Research Article - Biomedical Research (2017) Volume 28, Issue 16
The role of miR-25 in pediatric neuroblastoma
Chuantao Ren1,2#, Chengfei Li2#, Hongtao Zhang2, Lijian Gao3 and Aiwu Li1*
1Department of Pediatric Surgery, Qilu Hospital of Shandong University, Jinan, PR China
2Department of Pediatric Surgery, Dezhou People’s Hospital, Dezhou, PR China
3Department of Urology, Dezhou People's Hospital, Dezhou City, Dezhou, PR China
#These authors contributed equally to this study
- *Corresponding Author:
- Aiwu Li
Department of Pediatric Surgery
Qilu Hospital of Shandong University
PR China
Accepted date: July 27, 2017
Abstract
Aims: This study is to investigate the role of miR-25 in pediatric neuroblastoma.
Methods: The expression of miR-25 in pediatric neuroblastoma tissues and cell lines was determined by quantitative PCR. SH-SY5Y and IMR32 cells were transfected with miR-25 mimics and inhibitor, respectively. The effect of miR-25 on pediatric neuroblastoma was analysed by CCK-8 assay, flow cytometry and Transwell assay. Western blot was performed to detect the protein expression of TOB1, a potential target gene of miR-25. Dual luciferase reporter assay was carried out to determine the direct binding between miR-25 and TOB1.
Results: The expression of miR-25 was upregulated in pediatric neuroblastoma tissues and positively correlated to distant metastasis and clinical stages. The expression of miR-25 in the metastases derived SH-SY5Y cells was higher than that in the primary tumor derived IMR32 cells. In vitro experiments showed that over expression of miR-25 promoted the proliferation, G1/S phase transition and metastasis of IMR32 cells, while down regulation of miR-25 inhibited the proliferation, G1/S phase transition and metastasis of SH-SY5Y cells. Upregulation of miR-25 decreased the expression of TOB1 in IMR32 cells, while down-regulation of miR-25 increased the expression of TOB1 in SH-SY5Y cells. Dual luciferase reporter assay confirmed that miR-25 could bind to the 3’-UTR of TOB1, indicating that TOB1 is the direct target gene of miR-25.
Conclusions: The miR-25 expression in neuroblastoma and neuroblastoma cells is upregulated. MiR-25 can promote the proliferation, invasion and metastasis of neuroblastoma cells by regulating TOB1 expression.
Keywords
miR-25, Pediatric neuroblastoma, TOB1, Invasion, Metastasis
Introduction
Neuroblastoma (NB) is the most common tumor in children with an incidence rate of 10% [1], which is second only to leukemia and central nervous system tumors [2]. The prognosis of NB is poor [3]. NB belongs to neuroendocrine tumors, which is originated from the neural crest progenitors in the sympathetic nervous system, and usually occurs in the nerve tissues of the adrenal gland, neck, chest, and abdomen [4]. NB has the clinical features of high malignancy and heterogeneity, prone to metastasis and relapse and poor prognosis [5]. The clinical treatment effect for NB is limited and the patients’ 5 y survival rate was only 40%, except for a few patients who can spontaneously regress or completely resected at early stage [6,7]. Genetic abnormalities, such as V-Myc Avian Myelocytomatosis Viral Oncogene neuroblastoma derived homolog (MYCN) abnormal expression and missing of tumor suppressor gene sequences lp36.1 and lp36.2, are closely related to child NB classification, treatment and prognosis [8,9]. In addition, some new biomarkers were reported to be associated with the survival of NB. For instance, upregulation of ARID1B gene in NB patients suggests a worse prognosis [10]. The expression level of miR-149 was negatively correlated with the prognosis of NB patients and had potential clinical diagnostic values [11]. However, the molecular mechanisms of the occurrence and development of NB are still unclear.
MiRNA is a class of small non-coding RNA molecules with lengths of 18-22 nt and can bind to the 3’-UTR region of mRNA to inhibit its translation and regulate the protein expression [12,13]. Evidence shows that miRNA is abnormally expressed in NB, and many miRNA molecules are involved in the proliferation, invasion, metastasis and apoptosis of NB, suggesting that miRNAs play important roles in NB [14,15]. For example, Li et al. found that miR-1303 could promote SH-SY5Y cell proliferation and metastasis by targeting GSK3β and SFRP1 genes [14]. Li et al. indicated that miR-21 could promote NB cell proliferation and metastasis by regulating CHL1 gene [13]. MiR-25 belongs to the miR-106b-25 family and locates on human chromosome 7. It is upregulated in many tumors and plays important regulatory roles in the tumor occurrence [16-18]. Petroccaet al. found that miR-106b-25 cluster (miR-106b, miR-25 and miR-93) was highly expressed in gastric cancer cells and regulated by E2F1 regulator. Meanwhile, miR-106b and miR-93 can regulate the expression of E2F1, thereby forming a negative feedback path [19]. In Non-Small Cell Lung Cancer (NSCLC), the expression of miR-25 is upregulated, which promotes the proliferation, invasion and metastasis of tumor cells by targeting F-Box and WD Repeat Domain Containing 7 (FBXW7) gene, playing the oncogene functions [20]. These researches indicate that miR-25 plays some important roles in a variety of tumors, and closely related to tumor proliferation, invasion and metastasis. However, the role and mechanism of miR-25 in NB is still unclear.
In this study, the effect and mechanism of miR-25 in NB was investigated. The real-time quantitative PCR, Western blot, Transwell assay and flow cytometry were used.
Materials and Methods
Collection of neuroblastoma tissue samples
Fresh NB tissues were collected from 34 patients with NB between May, 2012 and December, 2015. Among these patients, 16 cases had distant metastasis, 22 cases were male and 12 cases were female. The average age of patients was 5.4 y old. All cases were diagnosed as NB by clinical, imaging and histopathological diagnosis. No patient was subjected to radiation therapy before surgery. The patients with stage III NB received chemotherapy and partially relieved before surgery, which complied with the surgical guidelines. NB tissues with 1 cm in diameter were taken during the surgery. The corresponding tumor adjacent tissue was taken 5 cm away from the cancer tissue. The clinical information and pathological data of the patients were collected. Prior written and informed consent was obtained from every patient and the study was approved by the ethics review board of Shandong University.
Cell culture
NB cell lines SH-SY5Y (metastatic tumor) and IMR-32 (primary tumor) were purchased from Shanghai cell bank of Chinese Academy of Sciences (Shanghai, China). Both cells were cultured in DMEM complete medium at 37°C under an atmosphere containing 5% CO2. Cell medium was changed every two days, and the cells were passaged when the confluency reached 80-90%.
RNA extraction and quantitative RT-PCR
The NB tissues and adjacent tissues were frozen by liquid nitrogen and ground into powder. The tissues were lysed by Trizol® isolation reagent, and the total RNAs were extracted by phenol chloroform method. The RNA concentration was determined by electrophoresis and the A260/A280 ratio was measured by UV spectrophotometer to determine RNA quality. RNA (1 μg) was then reverse transcribed into cDNA.
To detect miR-25 expression, RT-qPCR was performed. The reaction system (20 μL) contained 10 μLqRT-PCR-Mix, 0.5 μL upstream primer (miR-25, 5’-GCGGAGACTTGGGCAATTG-3’), 0.5 μL downstream universal primer (provided by the kit), 3 μL cDNA and 6 μL ddH2O. The reaction protocol was initial denaturation at 95°C for 10 min; 40 cycles of 95°C for 1 min and 60°C for 30 s (iQ5; Bio-Rad, Hercules, CA, USA). The 2-ΔΔCt method was used to calculate the relative expression of miR-25 against internal reference (U6, Forward: CTCGCTTCGGCAGCACA; Reverse: AACGCTTCACGAATTTGCGT). Each sample was tested in triplicate.
Transfection of miR-25 mimics and inhibitor into NB cells
The NB cell lines SH-SY5Y and IMR-32 were seeded in 24-well plates. MiR-NC and miR-25 inhibitor were transfected into SH-SY5Y, while miR-25 miR-NC and miR-25 mimics were transfected into IMR-32. Briefly, 1.25 μL miR-25 (20 μM) mimics/inhibitor, 2 μL lipo2000 and 50 μL Opti Memi were mixed and added to cells. After 6 h of incubation, cell medium was replaced with fresh medium containing 10% FBS. After another 48 h of incubation, cells were collected for further experiments.
CCK-8 assay
The cells were seeded in 96-well plates at a concentration of 3000/well. After transfection with miR-25 mimics or inhibitor, the cell proliferation was detected with CCK-8 assay every 24 h. CCK-8 reaction solution was added and incubated for 30 min, and then the absorbance at 450 nm was measured. Cell proliferation was monitored for three consecutive days and a proliferation curve was plotted.
Transwell assay
A 24-well Transwell chamber (Corning Co., USA) with a diameter of 8 μm was used in this experiment. To the upper wells, 200 μL of serum free DMEM medium containing 2 × 105 cells were added, while 500 μL DMEM medium containing 10% FBS was added to each bottom well. After 24 h of incubation, the cells in the upper wells were removed. The cells in the bottom wells were fixed with 4% formaldehyde for 10 min and stained with giemsa. The cells were observed under 200X magnifications. Five views were randomly selected to count the translocated cells in order to evaluate cell metastasis.
To determine cell invasive ability, matrigel (BD biosciences, USA) was use to simulate extracellular matrix environment. The matrigel was taken out from -20°C one day before the experiment and melted at 4°C overnight. The matrigel was diluted with serum free DMEM medium at 1:2 ratio, and 100 μL solution was evenly applied in the upper wells. The chamber was put into 37°C for 60 min to solidify the gel. All operation was performed on ice, and the pipette tips were pre-cooled at 4°C. All other procedures were the same as that of the cell metastasis assay, and the results were obtained after 72 h.
Flow cytometry
After transfected with miR-25 mimics/inhibitor 24 h, 1 × 106 cells were taken from each group and washed with pre-cooled PBS twice. Cell cycle was analysed using a cell cycle assay kit (BD Biosciences, USA) according to the manufacturer’s instruction. Briefly, 200 μL solution A was added to the cells and incubated for 10 min, and then 150 μL solution B was added and incubated for another 10 min. Finally, 120 μL solution C was added to incubate for 10 min in the dark. The cell cycle was detected by flow cytometry and the results were analyzed by Modifit software (BD Biosciences, NJ, USA).
Western blot
Proteins were isolated from cells after lysis with RIPA. Then, the proteins were separated by SDS-PAGE and transferred to PVDF membrane. The membrane was blocked with 50 g/L skim milk for 1 h at room temperature. After that, primary antibodies for Transducer of ERBB (rabbit anti-human TOB1 polyclonal antibody 1:1000) and mouse anti-human GAPDH monoclonal antibody (1:4000) were added and incubated at 4°C overnight. The membrane was washed with TBST (5 min × 5 times), and then HRP-labeled secondary antibodies (goat anti-mouse 1:3000, goat anti-rabbit 1:3000) were added and incubated at room temperature for 1 h. After washing with TBST (5 min × 5 times), the bands were developed with ECL luminous liquid. All antibodies were purchased from Abcam plc. (Cambridge, UK).
Dual luciferase reporter assay
The target gene of miR-25 was predicted using the Targetscan Online Prediction Software (www.targetscan.com). According to the bioinformatics prediction results, the wild-type and mutant seed regions of the 3’-UTR of the TOB1 gene containing miR-25 binding site were chemically synthesized in vitro. After added with the cleavage sites of Spe-1 and Hind-III at both ends, they were cloned into the pMIR-REPORT luciferase reporter plasmid. The plasmid containing wild-type and mutant 3’-UTR sequence were transfected together with miR-25 mimics into HEK293T cells using the liposome method. After 24 h of incubation, each group of cells was lysed according to the manufacturer’s instruction of Promega dual luciferase reporter assay kit (Promega, Madison, WI, USA). The luminance was recorded by GloMax 20/20 luminometer, using renilla as the internal standard.
Statistical analysis
The statistical analysis was performed with the statistical software SPSS 17.0. All data was expressed as mean ± Standard Deviation (SD). The t-test was used for comparisons between groups, and a p value less than 0.05 was considered as statistically significant.
Results
The expression of miR-25 in NB tissues and cells and the clinical significance
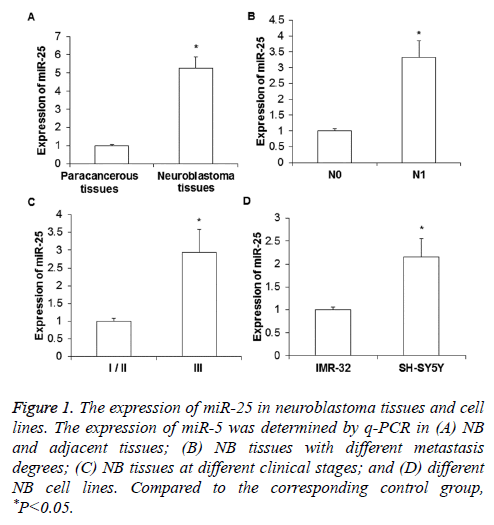
To determine the expression level of miR-25, real-time quantitative PCR was performed. As shown in Figure 1A, the expression of miR-25 in NB tissues was significantly increased compared to that in paracancerous tissues (P<0.01). The miR-25 expression in the tissues of NB patients with distant metastasis (N1) was significantly higher than that of NB patients without metastasis (N0, P<0.05) (Figure 1B). For the comparison between different clinical stages, it was found that miR-25 expression in the tissues of stage III NB was significantly higher than those in the tissues of stage I/II NB (P<0.05) (Figure 1C). In addition, the expression of miR-25 in the metastases-derived NB cell line SH-SY5Y was obviously higher than that in the primary tumor derived IMR-32 cells (P< 0.05) (Figure 1D).
Figure 1: The expression of miR-25 in neuroblastoma tissues and cell lines. The expression of miR-5 was determined by q-PCR in (A) NB and adjacent tissues; (B) NB tissues with different metastasis degrees; (C) NB tissues at different clinical stages; and (D) different NB cell lines. Compared to the corresponding control group, *P<0.05.
Effect of miR-25 on the proliferation of NB cells
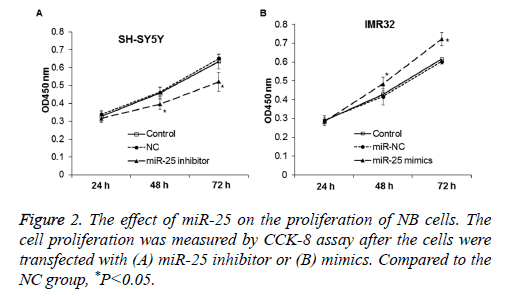
To determine the effect of miR-25 on cell proliferation, CCK-8 assay was performed. Compared with control and NC, the proliferation of NB cell line SH-SY5Y was significantly reduced after transfected with miR-25 inhibitor (P<0.05) (Figure 2A). This indicates that down-regulation of miR-25 may inhibit the proliferation of NB cell SH-SY5Y. On the other hand, expression of miR-25 mimics in IMR-32 cells significantly promoted cell proliferation (P<0.05) (Figure 2B).
Effect of miR-25 on the invasion and metastasis of NB cells
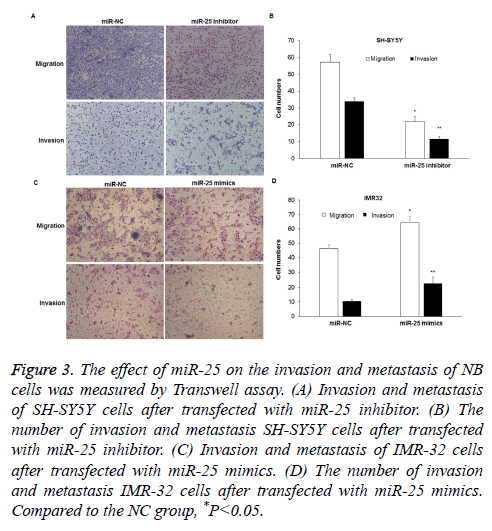
Transwell assay was performed to determine cell invasion and metastasis after transfection with miR-25 inhibitor or mimics. The metastasis ability of SH-SY5Y cells was significantly inhibited after transfected with miR-25 inhibitor, as the number of cells migrated through well membranes was less than that in the NC group (57.3 ± 4.5 vs. 21.7 ± 3.2) (P<0.05) (Figures 3A and 3B). The invasion experiment results showed that the number of SH-SY5Y cells invaded the Transwell membranes decreased (33.8 ± 2.34 vs. 11.5 ± 1.6) (P<0.05). However, when miR-25 expression in IMR32 cells was increased, the metastasis of cells enhanced, as the number of cells migrated the well membranes was higher than that of the NC group (64.3 ± 4.2 vs. 46.3 ± 2.6) (P<0.05) (Figures 3C and 3D). The invasion experiment results showed that the number of cells invaded the well membranes in the miR-25 mimics transfection group was higher than that in the NC group (22.7 ± 3.9 vs. 10.3 ± 1.25) (P<0.05).
Figure 3: The effect of miR-25 on the invasion and metastasis of NB cells was measured by Transwell assay. (A) Invasion and metastasis of SH-SY5Y cells after transfected with miR-25 inhibitor. (B) The number of invasion and metastasis SH-SY5Y cells after transfected with miR-25 inhibitor. (C) Invasion and metastasis of IMR-32 cells after transfected with miR-25 mimics. (D) The number of invasion and metastasis IMR-32 cells after transfected with miR-25 mimics. Compared to the NC group, *P<0.05.
MiR-25 promotes NB cell G1/S phase transition
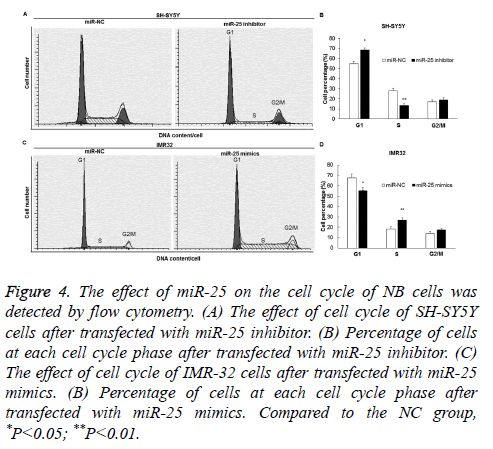
Flow cytometry was used to detect the effect of miR-25 on the cell cycle transition of SH-SY5Y and IMR-32 cells. The results showed that G1/S phase arrest occurred in the SH-SY5Y cells of the miR-25 inhibitor group (Figures 4A and 4B) (P<0.05), while G1/S phase transition was promoted in the IMR-32 cells of the miR-25 mimics group (P<0.05) (Figures 4C and 4D).
Figure 4: The effect of miR-25 on the cell cycle of NB cells was detected by flow cytometry. (A) The effect of cell cycle of SH-SY5Y cells after transfected with miR-25 inhibitor. (B) Percentage of cells at each cell cycle phase after transfected with miR-25 inhibitor. (C) The effect of cell cycle of IMR-32 cells after transfected with miR-25 mimics. (B) Percentage of cells at each cell cycle phase after transfected with miR-25 mimics. Compared to the NC group, *P<0.05; **P<0.01.
The expression of TOB1 protein
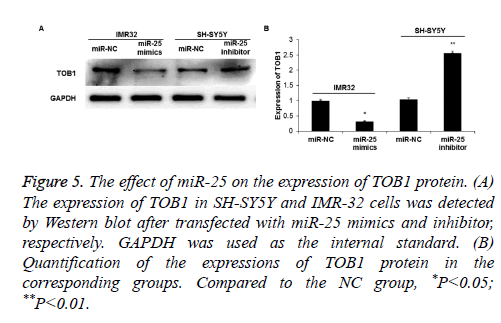
Bioinformatics predicted that the proliferation-related gene TOB1 was one of the target genes of miR-25. To further verify this, Western blot was used to detect TOB1 protein expression. The results showed that the expression of TOB1 in IMR-32 cells was significantly decreased when miR-25 was overexpressed (P<0.05) (Figures 5A and 5B). On the contrary, when miR-25 was down-regulated, the expression of TOB1 in SH-SY5Y cells significantly increased (P<0.05).
Figure 5: The effect of miR-25 on the expression of TOB1 protein. (A) The expression of TOB1 in SH-SY5Y and IMR-32 cells was detected by Western blot after transfected with miR-25 mimics and inhibitor, respectively. GAPDH was used as the internal standard. (B) Quantification of the expressions of TOB1 protein in the corresponding groups. Compared to the NC group, *P<0.05; **P<0.01.
Dual luciferase reporter assay
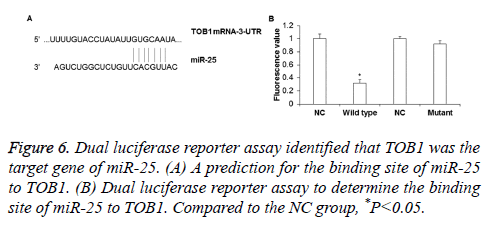
To determine whether TOB1 gene was the target gene of miR-25, dual luciferase reporter assay was performed. The binding sequence of miR-25 on TOB1 gene was shown in Figure 6A. The results showed that the fluorescence value of the wild type group was significantly lower than that in the NC group (P<0.05), while there was no obvious difference between the luminance in the mutant group and NC group (P>0.05) (Figure 6B). This indicates that miR-25 can directly bind to the 3’-UTR of wild type TOB1, while mutation of the 3’-UTR of TOB1 can inhibit miR-25 binding to TOB1. Thus, miR-25 can directly regulate the expression of TOB1 gene.
Discussion
In this study, the expression of miR-25 in NB tissues was detected by quantitative PCR and found to be upregulated. The expression of miR-25 was positively correlated to the distant metastasis and clinical stage of NB. In the in vitro experiment, miR-25 could obviously promote NB cell proliferation, invasion and metastasis. Bioinformatics predicted that TOB1 might be the target gene of miR-25. Upregulation and down-regulation of miR-25 could cause the decrease and increase of TOB1 expression, respectively. Dual luciferase reporter assay confirmed that miR-25 could directly bind to the 3’-UTR region of TOB1 to regulate its expression. Therefore, we suppose that miR-25 may play a role as an oncogene in NB and may promote the occurrence and development of NB through inhibiting TOB1 expression.
MiR-25 belongs to the miR-106b family, which is one of the recently discovered tumor-related miRNA molecules [21]. MiR-106b-25 family locates on Human chromosome 7 and encodes three mature miRNAs, including miR-106b, miR-25 and miR-93. MiR-106b and miR-93 have similar seed sequences and cancer-promoting gene function. However, miR-25 has a different sequence, suggesting it may have different functions [22]. MiR-25 is abnormally expressed in a variety of tumors and involved in the proliferation, invasion and metastasis of tumor. For example, Xiang et al. found that miR-25 was upregulated in Non-Small Cell Lung Cancer (NSCLC), and promoted cell proliferation and metastasis by regulating the expression of FBXW7 [20]. Chen et al. indicated that miR-25 could inhibit the apoptosis of NSCLC cells by regulating the expression of regulator of G-protein signaling 3 (RGS3) [23]. Peng et al. found that miR-25 could promote the proliferation and metastasis of gliomas by target regulating neurofilament (NEFL) gene [24]. These results demonstrate that miR-25 plays a role as oncogene. The present study indicated that miR-25 expression in NB tissues was obviously increased and positively correlated to the distant metastasis and clinical stages of the tumor. The expression of miR-25 in the metastases-derived NB cell line SH-SY5Y was obviously higher than that in the primary tumor cell line IMR32, suggesting miR-25 expression is correlated to the metastasis of NB. In vitro experiment showed that down-regulation of miR-25 expression could inhibit the proliferation, invasion and metastasis of SH-SY5Y, while upregulation of miR-25 could promote the proliferation, invasion and metastasis of IMR32. Although the results from cell lines could not reflect the actual status of in vivo tumor tissues, it still to some extent suggests that miR-25 may play an oncogene function in NB.
MiRNAs exert their biological functions through binding to the 3’-UTR region of target genes. We predicted the potential target gene of miR-25 by Targetscan (http://www.targetscan.org), and found that TOB1could be the potential target gene. TOB1 belongs to the Anti-Proliferative gene (APRO) family, containing TOB/BTG domain at the N-terminus, which is the main domain to exert biological functions [25]. It can regulate cell cycle through the Ras/MAPK signal pathway, and the combination with CCR4-Caf1 to form a complex and other ways. TOB1 expression in tumor is down-regulated, which will accelerate the cell cycle and promote tumor proliferation. Current researches [26] show that TOB1 could inhibit tumor cell proliferation. TOB1 is the transducer of ErbB-2 and exert functions through their interaction [26]. Wu et al. found that TOB1 could enhance the resistance to radiation therapy of breast cancer by regulating JNK and p38 signaling pathway [27]. Li et al. indicated that miR-25 could promote the proliferation, invasion and metastasis of gastric cancer [28]. Cheet al. also reported that overexpression of TOB1 could promote the resistance to radiation therapy of bronchial epithelial cells through MAPK/ERK pathway [29]. The above results indicate that TOB1 can inhibit tumor. However, whether TOB1 is mutated and its effects on NB are still not reported. In addition, there have been no reports of TOB1 mutation in any tumors, and whether TOB1 was regulated by miR-25 has not been reported either. In this study, the results demonstrated that miR-25 could regulate TOB1 expression in NB cells. Dual luciferase reporter assay further showed that miR-25 could directly bind to the 3’-UTR region of TOB1, indicating TOB1 is the direct target gene of miR-25.
Conclusions
In conclusion, miR-25 is upregulated in NB and NB cells. And, miR-25 can directly target TOB1. Additionally, miR-25 could promote the proliferation, invasion and metastasis of NB cells by regulating TOB1 expression.
Acknowledgement
None
Disclosures
All authors declare no financial competing interests.
All authors declare no non-financial competing interests.
References
- Daudigeos-Dubus E, Le Dret L, Bawa O, Opolon P, Vievard A, Villa I, Bosq J, Vassal G, Geoerger B. Dual inhibition using cabozantinib overcomes HGF/MET signaling mediated resistance to pan-VEGFR inhibition in orthotopic and metastatic neuroblastoma tumors. Int J Oncol 2017; 50: 203-211.
- Mari E, Mardente S, Morgante E, Tafani M, Lococo E, Fico F, Valentini F, Zicari A. Graphene oxide nanoribbons induce autophagic vacuoles in neuroblastoma cell lines. Int J Mol Sci 2016; 17: 1995.
- Gota V, Chinnaswamy G, Vora T, Rath S, Yadav A, Gurjar M, Veal G, Kurkure P. Pharmacokinetics and pharmacogenetics of 13-cis retinoic acid in Indian high-risk neuroblastoma patients. Cancer Chemother Pharmacol 2016; 78: 763-768.
- Murakami-Tonami Y, Ikeda H, Yamagishi R, Inayoshi M, Inagaki S, Kishida S, Komata Y, Jan K, Takeuchi I, Kondo Y, Maeda T, Sekido Y, Murakami H, Kadomatsu K. SGO1 is involved in the DNA damage response in MYCN-amplified neuroblastoma cells. Sci Rep 2016; 6: 31615.
- Sobhan PK, Zhai Q, Green LC, Hansford LM, Funa K. ASK1 regulates the survival of neuroblastoma cells by interacting with TLX and stabilizing HIF-1alpha. Cell Signal 2017; 30: 104-117.
- Turki S, Abouda M, Hachicha A, Ben Ghachem D. Neuroblastoma revealing cervical metastasis. Tunis Med 2016; 94: 164-165.
- Luessen DJ, Hinshaw TP, Sun H, Howlett AC, Marrs G, McCool BA, Chen R. RGS2 modulates the activity and internalization of dopamine D2 receptors in neuroblastoma N2A cells. Neuropharmacology 2016; 110: 297-307.
- Corroyer-Dulmont A, Falzone N, Kersemans V, Thompson J, Hill M, Allen PD, Beech J, Gilchrist S, Kinchesh P, Vojnovic B, Tullis I, Gaze MN, Smart S, Vallis KA. MRI-guided radiotherapy of the SK-N-SH neuroblastoma xenograft model using a small animal radiation research platform. Br J Radiol 2017; 90: 20160427.
- Rahman MA, Bishayee K, Habib K, Sadra A, Huh SO. 18alpha-Glycyrrhetinic acid lethality for neuroblastoma cells via de-regulating the Beclin-1/Bcl-2 complex and inducing apoptosis. Biochem Pharmacol 2016; 117: 97-112.
- Lee SH, Kim JS, Zheng S. ARID1B alterations identify aggressive tumors in neuroblastoma. Oncotarget 2017; 8: 45943-45950.
- Xu Y, Chen X, Lin L. MicroRNA-149 is associated with clinical outcome in human neuroblastoma and modulates cancer cell proliferation through Rap1 independent of MYCN amplification. Biochimie 2017; 139: 1-8.
- Srivastava S, Tsongalis GJ, Kaur P. Role of microRNAs in regulation of the TNF/TNFR gene superfamily in chronic lymphocytic leukemia. Clin Biochem 2016; 49: 1307-1310.
- Sorel O, Dewals BG. MicroRNAs in large herpesvirus DNA genomes: recent advances. Biomol Concepts 2016; 7: 229-239.
- Li Z, Xu Z, Xie Q, Gao W, Xie J, Zhou L. miR-1303 promotes the proliferation of neuroblastoma cell SH-SY5Y by targeting GSK3beta and SFRP1. Biomed Pharmacother 2016; 83: 508-513.
- Li Y, Shang YM, Wang QW. MicroRNA-21 promotes the proliferation and invasion of neuroblastoma cells through targeting CHL1. Minerva Med 2016; 107: 287-293.
- Liu Q, Wang Y, Yang T, Wei W. Protective effects of miR-25 against hypoxia/reoxygenation induced fibrosis and apoptosis of H9c2 cells. Int J Mol Med 2016; 38: 1225-1234.
- Song J, Li Y. miR-25-3p reverses epithelial-mesenchymal transition via targeting Sema4C in cisplatin-resistance cervical cancer cells. Cancer Sci 2017; 108: 23-31.
- Deng T, Yuan Y, Zhang C, Yao W, Wang C, Liu R, Ba Y. Identification of circulating mir-25 as a potential biomarker for pancreatic cancer diagnosis. Cell Physiol Biochem 2016; 39: 1716-1722.
- Petrocca F, Visone R, Onelli MR, Shah MH, Nicoloso MS, de Martino I, Iliopoulos D, Pilozzi E, Liu CG, Negrini M, Cavazzini L, Volinia S, Alder H, Ruco LP, Baldassarre G, Croce CM, Vecchione A. E2F1-regulated microRNAs impair TGFbeta-dependent cell-cycle arrest and apoptosis in gastric cancer. Cancer Cell 2008; 13: 272-286.
- Xiang J, Hang JB, Che JM, Li HC. MiR-25 is up-regulated in non-small cell lung cancer and promotes cell proliferation and motility by targeting FBXW7. Int J Clin Exp Pathol 2015; 8: 9147-9153.
- Jin Y, Yu D, Tolleson WH, Knox B, Wang Y, Chen S, Ren Z, Deng H, Guo Y, Ning B. MicroRNA hsa-miR-25-3p suppresses the expression and drug induction of CYP2B6 in human hepatocytes. Biochem Pharmacol 2016; 113: 88-96.
- He QY, Wang GC, Zhang H, Tong DK, Ding C, Liu K, Ji F, Zhu X, Yang S. miR-106a-5p suppresses the proliferation, migration, and invasion of osteosarcoma cells by targeting HMGA2. DNA Cell Biol 2016; 35: 506-520.
- Chen Z, Wu Y, Meng Q, Xia Z. Elevated microRNA-25 inhibits cell apoptosis in lung cancer by targeting RGS3. In Vitro Cell Dev Biol Anim 2016; 52: 62-67.
- Peng G, Yuan X, Yuan J, Liu Q, Dai M, Shen C, Ma J, Liao Y, Jiang W. miR-25 promotes glioblastoma cell proliferation and invasion by directly targeting NEFL. Mol Cell Biochem 2015; 409: 103-111.
- Shapouri F, Saeidi S, de Iongh RU, Casagranda F, Western PS, McLaughlin EA, Sutherland JM, Hime GR, Familari M. Tob1 is expressed in developing and adult gonads and is associated with the P-body marker, Dcp2. Cell Tissue Res 2016; 364: 443-451.
- Lee HS, Kundu J, Kim RN, Shin YK. Transducer of ERBB2.1 (TOB1) as a tumor suppressor: a mechanistic perspective. Int J Mol Sci 2015; 16: 29815-29828.
- Wu D, Zhou W, Wang S, Zhou Z, Chen L. Tob1 enhances radiosensitivity of breast cancer cells involving the JNK and p38 pathways. Cell Biol Int 2015; 39: 1425-1430.
- Li BS, Zuo QF, Zhao YL, Xiao B, Zhuang Y, Mao XH, Wu C, Yang SM, Zeng H, Zou QM, Guo G. MicroRNA-25 promotes gastric cancer migration, invasion and proliferation by directly targeting transducer of ERBB2, 1 and correlates with poor survival. Oncogene 2015; 34: 2556-2565.
- Che J, Lu YW, Sun KK, Feng C, Dong AJ, Jiao Y. Overexpression of TOB1 confers radioprotection to bronchial epithelial cells through the MAPK/ERK pathway. Oncol Rep 2013; 30: 637-642.