Research Article - Biomedical Research (2017) Volume 28, Issue 3
Expression and prognostic value of MIR-610 in patients with colorectal cancer
Feng Shi, Rumin Li and Ping Guo*Department of Oncology, the First Affiliated Hospital of Nanyang Medical College, China
- *Corresponding Author:
- Ping Guo
Department of Oncology
The First Affiliated Hospital of Nanyang Medical College, China
Accepted date: August 29, 2016
Abstract
Background: MIR-610 has been identified as a tumor suppressor in colorectal cancer (CRC). However, its prognostic value in colorectal cancer remains unclear. The aim of this study was to explore the significance of miR-610 expression in prognosis of colorectal cancer.
Methods: The expression of MIR-610 was determined in 209 pairs of CRC samples and the corresponding surrounding non-tumor tissues by Quantitative RT-PCR. Then, the association of miR-610 expression with clinicopathological factors and the prognosis of colorectal cancer patients were also analysed.
Results: The expression of MIR-610 was significantly down regulated in colorectal cancer when compared to the non-tumorous tissues (p<0.001). Low level of miR-610 was obviously related to large tumor size, positive lymph node metastasis, and advanced clinical stage. Moreover, Kaplan-Meier survival analysis showed that the overall survival rate of the low miR-610 expression group was significantly lower than that of the high miR-610 expression group (p<0.001). The multivariate analysis identified low MIR-154 expression as an independent predictor of poor survival.
Conclusions: MIR-610 may play an important role in the progression of colorectal cancer, and could be used as an independent prognostic biomarker for patients with colorectal cancer.
Keywords
MIR-610, Colorectal cancer, Prognosis, Overall survival.
Introduction
Colorectal Cancer (CRC) is the second most common type of cancer and the fourth leading cause of cancer-related mortality worldwide [1,2]. Recent progress in diagnosis and therapy methods has helped to cure this disease of many patients at early stage [3], but patients with advanced stage are refractory to existing therapies and have a poor prognosis. For advanced CRC patients, several novel molecular targeted therapies can be applied but with limited therapeutic effects [4,5].
Therefore, the identification of novel diagnostic and prognostic biomarkers for treatment response is eagerly desired. MicroRNAs (miRNAs) are a group of endogenous, small (18-24 nucleotides in length), non-coding RNAs which modulate gene expression by regulating mRNAs via binding to the 3′-untranslated regions (3′-UTR) [6-8].
MiRNAs have been shown to play fundamental roles in diverse biological and pathological processes, including cell proliferation, differentiation, apoptosis, and carcinogenesis [9,10]. Emerging evidence also shows that cancer is associated with very complex genetic alternations in oncogenes and tumor suppressors [11,12]. These findings suggest that miRs act not only as diagnostic and prognostic markers but also as potential therapeutic targets of CRC.
Recently, several studies showed that MiR-610 functions as a tumor suppressor miRNA and is deregulated in different types of cancers, including hepatocellular carcinoma (HCC), gastric cancer (GC) and lung cancer [13-15]. However, the role of miR-610 in CRC and its association with prognosis have not been reported. Here we examine MIR-610 expression in human CRC and investigate the clinical significance of MIR-610 expression.
Materials and Methods
Tissue samples
Surgical specimens of cancer tissue and paired adjacent normal mucosa were obtained from 209 patients with colorectal cancer who underwent surgery at Department of Oncology, the First Affiliated Hospital of Nanyang Medical College from 2007 to 2014. None of the patients had received radiation therapy or chemotherapy before surgery. After collection, all tissue samples were immediately stored in RNA stabilization reagent until use.
Survival time was calculated from the date of surgery to the time of death. The data of patients were shown in Table 1. Informed consent was obtained from all patients, and the research was approved by the Ethics Committee of the First Affiliated Hospital of Nanyang Medical College.
| Variables | Cases (n=209) | MIR-610 expression level | p value | |
|---|---|---|---|---|
| Low expression | High expression | |||
| Age (years) | ||||
| <55 | 109 | 58 | 51 | 0.37 |
| ≥55 | 100 | 47 | 53 | |
| Gender | ||||
| man | 112 | 52 | 60 | 0.236 |
| woman | 97 | 53 | 44 | |
| Tumor size (cm) | ||||
| <5 | 98 | 40 | 58 | 0.01 |
| ≥5 | 111 | 65 | 46 | |
| Histology/differentiation | ||||
| Well+moderate | 107 | 48 | 59 | 0.111 |
| Poor | 102 | 57 | 45 | |
| Depth of invasion | ||||
| T1+T2 | 91 | 41 | 50 | 0.188 |
| T3+T4 | 118 | 64 | 54 | |
| N classification | ||||
| Negative | 96 | 38 | 58 | 0.005 |
| Positive | 113 | 67 | 46 | |
| TNM stage | ||||
| I | 43 | 14 | 29 | 0.003 |
| II | 65 | 29 | 36 | |
| III | 101 | 62 | 39 | |
Table 1: A comparison of miR-610 expression in CRC and clinicopathological features.
Total RNA isolation and quantitative reverse transcription-polymerase chain reaction (qRT-PCR)
To detect miRNA expression by qRT-PCR, total RNA was extracted from tissue samples using the Trizol reagent (Invitrogen, Carlsbad, CA, USA). Quantification of miRNAs was performed using the miRscript SYBR Green PCR kit (Qiagen, Hilden, Germany). The PCR program for detecting miRs was as follows: 50°C for 2 min, 95°C for 10 min, 40 cycles of 95°C for 15 s, and 60°C for 1 min. the U6 gene was used as an internal control to normalize variances. The fold changes in miRNAs were calculated by the equation 2-ΔCt, where ΔCT = (CTmiR-124 - CTU6).
Statistical analysis
The comparison of the expression level of MIR-610 between colorectal cancer tissue and adjacent normal tissue was performed using the two-sample Student’s t test. Either independent sample t test or one-way ANOVA was used to evaluate the possible relationships between the expression levels of miR-610 and various clinicopathological parameters. Overall Survival (OS) curves were analyzed using the Kaplan- Meier method, and the differences were examined using logrank tests. A multivariate survival analysis was performed for all parameters that were significant in the univariate analyses using the Cox regression model. P values<0.05 were considered as statistically significant. Statistical analyses were performed by SPSS Windows version 19.
Results
Expression of miR-610 was downregulated in CRC tissues
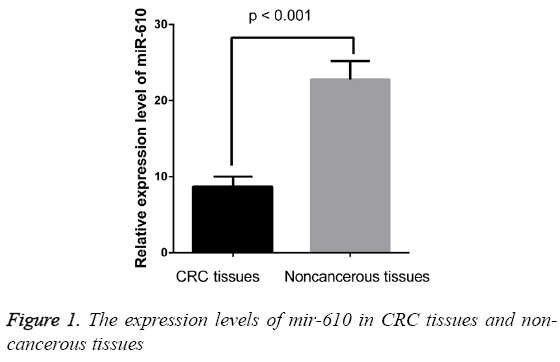
The expression levels of MIR-610 were detected in 209 pairs of CRC and adjacent normal colorectal tissues normalized to RNU6B. As shown in Figure 1, we found the expression levels of MIR-610 to be distinctly decreased in tissues with CRC compared with adjacent normal colorectal tissues. The results showed that MIR-610 expression level in human CRC tissues was significantly lower than that in adjacent non-tumorous tissues (p<0.01).
The association between miR-610 expression and clinical-pathological parameters
The mean expression level of MIR-610 was used to classify the patients with CRC into high-MIR-610 group (>0.52) and low- MIR-610 group (<0.52). As shown in Table 1, we observed that the expression level of MIR-610 was positively correlated with tumor size (p=0.01), lymph node metastasis (p=0.005), and advanced clinical stage (p=0.003). No statistically significant association of MIR-610 expression with patients’ age, gender, tumor differentiation, and depth of invasion.
Low MIR-610 expression predicts poor prognosis in patients with CRC
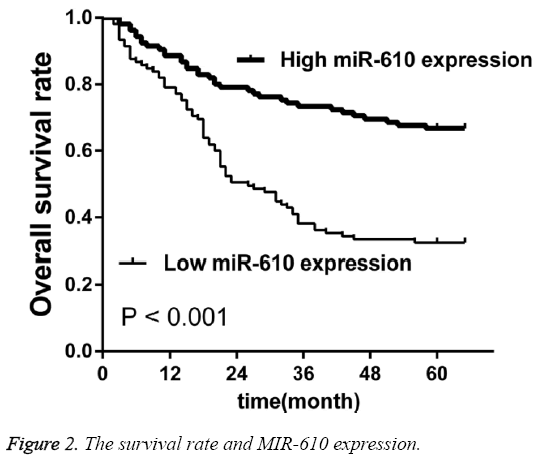
To investigate the prognostic value of MIR-610 for CRC using the Kaplan-Meier method and log-rank test, we observed that OS was significantly lower in the patients with low MIR-610 level than in those with high levels (p<0.001, Figure 2). Multivariate analysis revealed MIR-610 (p=0.017), lymph node metastasis (0.004) and TNM stage (p=0.002) played a significant role of independent prognostic markers in overall survival rates (Table 2).
| Variable | Univariate analysis | Multivariate analysis | ||||
|---|---|---|---|---|---|---|
| HR | 95 % CI | P value | HR | 95 % CI | P value | |
| Age (years) | 1.521 | 0.331-3.761 | 0.312 | - | - | - |
| Gender | 1.235 | 0.422-2.561 | 0.841 | - | - | - |
| Histological grading | 1.663 | 0.681-3.631 | 0.164 | - | - | - |
| Tumor size | 2.778 | 1.238-5.313 | 0.031 | 2.264 | 0.893-4.717 | 0.089 |
| Depth of invasion | 2.442 | 1.212-5.668 | 0.036 | 1.763 | 0.732-3.414 | 0.253 |
| Lymph node metastasis | 5.617 | 2.015-11.441 | 0.003 | 5.226 | 2.427-11.891 | 0.004 |
| Clinical staging | 3.164 | 1.448-5.782 | 0.019 | 5.671 | 2.341-11.991 | 0.002 |
| MIR-610 | 3.771 | 1.679 | 0.002 | 2.984 | 1.212-4.883 | 0.017 |
Table 2: Univariate and multivariate analyses of prognostic factors in colorectal cancer.
Discussion
Although various treatment improvements have been developed for CRC, including molecular targeted therapy, gene therapy, and immunotherapy. However, the 5-year overall and disease-free survival rates for CRC patients are around 50-60% [16]. More and more studies have shown that miRNAs could serve as an efficient prognostic factor for cancer patients. In our present study, we explore the prognostic value of MIR-610 in CRC patients. To our knowledge, this is the first time to investigate the clinical significance of MIR-610 expression in CRC. Our results showed that the expression levels of MIR-610 in tumor samples were much lower than those in normal colonic mucosa samples (p<0.001) and the lowexpression level of MIR-610 was correlated with large tumor size (p=0.01), positive lymph node metastasis (p=0.005), and advanced clinical stage (p=0.003). Furthermore, Patients with lower miR-610 expression had significantly poorer OS. In a multivariate Cox model, we found that MIR-610 expression was an independent predictor of poor prognosis. All those results suggest that miR-610 play an important role in prognostic of patients with CRC.
MIR-610 downregulation has been reported to play important roles in cancer progression. For instance, Zeng et al. reported MIR-610 was downregulated in human HCC cells and tissues and overexpression of miR-610 reduced, HCC cell proliferation and tumorigenicity both in vitro and in vivo [13]. Wang et al. found that MIR-610 function as a tumor suppressor by repressing Vasodilator-Stimulated Expression Phosphoprotein (VASP) [14]. Jin et al. observed that MIR-610 suppresses lung cancer cell proliferation and invasion by targeting GJA3 [15]. Those results informed that MIR-610 expression may be correlated with tumor progression. Recently, Li et al. firstly reported that MIR-610 was downregulated in CRC [17]. Furthermore, Sun et al. found that MIR-610 expression in CRC tissues was significantly lower than that in matched normal adjacent tissues and identified Hepatoma-derived Growth Factor (HDGF) as a direct target of MIR-610 in CRC cells [18]. Those results indicated that miR-610 may function as a potential tumor suppressor. Suggesting that miR-610 could be used as a marker of poor prognosis in CRC patients.
Conclusion
Our results revealed that MIR-610 was down-expressed in glioma tumors and that high expression level of MIR-610 was strongly associated with poor prognosis. MIR-610 may thus be useful for evaluating prognosis and may provide a novel target for the treatment of patients with CRC. Hence, the detailed biological mechanism of MIR-610 down-regulation in CRC tumorigenesis and progression deserves further study.
References
- Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. CA Cancer J Clin 2011; 61: 69- 90.
- Haggar FA, Boushey RP. Colorectal cancer epidemiology: Incidence, mortality, survival, and risk factors. Clin Colon Rectal Surg2009; 22: 191-197.
- Cunningham D, Humblet Y, Siena S. Cetuximabmonotherapy and cetuximab plus irinotecan in irinotecan-refractory metastatic colorectal cancer. NEJM 2004; 351: 337–345.
- Hill DA, Furman WL, Billups CA, Riedley SE, Cain AM, Rao BN, Pratt CB, Spunt SL. Colorectal carcinoma in childhood and adoles¬cence: a clinicopathologic review. J ClinOncol 2007; 25: 5808-5814.
- Zhang J, Fei B, Wang Q, Song M, Yin Y, Zhang B, Ni S, Guo W, Bian Z, Quan C, Liu Z, Wang Y, Yu J. MicroRNA-638 inhibits cell proliferation, invasion and regulates cell cycle by targeting tetraspanin 1 in human colorectal carcinoma. Oncotarget 2014.
- Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004; 116: 281-297.
- Ma Y, Zhang P, Yang J, Liu Z, Yang Z, Qin H. Candidate microRNA biomarkers in human colorectal cancer: systematic review profiling studies and experimental validation. Int J Cancer 2012; 130: 2077-2087.
- Lee YS, Dutta A. MicroRNAs in cancer. Annu Rev Pathol 2009; 4: 199–227.
- Krol J, Loedige I, Filipowicz W. The widespread regulation of microRNA biogenesis, function and decay. Nat Rev Genet. 2010; 11: 597–610.
- Chen CZ, Li L, Lodish HF, Bartel DP. MicroRNAs modulate hematopoietic lineage differentia¬tion. Science 2004; 303: 83-86.
- Babashah S, Soleimani M. The oncogenic and tumour suppressive roles of microRNAs in cancer and apoptosis. Eur J Cancer 2011; 47: 1127–1137.
- Hollis M, Nair K, Vyas A, Chaturvedi LS, Gambhir S, Vyas D. MicroRNAs potential utility in colon cancer: early detection, prognosis, and chemosensitivity. World J Gastroenterol 2015; 21: 8284–8292.
- Zeng XC, Liu FQ, Yan R, Yi HM, Zhang T, Wang GY, Li Y, Jiang N. Downregulation of miR-610 promotes proliferation and tumorigenicity and activates Wnt/beta-catenin signaling in human hepatocellular carcinoma. Mol Cancer 2014; 13: 261.
- Wang J, Zhang J, Wu J, Luo D, Su K, Shi W, Liu J, Tian Y, Wei L. MicroRNA-610 inhibits the migration and invasion of gastric cancer cells by suppressing the expression of vasodilator-stimulated phosphoprotein. Eur J Cancer 2012; 48: 1904-1913.
- Jin J, Li C, You J, Zhang B. miR-610 suppresses lung cancer cell proliferation and invasion by targeting GJA3. ZhonghuaZhong Liu ZaZhi 2014; 36: 405-411.
- Amano R, Yamada N, Nakata B. A prognostic indicator for the resection of liver metastasis of colorectal cancer. Surg Today. 2014; 44:1287–1292.
- Li E, Ji P, OuyangN, Zhang Y, Wang XY, Rubin DC, Davidson NO, Bergamaschi R, Shroyer KR, Burke S, Zhu W, Williams JL. Differential expression of miRNAs in colon cancer between African and Caucasian Americans: implications for cancer racial health disparities. Int J Oncol 2014; 45: 587-594
- Sun B, Gu X, Chen Z, Xiang J. MiR-610 inhibits cell proliferation and invasion in colorectal cancer by repressing hepatoma-derived growth factor. Am J Cancer Res. 2015; 5: 3635-3644.