delivery process and the degree of silencing achieved are main aspects before consider it as a cardiovascular disease therapy. Presently, we will give a brief outline of the current understanding of the RNAi mechanism and the delivery system to the heart. We describe the use of lentivirus for a functional silencing of cardiac proteins in the study of a pathophysiological process, the slow force response to cardiac stretch.
Keywords
siRNA, Heart, Lentivirus, Cardiomyocyte, Slow force response
Abbreviations
RNAi: RNA interference; siRNA: small interfering RNA; dsRNA: double-stranded RNA; shRNA: short hairpin RNA; RISC: RNA-Induced Silencing Complex; SFR: Slow Force Response; CH: Cardiac Hypertrophy; NHE: Na+/H+ Exchanger; MR: Mineralocorticoid Receptor; EGFR: Epidermal Growth Factor Receptor; TRH: Thyrotropin Releasing Hormone
Introduction
Understanding of RNA biology has deeply expanded during last years. Since new RNA structures with different functions were discovered, it is now possible to conceive it as a structure with multiples signaling functions. These advances allowed using it as an experimental tool or even in the therapeutic field of clinical medicine. In fact, ribozymes, antisense RNA, aptamers, micro RNA (miRNA) or small interfering RNA (siRNA) are all used to decrease protein expression and characterize its function in gain or loss strategy or ultimately test its efficacy as novel therapy.
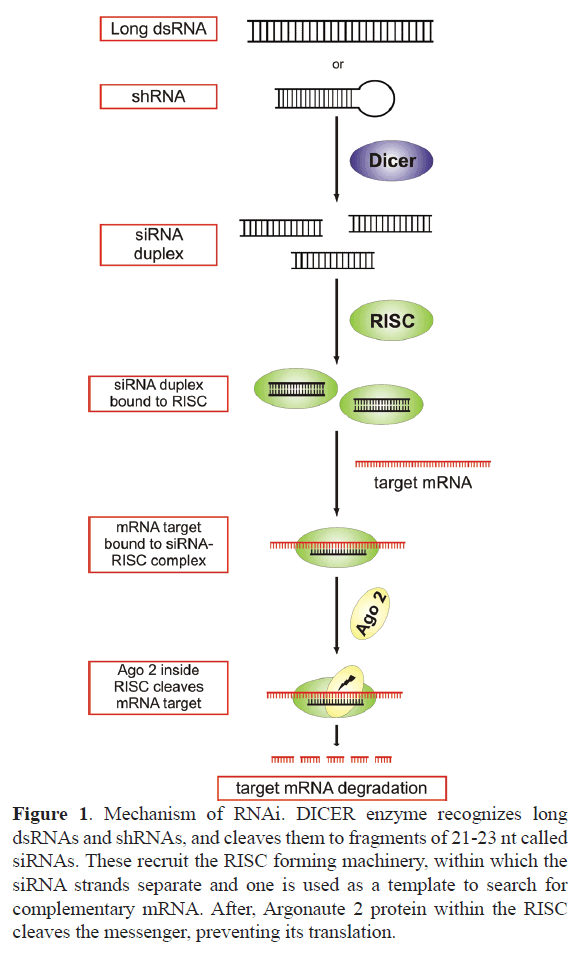
Almost 20 years ago, it was demonstrated that introduction of double strand RNA (dsRNA) into eukaryote cells triggered a sequence-specific gene silencing (Fire A, 1998). The dsRNA is processed by pre-existing intracellular machinery, along a pathway that forms the interfering RNA which mediates mRNA degradation. Briefly, in the cytosol, the dsRNA is cleaved by DICER endoribonuclease into small fragments of 21-23 nt, called siRNA. These recruit Argonaute-2 and other proteins forming the RISC complex (RNA-induced silencing complex). Within the complex, RNA strands are separated, being one of them a template to find an mRNA with a complementary sequence (Tang, 2005). Once the matching mRNA is found, Argonaute-2 cleaves it (Grimm, 2009) and prevents its translation (Figure 1). Detailed description of the mechanism of functioning of these RNA molecules can be found in the works of Ipsaro and Joshua- Tor (Ipsaro and Joshua-Tor, 2015) and Kobayashi and Tomari (Kobayashi and Tomari, 2016).
Figure 1. Mechanism of RNAi. DICER enzyme recognizes long dsRNAs and shRNAs, and cleaves them to fragments of 21-23 nt called siRNAs. These recruit the RISC forming machinery, within which the siRNA strands separate and one is used as a template to search for complementary mRNA. After, Argonaute 2 protein within the RISC cleaves the messenger, preventing its translation.
RNAi technique offers several advantages to modulate gene expression in whole animals compared to the widely used knockout models, to decrease protein function. Working with siRNAs is simpler and cheaper than generation and maintenance of knockout models, albeit some genes cannot be knocked-out. Despite these advantages, RNA silencing involves certain general challenges, as efficient siRNA delivery into the target cell, enough expression levels within the cell and cell specificity. The latter is important to avoid siRNA expression into other organs, and even more important when the objective is to target a specific cell within a desired organ (Whitehead et al, 2009). Finally, the intervention to express the siRNA must be safe, avoiding collateral alterations within the cell or in the surrounding tissue induced by immune response.
Exogenous siRNA could be express in vivo using different strategies. The most direct way is infusing synthetic siRNA, conjugated with particles that stabilize its structure, preventing early degradation in the tissue (Chira et al, 2015, Jasinski et al, 2017). siRNA could be stabilized by conjugation with cholesterol or with different kind of nanoparticles as aptamers, micelles, liposomes, cationic polymers. This is a factor to consider not only for a systemic but local administration of siRNA, since will improve the rate of cellular uptake, prevent immune system activation, avoid renal elimination, and confers pH and thermal stability, enhancing siRNA delivery efficiency (Tatiparti et al, 2017).
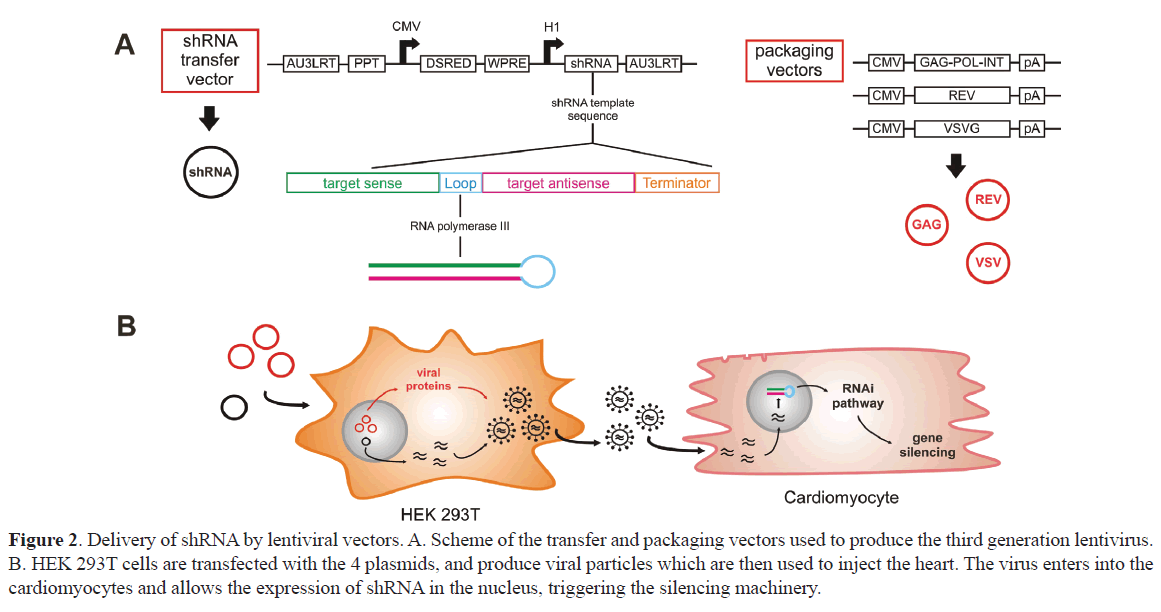
Other options consider to generate a short hairpin RNA (shRNA), introduced in the cells with viral vectors. shRNA is synthesized from a DNA cassette consisting of two sections of complementary sequences joined by a loop sequence. This cassette is transcribed by RNA polymerase III from the host cell, and its expression is dependent on a promoter like U6, H1 or 7SK (Figure 2). The hairpin structure is recognized by the enzyme DICER, which cleaves it generating a siRNA duplex (McIntyre and Fanning, 2006). The following sections of this review will focus on cardiac delivery of siRNA and a brief discussion of lentiviral vectors to express shRNA in vivo in the heart, during pathophysiological processes.
Figure 2. Delivery of shRNA by lentiviral vectors. A. Scheme of the transfer and packaging vectors used to produce the third generation lentivirus. B. HEK 293T cells are transfected with the 4 plasmids, and produce viral particles which are then used to inject the heart. The virus enters into the cardiomyocytes and allows the expression of shRNA in the nucleus, triggering the silencing machinery.
Cardiac Sirna Delivery
Transient siRNA expression
Cardiomyocytes are approximately 75% of the heart’s volume, but less than 60% of the total cells in the heart. Other cell types like fibroblasts and vessel cells constitute anatomic barriers that prevent proper siRNA delivery. Furthermore, non-myocyte cells could be also potential targets to compete with cardiomyocytes for siRNA transfection or vector transduction. The correct choice of a siRNA delivery route to the heart is a key issue to consider to a successful cardiomyocyte expression. If the intention is a silencing effect restricted to the heart, intravenous delivery of naked siRNA is certainly not an option, since it would be distributed throughout the organism. Several studies have shown that intraperitoneal injection, subcutaneous delivery through pumps or direct infusion by the blood stream of siRNA molecules not only had effects on the heart but also affect other organs such as lung and kidney (Arnold et al, 2007, Clemente et al, 2007, Wang et al, 2009). This inconvenience could be avoided by directly injecting the siRNA in vivo into the myocardial wall. We found that direct intramyocardial injection of naked siRNA in the left ventricle to silence the membrane Na+/H+ exchanger (NHE1) produced a reduction of cardiac protein expression without affecting other organs (Morgan et al, 2011). Similarly, other authors found that this delivery strategy to target other cardiac proteins such as epidermal growth factor receptor (EGFR) (Feng et al, 2012) and thyrotropin releasing hormone (TRH) (Schuman et al, 2011) led to protein expression reduction with functional consequences.
It is important to highlight that naked siRNA without any stabilizing modification is rapidly degraded outside the cell, in the tissue. With this strategy, the silencing effect is greater between 48-72 hours, achieving an expression reduction of approximately 55-70% (Morgan et al, 2011, Feng et al, 2012). The broad silencing of protein expression along the left ventricle could result from the diffusion of siRNA molecules from the injection site in the apex along the myocardium. Also it could be possible that siRNA molecules travel from one cell to another through gap junctions containing connexin 43 (Kizana et al, 2009). The low stability of naked siRNA and the specific protein turnover rate, determines that the effect vanishes around a week later. Therefore, a potential chronic treatment would need periodical repeat of the procedure, although the reversibility of the effect could be considered as an advantage for temporary inhibition. siRNA conjugation with different kind of nanoparticles, although widely used in cancer research and as a potential therapeutic tool was little utilized in the heart field. Recently, cholesterol conjugated siRNA was injected into the heart wall to decrease the expression of renalase (Huang et al, 2016). After 24 h, protein expression was ~75% decreased and the siRNA fluorescence tag signal detected, indicative of prolonged stability.
The use of siRNA in vitro showed in some cases activation of the interferon system (Samuel-Abraham and Leonard, 2010), which could mask a specific response. Delivery of siRNA through the hydrodynamic pulse achieved a reduction of the protein without affecting similar proteins or inducing an inflammatory response in the heart (Arnold et al, 2007, Clemente et al, 2007). From the clinical perspective, the highly invasive nature of this procedure force to consider that the potential therapeutic application of the technique is still too far.
Long term siRNA expression
Due to the low stability of the naked siRNA, long term silencing could be obtained with viral vectors that transduce the target cell with the coding sequence of a specific shRNA. The three most commonly viral systems used are adenovirus (Wu et al, 2014, Gao et al, 2017), lentivirus (Zhang et al., 2015, Brea et al, 2016) and adeno-associated virus (Suckau et al, 2009, Miyazaki et al, 2012). These vectors have the advantage of possessing a better efficiency for siRNA delivering and with adequate modifications, minimize an immune response that could compromise a correct delivery or mask the siRNA effect through off-target effects (Merentie et al, 2016). Some viral vectors have the possibility of being intravenously injected, since though the vectors are distributed throughout the whole organism, the shRNA promoter can only be expressed in the target organ. In addition, the vector could also have a special tropism towards the heart. In other words, despite being distributed throughout the organism, only the heart will be transduced.
Lentivirus, which does not have cardiac tropism, could limit their expression to the heart by an intraventricular injection. Lentiviral vectors derived from human immunodeficiency virus type 1 (HIV-1) are capable of transducing a wide variety of dividing and terminals cells, where they are stably integrated into the host genome, allowing prolonged expression of the transgene. The general strategy used to produce a recombinant lentiviral vector has been to eliminate elements of the virus genome limiting its replication and pathogenicity. This is crucial to avoid possible recombination and immune response (Tiscornia et al, 2003).
Third generation lentiviral vectors used by us were generated with four plasmids (Figure 2). The transfer vector contains the transgene of interest, and all cis-elements required for RNA production and packaging. The packaging system involves three additional plasmids, which provide the required trans-factors, called Gag-Pol, envelope protein and Rev, respectively. Gag-Pol encodes for an integrase, a reverse transcriptase, and structural proteins. Structural proteins are required for particle production, whereas integrase and reverse transcriptase are packaged within the viral particle, becoming active during the process of infection and integration. Rev interacts with the response element Rev (RRE), which is a sequence contained in the transfer vector that increases the export of viral genomic RNA from the nucleus, favoring the increase of the viral titer (Tiscornia et al, 2003). Vesicular Stomatitis Virus G-protein (VSV-G) incorporated in the envelope of the viral particles confer them the ability to transduce a variety of cell types (Figure 2).
Studying Slow Force Response by Gene Silencing
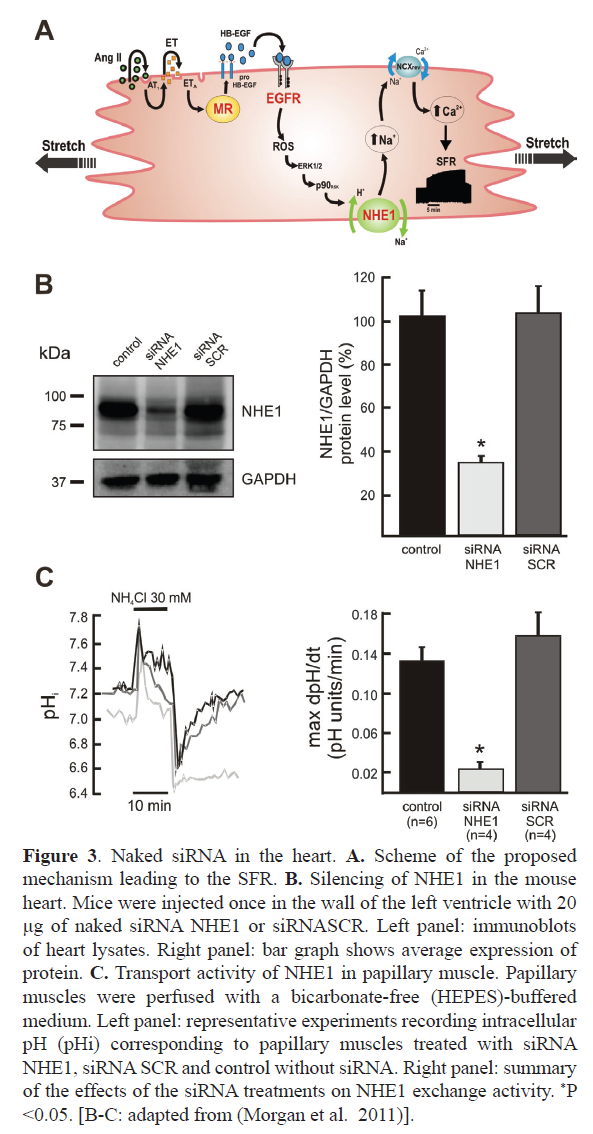
Hemodynamic increase induce a cardiac output adaptation that activates intracellular pathways, that if chronically sustained lead to pathological consequences as cardiac hypertrophy (CH) and heart failure. Acute myocardial stretch, originates an initial rise in contractility, followed by a slowly continuous force increase. This slow force response (SFR) is originated by a stretch-triggered activation of Angiotensin II receptor 1a (AT1R), as in CH. A proposed signaling pathway for the SFR includes a sequence of: 1) Stretch-triggered release of Angiotensin II/AT1-R activation, 2) Release/formation of endothelin, 3) activation of the mineralocorticoid receptor (MR), 4) transactivation of EGFR, 5) increased formation of mitochondrial reactive oxygen species (ROS), 6) activation of redox-sensitive kinases, 7) NHE1 activation, 8) increase in intracellular Na+ concentration, and 9) increase in Ca2+ transient amplitude through the Na+/Ca2+ exchanger (Figure 3A). Experimental evidence indicates that a good option to treat CH and heart failure would be preventing NHE1 or MR activation (Pitt et al, 2001, Zannad et al, 2011, Karmazyn, 2013), two proteins that in addition to EGFR were target for gene silencing. Following we discuss results of siRNA use against NHE1, MR and EGFR.
Figure 3. Naked siRNA in the heart. A. Scheme of the proposed mechanism leading to the SFR. B. Silencing of NHE1 in the mouse heart. Mice were injected once in the wall of the left ventricle with 20 μg of naked siRNA NHE1 or siRNASCR. Left panel: immunoblots of heart lysates. Right panel: bar graph shows average expression of protein. C. Transport activity of NHE1 in papillary muscle. Papillary muscles were perfused with a bicarbonate-free (HEPES)-buffered medium. Left panel: representative experiments recording intracellular pH (pHi) corresponding to papillary muscles treated with siRNA NHE1, siRNA SCR and control without siRNA. Right panel: summary of the effects of the siRNA treatments on NHE1 exchange activity. *P <0.05. [B-C: adapted from (Morgan et al. 2011)].
The NHE1 is an integral protein of the cardiomyocyte plasma membrane that removes H+ from the cytosol in exchange for Na+. Hyperactivity of cardiac NHE1 induces SFR, CH and heart failure. Pharmacological inhibition of NHE1 has been shown to be beneficial in animal models of cardiac pathologies. However, clinical trials with NHE1 inhibitors failed to yield positive results or worst were associated to undesired side effects probably due to a tissue unspecific NHE1 inhibition.
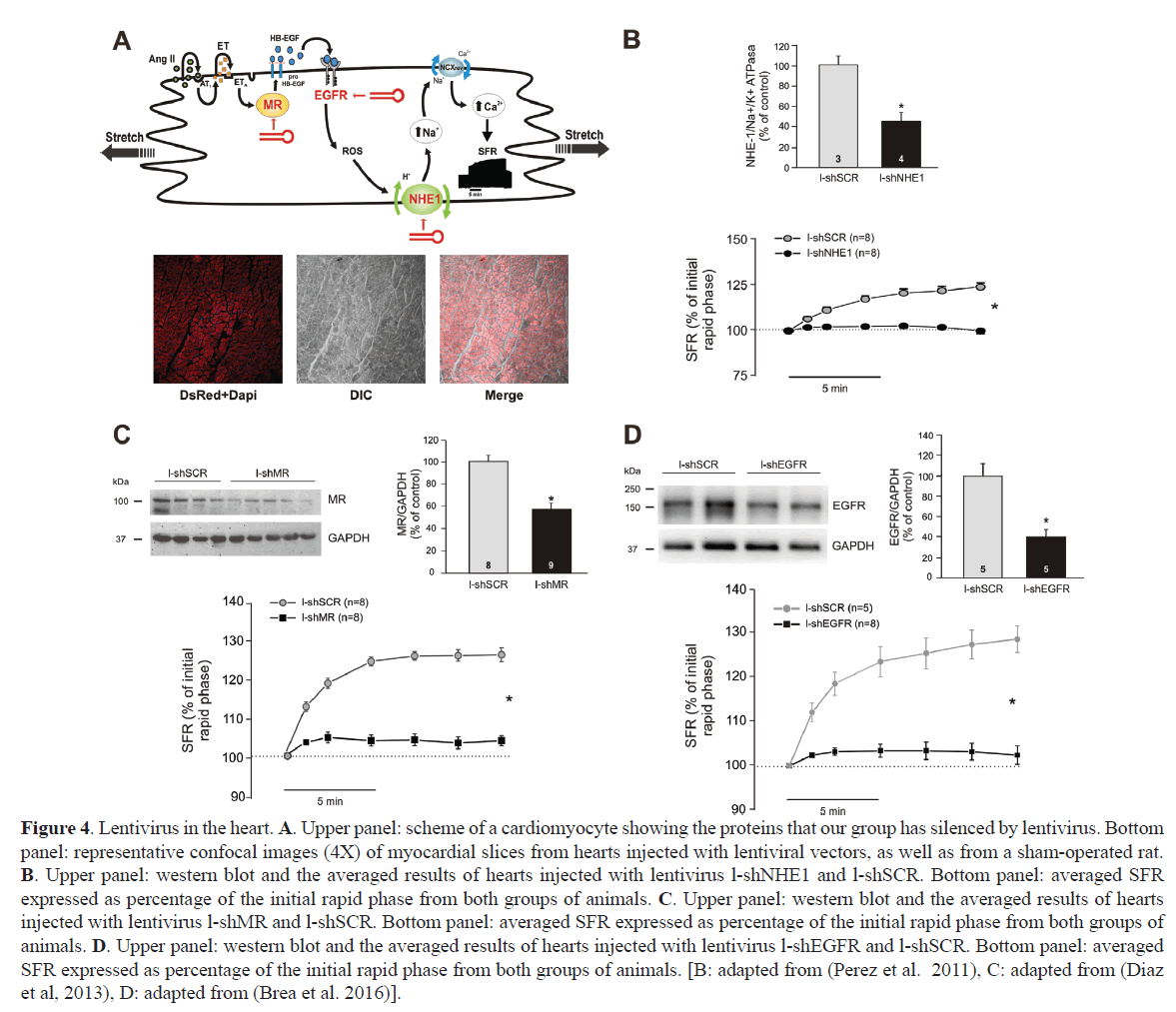
Direct injection of naked siRNA into the mouse heart wall, temporarily decreases NHE1 expression, producing functional silencing (Figure 3B and 3C) (Morgan et al, 2011). NHE1 dependent proton efflux after an acute acidic challenge was minimized. Protein expression reduction was not even throughout the entire myocardium, rather, it gets reduced from the injection site. Expression in liver, lung or brain was not changed, indicating that the silencing was limited to the injected organ. For a long-term silencing, the same siRNA sequence was incorporated into a third-generation lentivirus. A lentivirus carrying a shRNA-NHE1 and a reporter protein was injected at two sites in the free wall of rat heart left ventricle (Perez et al, 2011). A month later, red fluorescence staining of the reporter protein was observed by confocal microscopy in both the apex and base of the heart, as well as in the papillary muscles, where ex vivo, the SFR was studied (Figure 4A). A significant reduction of NHE1 protein expression together with cancellation of the SFR was observed (Figure 4B). The same combination of lentivirus and intramyocardial injection was used to verify the effect of NHE1 silencing in the hypertrophic myocardium of SHR rats. Long term NHE1 silencing affected ~50% of the protein expression, reversing the pathologic CH and producing a significant reduction in cardiac parietal stress (Nolly et al, 2015). As with naked siRNA, the silencing effect was specific to the heart, since NHE1 expression was not modified in lung or liver.
Figure 4. Lentivirus in the heart. A. Upper panel: scheme of a cardiomyocyte showing the proteins that our group has silenced by lentivirus. Bottom panel: representative confocal images (4X) of myocardial slices from hearts injected with lentiviral vectors, as well as from a sham-operated rat. B. Upper panel: western blot and the averaged results of hearts injected with lentivirus l-shNHE1 and l-shSCR. Bottom panel: averaged SFR expressed as percentage of the initial rapid phase from both groups of animals. C. Upper panel: western blot and the averaged results of hearts injected with lentivirus l-shMR and l-shSCR. Bottom panel: averaged SFR expressed as percentage of the initial rapid phase from both groups of animals. D. Upper panel: western blot and the averaged results of hearts injected with lentivirus l-shEGFR and l-shSCR. Bottom panel: averaged SFR expressed as percentage of the initial rapid phase from both groups of animals. [B: adapted from (Perez et al. 2011), C: adapted from (Diaz et al, 2013), D: adapted from (Brea et al. 2016)].
Mineralocorticoid receptor (MR) is overexpressed in the failing heart where the clinical efficacy of MR inhibitors is well established (Pitt et al, 1999, Pitt et al, 2001, Zannad et al, 2011). However, the exact mechanisms by which MR antagonists provide cardiovascular protection in patients with heart failure are not completely understood. Its participation in SFR had been demonstrated using the inhibitors spironolactone and eplerenone (Caldiz et al, 2011). Since those are non-selective compounds and have been described as inverse agonist of the MR, siRNA was used as a specific inhibitory strategy. shRNA-MR incorporated into a lentivirus was injected into the left ventricle wall, reducing MR expression at both cardiac wall and papillary muscle (Figure 4C). The functional consequence of MR silencing was the cancellation of the SFR (Figure 4C) (Diaz et al, 2013). Alternatively, lentiviruses expressing shRNA-MR were injected into mice oocytes to generate inducible MR knock-down animals, allowing shRNA temporary expression control (Montes-Cobos et al, 2015). shRNA was expressed in all tissues analyzed and produced after 15 days, at least in heart and kidney a ~50% MR reduction, preventing CH.
EGFR belongs to a tyrosine kinase family involved in numerous biological processes. It was extensively studied in cancer, diabetes and cardiovascular diseases (Hervent and De Keulenaer, 2012, Forrester et al, 2016). Different siRNA delivery strategies were used to specifically silence this receptor in the heart. In our hands, lentivirus expressing shRNA-EGFR was injected in the cardiac left ventricle wall of rats (Brea et al, 2016). Expression of EGFR decreased by 60%, without affecting other family member receptors like ErbB2 or ErbB4 (Figure 4D). As for NHE1 or MR silencing, reduced protein expression correlated with a minimized SFR in papillary muscles expressing shRNA-EGFR (Figure 4D). Other authors have shown that siRNA duplex injected directly into the cardiac wall produces 45% decrease of EGFR expression after 48 hours (Feng et al, 2012). The functional consequence was a reduction of the incidence of ventricular fibrillation induced after reperfusion. Nevertheless, there is no evaluation of silencing of other family receptors or of EGFR in other organs.
Conclusion
The RNAi technique allows studying a protein function specifically localized in the heart. Different strategies have been used to express siRNA in cardiomyocytes including administration of naked RNA duplexes or conjugated to nanoparticles to improve stability, safety and cellular uptake. Viral vectors such as adenoviruses, adeno-associated viruses and lentiviruses have effectively been used to inhibit long-term protein expression and study its functional consequences. The gene silencing results obtained in animals are encouraging to consider it as an alternative therapeutic tool. The scarce clinical trials with interference RNA are directed to treat different pathologies that do not include the heart as target organ. Heart access presents serious difficulties to obtain an efficient and specific silencing minimizing a potential immune response. Intramyocardial injection presented here with small animals, although highly invasive, may probably be considered as an option during a simple catheterism intervention in higher animals.
Source of Funding
This work was supported in part by grant PICT 2012-2396 from Agencia Nacional de Promoción Científica of Argentina and PIP0750 from CONICET to Dr. NG Pérez.
Competing Interests
The authors have no competing interests.
References
- Fire A. 1998. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature. 391, 806-811
- Tang G. 2005. siRNA and miRNA: an insight into RISCs. Trends Biochem Sci. 30, 106-114.
- Grimm D. 2009. Small silencing RNAs: state-of-the-art. Adv Drug Deliv Rev. 61, 672-703.
- Ipsaro JJ and Joshua-Tor L. 2015. From guide to target: molecular insights into eukaryotic RNA-interference machinery. Nat Struct Mol Biol. 22, 20-28
- Kobayashi H and Tomari Y. 2016. RISC assembly: Coordination between small RNAs and Argonaute proteins. Biochim Biophys Acta. 1859, 71-81
- Whitehead KA, Langer R and Anderson DG. 2009. Knocking down barriers: advances in siRNA delivery. Nat Rev Drug Discov. 8, 129-138
- Chira S, Jackson CS, Oprea I, et al. 2015. Progresses towards safe and efficient gene therapy vectors. Oncotarget. 6, 30675-30703
- Jasinski D, Haque F, Binzel DW, et al. 2017. Advancement of the Emerging Field of RNA Nanotechnology. ACS Nano. 11, 1142-1164
- Tatiparti K, Sau S, Kashaw SK, et al. 2017. siRNA Delivery Strategies: A Comprehensive Review of Recent Developments. Nanomaterials. 7, 4
- McIntyre GJ and Fanning GC. 2006. Design and cloning strategies for constructing shRNA expression vectors. BMC Biotechnol. 6, 1
- Arnold AS, Tang YL, Qian K, et al. 2007. Specific beta1-adrenergic receptor silencing with small interfering RNA lowers high blood pressure and improves cardiac function in myocardial ischemia. J Hypertens. 25, 197-205
- Clemente CF, Tornatore TF, Theizen TH, et al. 2007. Targeting focal adhesion kinase with small interfering RNA prevents and reverses load-induced cardiac hypertrophy in mice. Circ Res. 101, 1339-1348
- Wang X, Oka T, Chow FL, et al. 2009. Tumor necrosis factor-alpha-converting enzyme is a key regulator of agonist-induced cardiac hypertrophy and fibrosis. Hypertension. 54, 575-582
- Morgan PE, Correa MV, Ennis IL, et al. 2011. Silencing of sodium/hydrogen exchanger in the heart by direct injection of naked siRNA. J Appl Physiol. 111, 566-572
- Feng M, Xiang JZ, Ming ZY, et al. 2012. Activation of epidermal growth factor receptor mediates reperfusion arrhythmias in anaesthetized rats. Cardiovasc Res. 93, 60-68
- Schuman ML, Landa MS, Toblli JE, et al. 2011. Cardiac thyrotropin-releasing hormone mediates left ventricular hypertrophy in spontaneously hypertensive rats. Hypertension. 57, 103-109
- Kizana E, Cingolani E and Marban E. 2009. Non-cell-autonomous effects of vector-expressed regulatory RNAs in mammalian heart cells. Gene Ther. 16, 1163-1168
- Huang K, Liu J, Zhang H, et al. 2016. Intramyocardial injection of siRNAs can efficiently establish myocardial tissue-specific renalase knockdown mouse model. Biomed Res Int. 2016, 1267570
- Samuel-Abraham S and Leonard JN. 2010. Staying on message: design principles for controlling nonspecific responses to siRNA. FEBS J. 277, 4828-4836
- Wu W, Hu Y, Li J, et al. 2014. Silencing of Pellino1 improves post-infarct cardiac dysfunction and attenuates left ventricular remodelling in mice. Cardiovasc Res. 102, 46-55
- Gao H, Yin J, Shi Y, et al. 2017. Targeted P2X7 R shRNA delivery attenuates sympathetic nerve sprouting and ameliorates cardiac dysfunction in rats with myocardial infarction. Cardiovasc Ther. 35, 2
- Zhang Y, Bao M, Dai M, et al. 2015. Cardiospecific CD36 suppression by lentivirus-mediated RNA interference prevents cardiac hypertrophy and systolic dysfunction in high-fat-diet induced obese mice. Cardiovasc Diabetol. 14, 69
- Brea MS, Díaz RG, Escudero DS, et al. 2016. Epidermal Growth Factor Receptor Silencing Blunts the Slow Force Response to Myocardial Stretch. J Am Heart Assoc. 5,10
- Suckau L, Fechner H, Chemaly E, et al. 2009. Long-term cardiac-targeted RNA interference for the treatment of heart failure restores cardiac function and reduces pathological hypertrophy. Circulation. 119, 1241-1252
- Miyazaki Y, Ikeda Y, Shiraishi K, et al. 2012. Heart failure-inducible gene therapy targeting protein phosphatase 1 prevents progressive left ventricular remodeling. PLoS One. 7, e35875
- Merentie M, Lottonen-Raikaslehto L, Parviainen V, et al. 2016. Efficacy and safety of myocardial gene transfer of adenovirus, adeno-associated virus and lentivirus vectors in the mouse heart. Gene Ther. 23, 296-305
- Tiscornia G, Singer O, Ikawa M, et al. 2003. A general method for gene knockdown in mice by using lentiviral vectors expressing small interfering RNA. Proceedings of the National Academy of Sciences. 100, 1844-1848
- Pitt B, Williams G, Remme W, et al. 2001. The EPHESUS trial: eplerenone in patients with heart failure due to systolic dysfunction complicating acute myocardial infarction. Eplerenone Post-AMI Heart Failure Efficacy and Survival Study. Cardiovasc Drugs Ther. 15, 79-87
- Zannad F, McMurray JJ, Krum H, et al. 2011. Eplerenone in patients with systolic heart failure and mild symptoms. N Engl J Med. 364, 11-21
- Karmazyn M. 2013. NHE-1: still a viable therapeutic target. J Mol Cell Cardiol. 61, 77-82.
- Pérez NG, Nolly MB, Roldan MC, et al. 2011. Silencing of NHE-1 blunts the slow force response to myocardial stretch. J Appl Physiol. 111, 874-880
- Nolly MB, Pinilla AO, Ennis IL, 2015. Cardiac hypertrophy reduction in SHR by specific silencing of myocardial Na(+)/H(+) exchanger. J Appl Physiol. 118, 1154-1160
- Pitt B, Zannad F, Remme WJ, et al. 1999. The effect of spironolactone on morbidity and mortality in patients with severe heart failure. Randomized Aldactone Evaluation Study Investigators. N Engl J Med. 341, 709-717
- Caldiz CI, Díaz RG, Nolly MB, et al. 2011. Mineralocorticoid receptor activation is crucial in the signalling pathway leading to the Anrep effect. J Physiol. 589, 6051-6061
- Díaz RG, Pérez NG, Morgan PE, et al. 2013. Myocardial Mineralocorticoid Receptor Activation by Stretching and Its Functional Consequences. Hypertension. 63, 112-118
- Montes-Cobos E, Li X, Fischer HJ, et al. 2015. Inducible Knock-Down of the Mineralocorticoid Receptor in Mice Disturbs Regulation of the Renin-Angiotensin-Aldosterone System and Attenuates Heart Failure Induced by Pressure Overload. PLoS One. 10, e0143954
- Hervent AS and De Keulenaer GW. 2012. Molecular mechanisms of cardiotoxicity induced by ErbB receptor inhibitor cancer therapeutics. Int J Mol Sci. 13, 12268-12286
- Forrester SJ, Kawai T, O'Brien S, et al. 2016. Epidermal Growth Factor Receptor Transactivation: Mechanisms, Pathophysiology, and Potential Therapies in the Cardiovascular System. Annual Review of Pharmacology and Toxicology. 56, 627-653.