Research Article - Biomedical Research (2017) Volume 28, Issue 14
Development of Zaocys dhumnades (Cantor) DNA test kit and its application in quality inspection of commercial products
Li Zitong1, Sun Jingyu2, Ai Jinxia3, Li Yingnuo1, Li Mingcheng3, Zhang Lihua4 and Yuan Guangxin1*1College of Pharmacy, Beihua University, Jilin, PR China
2Research Institute of Tuberculosis Control, Changchun, PR China
3School of Laboratory Medicine, Beihua University, Jilin, PR China
4LeiBo Technology Co., Ltd., Jilin, PR China
Accepted date: June 15, 2017
Abstract
Purpose: To develop a Zaocys dhumnades (Cantor) DNA test kit, evaluate its quality indexes including specificity, stability, sensitivity and repeatability, and inspect the qualities of commercial Zaocys dhumnades samples.
Methods: Modern DNA fingerprint technology was used to optimize and improve the detection method of Zaocys dhumnades recorded in Pharmacopoeia of People’s Republic of China (2015), Zaocys dhumnades DNA extraction and detection reagents were research and developed, a Zaocys dhumnades DNA test kit were developed, and detection parameters of the kit were also evaluated. Eighteen samples from Beijing, Tianjin, Changchun and Jilin Cities were detected using the kit, and a comparative test of using the kit with the method recorded in the pharmacopoeia was conducted at the same time.
Results: The Zaocys dhumnades DNA detection kit was still effective after repeated freezing and thawing 5, 10 and 20 times. Fourteen of the 18 commercial Zaocys dhumnades samples were the certified products and 4 of them were counterfeits. Repeated tests demonstrated that the specificity, stability and repeatability of the kit were exactly the same. The results by the kit method were consistent with those by the pharmacopoeia method, and the DNA testing could be completed using the kid in a common laboratory.
Conclusion: The Zaocys dhumnades identification using the Zaocys dhumnades DNA detection kit is simple, convenient, accurate and effective, suitable for the rapid detection of Zaocys dhumnades DNA.
Keywords
Zaocys dhumnades, DNA detection kit, Pharmacopoeia method, PCR.
Introduction
Zaocys dhumnades is the dried bulk of Am Family animals Zaocys dhumnades (Cantor). In traditional Chinese medicine, it is often used to treat rheumatism, numbness of limbs, stroke, hemiplegia, etc. [1]. It is mainly distributed in Fujian and Yunnan. It primarily contains protein, fat and other ingredients. At present, Zaocys dhumnades is bred in a way of semi-farmed and semi-wild, with a high production cost and the production of Zaocys dhumnadesis is decreasing year by year. with the accelerated pace of aging in China, cerebrovascular and cardiovascular patients are also gradually increasing, and Zaocys dhumnadesisthe preparations have been a good choice for the treatment of these diseases, so that the market demand for it is increasing and at the same time the counterfeit is increasing in the market, seriously harming the interests of consumers. Thus it is necessary to look for a quick and accurate method to identify the authenticity of Zaocys dhumnades snakes [2].
Pharmacopoeia of People’s Republic of China (Ch.P) 2015 included biological methods for the identification of Zaocys dhumnades, but they are rarely used by the drug inspection departments nationwide. This method for the identification of Zaocys dhumnades requires a better environment and more complex operations. Based on the pharmacopoeia method, we improved the DNA extraction and identification by PCR, and developed a Zaocys dhumnades DNA detection kit [3]. In the experiment, we used the kit to identify genuine Zaocys dhumnades and commercially available samples, respectively, and then the kit method was compared with the Pharmacopoeia method. The results confirmed that the modified DNA Zaocys dhumnades kit method should be more convenient and quicker than the pharmacopoeia method, without needing the Zaocys dhumnades reference crude material and with a better reproducibility and more accurate results.
Materials and Methods
Medicinal herbs
Genuine Zaocys dhumnades medicinal herbs (verified as the certified product by the method recorded in Ch.P (2015). From Beijing, Tianjin, Changchun and Jilin, we randomly selected 18 Zaocys dhumnades samples (Table 1).
| Number | Source | Size/g | Genuine or counterfeit |
|---|---|---|---|
| BJWSS-1 | Beijing | 5 | Genuine |
| BJWSS-2 | Beijing | 5 | Genuine |
| BJWSS-3 | Beijing | 5 | Genuine |
| TJWSS-1 | Tianjin | 5 | Genuine |
| TJWSS-2 | Tianjin | 5 | Genuine |
| TJWSS-3 | Tianjin | 5 | Genuine |
| TJWSS-4 | Tianjin | 5 | Counterfeit |
| TJWSS-5 | Tianjin | 5 | Genuine |
| TJWSS-6 | Tianjin | 5 | Counterfeit |
| TJWSS-7 | Tianjin | 5 | Genuine |
| CCWSS-1 | Changchun | 5 | Genuine |
| CCWSS-2 | Changchun | 5 | Genuine |
| CCWSS-3 | Changchun | 5 | Counterfeit |
| CCWSS-4 | Changchun | 5 | Genuine |
| CCWSS-5 | Changchun | 5 | Genuine |
| CCWSS-6 | Changchun | 5 | Genuine |
| JLWSS-1 | Jilin | 5 | Genuine |
| JLWSS-2 | Jilin | 5 | Counterfeit |
Table 1: Sources and identification results of commercially available Zaocys samples.
Instrument
UV WHITE-2020D Gel imaging analyzer (BioRad, USA); UV-2550 analyzer (SHIMADZU International Trade Co., LTD., Shanghai); Gene AmpR PCR System 2700 PCR instrument (Per-kin-Elmer Company, USA); H2050R table high speed refrigerated centrifuge (Xiang-yi Instrument Centrifuge Instrument Co., LTD, Changsha); UV gel imaging analyzer (Biorad Company, USA); Sha-B water bath temperature oscillator (Jianfsu Province Jintan Medical Instrument Factory, Jiangsu); EDC-810 PCR ABI Company, USA); DYY-4 voltage steady flow electrophoresis apparatus (Beijing City Liuyi Instrument Factory, Beijing).
Reagent Kit
The amount of reagents in the kit could be used 20 times for the detection, which consisted of 8 parts, including DNA extraction and PCR amplification (Table 2).
| Name | Specification | Function |
|---|---|---|
| P1 | 50ml | Destroy cell well, lease DNA |
| P2 | 50ml | Surface active agent DNA |
| P3 | 50ml | Catalysis DNA |
| P4 | 50ml | Precipitate DNA |
| P5 | 50ml | Precipitate DNA |
| P6 | 50ml | Dissolve DNA |
| W-positive | 50μl | As the positive control |
| W-negative | 50μl | As the negative control |
| W-reaction | 23μl | PCR reaction system |
Table 2: Make-up of Zaocys dhumnades DNA detection kit.
Methods
Kit method
Genomic DNA extraction of Zaocys dhumnades samples: Zaocys dhumnades samples were cut up to pieces with a size about 1 mm3, and then 0.05 g of the samples was dissolved in 500 μL P1, 30 μL P2 and 15 μL P3, mixed evenly and shaken at 56 in a water bath for 16-18 h. The solution was added to 500 μL P4, which was gently shaken for 10 min and centrifuged at 4°C (11000 rpm × 10 min), and the obtained supernatant was added to the same volume of P5 and placed at -20°C for 1 h. Then the supernatant solution was centrifuged at 4°C (11000 rpm × 10 min), the supernatant was discarded and 80 μL P6 was added to the precipitation for dissolving the DNA, which would be used as the DNA test solution [4-6].
Detection of genomic DNA concentration and purity: The DNA extracte was properly diluted, the absorbances at 260 nm (A260) and 280 nm (A280) were measured by UV spectrophotometer, and the ratio of A260 to A280 was used to determine the concentration and purity of DNA.
PCR amplification: 2 μL of the DNA test solution, positive control solution and negative control solution each were placed in PCR reaction tubes, respectively. The PCR parameters were adjusted to 95°C for 5 min for the pre-denaturation, circular reaction 30 times (95°C for 30 s, 63°C for 45 s, and 72°C for 30 s) and 72°C for 5 min for the extending.
Detection by agarose gel electrophoresis: 12 μL PCR reaction product and 2 μL 6 × loading buffer were evenly mixed, and then the mixed solution was loaded on 2% agarose gel plate. The voltage was 15 V·cm-1 and the running duration was 20-30 min, and the electrophoresis was stopped when the blue belt moved to 2-4 cm away from the plastic front. The gel was observed and photographed by an ultraviolet analyzer.
Pharmacopoeia method
In the same laboratory and under the same experimental conditions as those in the kit method, the same tester detected the commercial simples and the certified products identified by The National Institute for The Control of Pharmaceutical and Biological Products according to the Pharmacopoeia method.
Evaluation of kit detection parameters
Specificity: The selected 16 Zaocys dhumnades genuine and 4 commercial counterfeits were randomly labeled, and tested with the test kit method, respectively.
Stability: A kit kept at -20°C was randomly selected and thawed at room temperature, and then replaced at -20°C for the refrigeration, which was repeated 5, 10 and 20 times, respectively, and then the DNA extraction reagents, the positive reference substance and the negative reference substance were assayed.
Repeatability: Three genuine Zaocys dhumnades products and one counterfeit were randomly selected, and tested three times under the same laboratory conditions by the same tester in line with the kit method.
Experimental results
Judgment basis
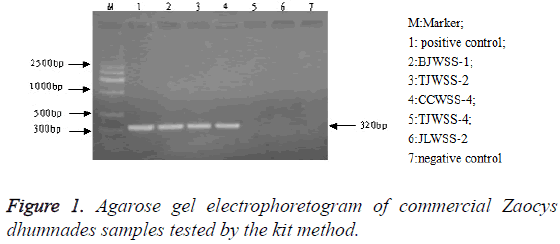
Ch. P (2015) specifies that there is a band of DNA at 300-400 bp in the gel electrophoretogram of Zaocys dhumnades tested by DNA fingerprint identification technology, which is corresponding to that of reference crude herb, but no band in that of the negative control, indicating that the sample could be a genuine Zaocys dhumnades.
Comparison on results by kit method with those by Pharmacopoeia method
Genomic DNA concentration and purity detection by the two methods: The genomic DNA concentration and purity tested by the two methods both were within 1.80 ± 0.23, showing that there was no protein contamination in the samples extracted by the two methods (Table 3).
| Med | A260 | A280 | Purity | ρ/μg·L-1 | ||||
|---|---|---|---|---|---|---|---|---|
| Mat | Method of kit | Method of pharmacopoeia | Method of kit | Method of pharmacopoeia | Method of kit | Method of pharmacopoeia | Method of kit | Method of pharmacopoeia |
| ZPWSS-1 | 0.343 | 0.352 | 0.204 | 0.215 | 1.681 | 1.637 | 1.71 | 1.76 |
| ZPWSS-2 | 0.348 | 0.332 | 0.211 | 0.203 | 1.649 | 1.635 | 1.74 | 1.66 |
| BJWSS-1 | 0.341 | - | 0.204 | - | 1.672 | - | 1.70 | - |
| BJWSS-2 | 0.338 | - | 0.206 | - | 1.641 | - | 1.69 | - |
| BJWSS-3 | 0.339 | - | 0.207 | - | 1.638 | - | 1.69 | - |
| TJWSS-1 | 0.340 | - | 0.204 | - | 1.667 | - | 1.70 | - |
| TJWSS-2 | 0.340 | - | 0.205 | - | 1.659 | - | 1.70 | - |
| TJWSS-3 | 0.337 | - | 0.206 | - | 1.636 | - | 1.68 | - |
| TJWSS-4 | 0.336 | - | 0.201 | - | 1.672 | - | 1.68 | - |
| TJWSS-5 | 0.344 | - | 0.206 | - | 1.670 | - | 1.72 | - |
| TJWSS-6 | 0.350 | - | 0.214 | - | 1.636 | - | 1.75 | - |
| TJWSS-7 | 0.350 | - | 0.213 | - | 1.643 | - | 1.75 | - |
| CCWSS-1 | 0.338 | - | 0.205 | - | 1.649 | - | 1.69 | - |
| CCWSS-2 | 0.347 | - | 0.208 | - | 1.668 | - | 1.73 | - |
| CCWSS-3 | 0.349 | - | 0.210 | - | 1.662 | - | 1.75 | - |
| CCWSS-4 | 0.335 | - | 0.201 | - | 1.667 | - | 1.67 | - |
| CCWSS-5 | 0.342 | - | 0.202 | - | 1.693 | - | 1.71 | - |
| CCWSS-6 | 0.351 | - | 0.215 | - | 1.633 | - | 1.75 | - |
| JLWSS-1 | 0.332 | - | 0.203 | - | 1.635 | - | 1.66 | - |
| JLWSS-2 | 0.341 | - | 0.202 | - | 1.688 | - | 1.70 | - |
Table 3: Results of purities and concentrations detection of 18 commercially available Zaocys dhumnades (Cantor) samples.
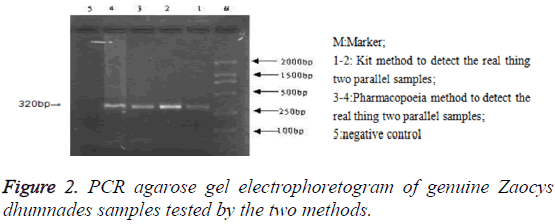
PCR products agarose gel electrophoresis: After PCR amplification with specific primer, both the genuine and 14 commercial simples had specific binding sites. The amplified products were 300-400 bp and their location was consistent with that of the positive samples, but there was no band in the electrophoretogram of negative control and 4 counterfeit samples (Figure 1). The testing results of the two methods were identical (Figure 2).
Evaluation of kit detection parameters
Specificity: Fourteen genuine Zaocys dhumnades and 4 counterfeits were detected by the kit method. The results showed that all the commercially certified products presented an amplified band at 300-400 bp, but the counterfeits not, indicating that the kit method had a good specificity.
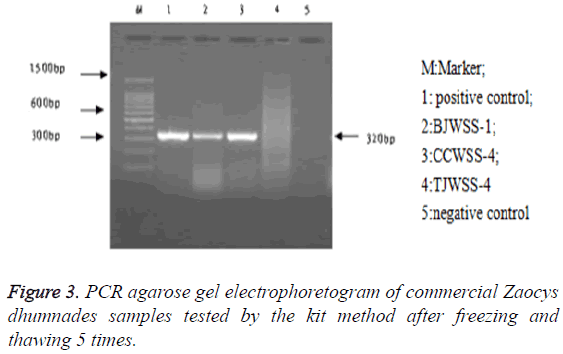
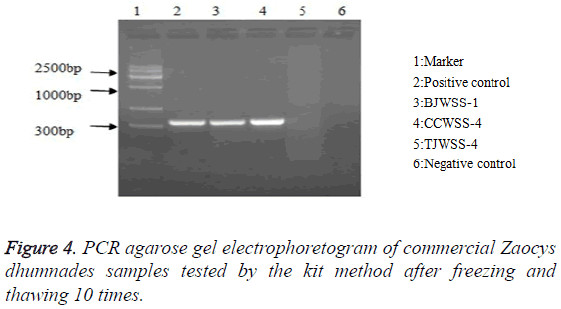
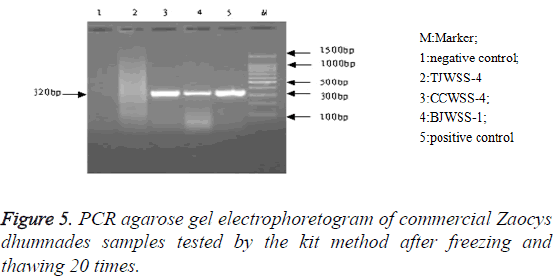
Stability: After repeated freezing and thawing 5, 10 and 20 times, the kit could be still used to extract the genomic DNA, with a target band between 300-400 bp, showing a good stability of the kit (Figures 3-5).
Repeatability: Three commercial genuine Zaocys dhumnades samples and one counterfeit sample were repeatedly tested three times. All the 3 genuine samples showed amplified bands between 300-400 bp and their location was consistent with that of the positive control, and 4 counterfeits and the negative control did not show the amplified band, suggesting that kit should have a good repeatability.
Discussion
There are many methods to evaluate the quality of Chinese herbal medicine Zaocys dhumnades currently, but all of them show some limitations. The traditional identification method is used to identify drugs by observing the characters of genuine and counterfeit herbal medicines, simple and feasible, but with a strong subjectivity, and its accuracy depends on the authenticators’ experience, so its identification is low. The repeatability of thin layer chromatography (TLC) is better, but for Zaocys dhumnades that ingredients of its counterfeits are similar to those of its certified products; so far it is difficult for it to be identified. The main advantage of infrared spectra analysis is that it cannot destroy samples and test results can provide rich information, but the spectrum analysis requires experiment personnel who have relatively rich experiences. High performance liquid chromatography (HPLC) has high detection sensitivity, but the cost for the analysis and daily instrument maintenance is very high. Two-phase titration method identifies genuine and counterfeit Zaocys dhumnades by measuring the content of alkaloids contained in Zaocys dhumnades, fast and convenient, but the method is influenced by the external conditions greatly, so that it is not stable enough [7].
In Ch. P (2015), DNA fingerprint detection method was proposed as the national standard of Zaocys dhumnades detection. DNA fingerprinting identification is a molecular identification method of traditional Chinese medicines based on DNA molecule. The DNA sequences of living bodies cannot change whether in the process of growth or through artificial processing and specific species have their own specific DNA sequences, so that the method is especially suitable for use. However, the promotion of the pharmacopoeia method is restricted by many factors. First of all, the pharmacopoeia method has high requirements on the experimental environment since DNA molecular identification method is a kind of micro identification method, in which to avoid contamination is a key step. Secondly, the process of the pharmacopoeia method is tedious, complicated and timeconsuming.
In view of the above several factors, now a simple, rapid and accurate method is urgently needed to regulate the Zaocys dhumnades medicinal material market. On the basis of the pharmacopoeia method, our team independently researched and developed a Zaocys dhumnades DNA detection kit, both the pharmacopoeia method and the kit method were used to detect the standard and reference Zaocys dhumnades at the same time in this study, and the results showed that the results obtained by the two identification methods were consistent, indicating that the kit method used to identify Zaocys dhumnades should be reliable. The comparison of the 2 methods indicates that the kit method can not only reserve the advantages of the pharmacopoeia method, but also meet the requirements for the simple, rapid and accurate detection, with an advantage of widespread in grassroots units [8].
This kit can be used in the whole process from the DNA extraction to the identification by PCR. First, the operation steps can be optimized, so that the original repeated process becomes simple, the authenticator’s identification workload is reduced at the greatest extent, the identification period is shortened, and the possibility of DNA contamination caused by the operation of experimental personnel is lowered, so as to achieve the conditions in which the method can be popularized. Secondly, there are uniform reagents in the reagent kit, which can avoid the influence of various reagents from multiple sources on the stability of experiment. Finally, the detection accuracy is high, without subjective dependence. The results are intuitive and easy to judge. Therefore, this kit can be widely applied to the testing of herbal medicines, especially in grassroots units.
Acknowledgement
This work was funded by the Strategic New Industries and High-tech Development Project of Jilin Province (No. 2013G030No.2015Y077) and the Science and Technology Project Development Finance Project of Jilin province (20160204004NY) and Science and Technology Innovation Development Project by Jilin City (20162006) and The Young teacher promotion program of Beihua University.
References
- Lin Z, Hu LN, Li N. Animal medicine research Zaocys dhumnades. J Traditional Chinese Med 2009; 29: 982-984.
- Gu YG, Zhang LH, Fu GL. Identification of velvet antler by mitochondrial DNA fingerprint.Chin Pharm J 2013; 48: 170-173.
- Zhang X, Song L, Yu W, Zhou T, Li M. Development and evaluation of a DNA detection kit on authentication of Zaocys dhumnades based on a bioinformatics method and PCR technology. 7th International Conference on Biomedical Engineering and Informatics, Dalian, 2014.
- Liu TH, Wang J, Zhang LH. Identification of cytochrome b bene of Chinese herb Testudinis Carapax et plastrum. Chin Pharm J 2012; 47: 182-185.
- Liu Y, Yang MY, Zhang LH. Identification and characterization of DNA finger print between penis ettestis of cervusnippon temm ink and bullwhip. Lishizhen Med Mater Med Res 2010; 21: 993-995.
- Wang S,Wang H, Zhang LH. Identification of Panax ginsheng C. A. Meyer Cv. Silvatica and cultivated Panax ginseng by DALP fin-gerprint. Chin Pharm J 2013; 48: 677-680.
- Zhou TT, Yu WJ, Li MC. The development of the unibract fritillary bulb DNA testing kits and evaluation. Chin Pharm J 2014; 49: 501-504.
- Yu WJ, Zhou TT, Zhang LH. Evaluation and application agkistrodon DNA testing kit. Chin Pharm J 2014; 49: 999-1003.