Research Article - Biomedical Research (2018) Volume 29, Issue 14
Prevalence of multidrug-resistant Salmonella typhi in typhoid patients and detection of blaCTX-M2 and blaCTX-M9 genes in cefetoxime-mediated extended spectrum β-lactamase-producing Salmonella typhi isolates
Rahul Narasanna, Manjunath Chavadi, Kelmani Chandrakanth*Department of Biotechnology, Kalaburagi University, Kalaburagi, Karnataka, India
- *Corresponding Author:
- Kelmani Chandrakanth
Department of Biotechnology
Kalaburagi University
Karnataka, India
Accepted date: July 26, 2018
DOI: 10.4066/biomedicalresearch.29-18-794
Visit for more related articles at Biomedical ResearchAbstract
Objectives: Prevalence of multidrug-resistant (MDR) S. typhi in typhoid-positive patients, phenotypic screening and molecular detection of blaCTX-M2, and blaCTX-M9 ESBL coding genes.
Materials and Methods: Sixty-six of the 650 clinical blood samples collected have turned out to be positive for S. typhi. Antimicrobial susceptibility and MIC was done as recommended by CLSI. Phenotypic screening of ESBL production and the PCR amplification ESBL coding genes blaCTX-M2, and blaCTX-M9 was also achieved.
Results: Sixty-six out of 650 clinical blood samples have turned out to be positive for S. typhi with an incidence rate of 10.15%. Among 66 isolates (n=44) isolates were MDR. The significant increase in MIC was observed between 128-256 μg/ml to cefetoxime, among the isolates, 9 S. typhi isolates reported positive for cefetoxime-mediated ESBL production. The PCR amplification of ESBL coding genes blaCTX-M2 and blaCTXM-9 yielded amplicon size of 884 and 692 bp, respectively. The ciprofloxacin resistant and cefetoxime sensitive standard MTCC S. typhi 734 and S. typhi BST 63 strain showed absence of both blaCTX-M2 and blaCTXM-9 genes.
Conclusion: The study shows an increase in the incidence rate of MDR S. typhi infection and ESBL production in S. typhi clinical isolates.
Keywords
Salmonella typhi, Multidrug resistant (MDR), ESBL, blaCTX-M2, blaCTX-M9.
Introduction
Majority of Salmonella sp. causes various types of clinical syndromes in humans and have been broadly categorized into two groups. The first, enteric fever, is caused by Typhoidal salmonella (TS) like S. typhi (typhoid fever), S. paratyphi A, and S. paratyphi B (Paratyphoid fever) and are mostly transmitted through contaminated water and food. The second type of clinical syndromes like diarrhea and gastroenteritis are caused by non-typhoidal Salmonella serovars [1]. In India, enteric diseases are most prevalent in urban areas, with the incidence approaching one percent of the population annually in some endemic areas [2]. Earlier studies have shown that majority of enteric fevers are prevalent in most of the developing countries like India, Pakistan, Bangladesh, and several African nations. In India, some reports suggest the incidence of typhoid fever in urban slum areas was as high as 0.2% (2 typhoid positive patients per 1000 population) per year among children less than 5 years of age, and 0.51% (5.1 typhoid positive patients per 1000 population) per year among those under 10 years of age [3]. Incidence of typhoidal infection varies with age. Studies from Kolkata have shown that the incidence of typhoid fever is higher in persons with average age of 14.7 y [4]. In contrast, the incidence of paratyphoid fever is higher among persons mean age of 17.1 y [4]. A study conducted in north India shows that incidence ratio of typhoidal fever is higher in children between 5 and 12 y of age, and the ratio is far higher in children aged less than 5 y; 24.8% of cases per year were reported among the children aged less than 5 y [5].
Extended Spectrum Beta-Lactamases (ESBL) were originally recognized in the year 1983 [6]. They belong to group of amber-class A β-lactamase enzyme that potentially hydrolyze β-lactam antibiotics with an oximinino group such as thirdgeneration cephalosporin’s and aztreonem, and are completely inactivated by β-lactamase inhibitors. The screening of the positive ESBL production in S. enterica serovars was detected and reported from various countries like Italy, France, and Nepal [7]. The significant increase in the production of CTXM- β-lactamases, a family of ESBLs, preferably hydrolyze cefetoxime (CTX) has been recognized and reported with increasing frequency [8]. CTX-M type of ESBLs or cefotaximases encoded by blaCTX-M genes located in a plasmid or on the chromosome belongs to type of class A β- lactamases [9]. This widespread resistance mechanism across the most part of the globe, from Europe, Africa, Asia, South America, and most recently from North America with several reports of clinical isolates producing these lactamases [8,10].
The CTX-M-type ESBLs causing serious concerns by displaying significantly increased level of resistance to both cefetoxime (CTX) and ceftriaxone (CTR) than that to ceftazidime (Caz). Approximately, 40 various types of CTX-M enzymes have been identified, and classified into five groups based on their amino-acid sequence homology, subsequently, been reported in different enterobacteria [11]. The first cefotaximase to be reported in Salmonella sp. was CTX-M-2, subsequently, over ten other CTX-M enzymes have been identified and reported in different Salmonella serovars with S. enterica serovar typhimurium being the most frequent, producing CTX-M-type enzymes such as CTX-M-5 and CTXM- 15 [6,12,13].
Presently, ciprofloxacin is preferred as a drug of choice for the treatment of typhoid infections. Its frequent administration has resulted in dramatic emergence of S. typhi with reduced susceptibilities to fluoroquinolones [2]. A study conducted in Kolkata has given a strong evidence of fluoroquinoloneresistant S. typhi with the Minimum Inhibitory Concentration (MIC) as high as 16 μg/ml to ciprofloxacin and norfloxacin [14]. Fluoroquinolone resistance in India is similar in pattern to other Asian countries [15].
The principal objective of the conducted study was to determine the incidence of multidrug resistance (MDR) and screening of extended spectrum β-lactamase (ESBL) in S. typhi strains isolated from typhoid patients from Kalaburagi region, Karnataka, India, and PCR amplification of plasmid-encoded blaCTX-M2 and blaCTX-M9 genes majorly responsible for cefetoxime-mediated resistance in S. typhi isolates.
Materials and Methods
Isolation of S. typhi from clinical samples
In the present study, 650 blood samples were collected from typhoid-suspected patients confirmed by Widal test from Government Hospital, Kalaburagi, India and Pooja Diagnostic Centre, Kalaburagi, India during July 2013-December 2013. The samples were enriched in tryptic soy broth with 100 units of streptokinase to dissolve clot. They were processed and tested biochemically according to standard guidelines for S. typhi isolation [16]. The isolated S. typhi strains were screened in relation to Standard MTCC S. typhi 734 and genotypically confirmed by 16S rDNA sequencing at Chromous biotech laboratory, Bangalore, India, with universal primers.
Antimicrobial susceptibility testing
The S. typhi isolates and standard MTCC S. typhi 734 were tested for antibiotic susceptibility and resistance for various groups of antibiotics by Kirby-Bauer disk diffusion method as per the standard guidelines of Clinical Laboratory Standard Institute (CLSI) [17,18]. The 16 antibiotic disks (Himedia Pvt. Ltd., Mumbai, India) selected for this study are listed in Table 1. The zone of inhibition post incubation was measured using standard antibiogram scale.
| Sl. no | Antibiotic | Concentration (µg) |
|---|---|---|
| 1 | β-Lactams | |
| Ampicillin | 10 | |
| Azeotrenem | 30 | |
| 2 | Cephalosporins | |
| Cefixime | 5 | |
| Ceftriaxone | 30 | |
| Ceftazidime | 30 | |
| Cefetoxime | 30 | |
| 3 | Quinolones | |
| Nalidixic acid | 30 | |
| Ciprofloxacin | 5 | |
| Levofloxacin | 5 | |
| Gatifloxacin | 5 | |
| 4 | Cotrimoxizole | 25 |
| 5 | Aminoglycosides | |
| Streptomycin | 10 | |
| Amikacin | 30 | |
| 6 | Macroloides | |
| Azithromycin | 15 | |
| 7 | Tetracycline | 30 |
| 8 | Chloromphenicol | 30 |
Table 1: Antibiotics selected for the study.
Determination of MIC
MIC was determined by agar dilution method and broth dilution method following CLSI standard guidelines 2012 [18] of five selected antibiotics: ciprofloxacin, nalidixic acid, chloramphenicol, cefetoxime, ampicillin (commercially available as ciprofloxacin hydrochloride monohydrate), nalidixic acid-free acid, chloramphenicol, cefetoxime sodium salt, ampicillin sodium salt (Himedia laboratories, Mumbai, India). Stock solutions of the required antibiotics were prepared as follows: ciprofloxacin (2 mg/ml), nalidixic acid (2 mg/ml), chloramphenicol (1 mg/ml), cefetoxime (1 mg/ml), ampicillin (1 mg/ml) [19]. Significant MIC breaking points to the respective antibiotics were interpreted.
Screening of ESBL production in S. typhi (Extended spectrum ß-lactamases)
Combination disc method: ESBL production in S. typhi isolates was determined by combination disc method with reference to Drieux et al. [20] on MHA (Muller Hinton Agar) medium seeded with S. typhi test organism. A combination of cephalosporin’s/clavulanic acid, i.e. cefetoxime (30 μg) and ceftazidime (30 μg) in combination with clavulanic acid (30 μg +10 μg), was placed 30 mm apart as suggested in standard CLSI guidelines, 2012 [18] and incubated at 37°C for 24 h. A greater by 3 mm increase in zone diameter of either respective clavulanic acid combination discs versus its zone diameter when tested alone confirmed as a significant ESBL production.
Double disc synergetic test: Double disc synergy test is the confirmatory test to combination disc method and was performed by standard protocol of Drieux et al. [20]. This test was performed on MHA seeded with S. typhi isolates and impregnated with amoxiclav or augmentin (combination of amoxicillin with 10 μg of clavulanic acid) at the center surrounded by cephalosporin antibiotic discs like cefetoxime (30 μg), ceftazidime (30 μg), cefexime (30 μg) set 30 mm from the center (Standard CLSI Guidelines, 2012) and incubated at 37°C for 24 h. A significant enhancement in the inhibition zone with either one of the test antibiotics to Augmentin disc is confirms positive for ESBL production.
PCR detection of ESBL-resistant blaCTX-M2 and blaCTX-M9 genes
PCR analysis was carried out to detect plasmid-encoded blaCTX-M2 and blaCTX-M9 responsible for cefetoximemediated drug resistance in S. typhi. The primers, sourced from Chromous Biotech Pvt. Ltd, Bangalore, India, were for blaCTX-M2, 5’-CGGTGCTTAAACAGAGCGAG-3’ (Forward), blaCTX-M2 (reverse) 5’ CCATGAATAAGC AGCTGATTGCCC-3’, blaCTX-M9 (forward) 5’- GATTGACCGTATTGGGAGTTT-3’, blaCTX-M9 (reverse) 5’- CGGCTG GGTAAAATAGGTCA-3’ [21]. For PCR amplification, about 100 ng of DNA was added to 50 μl mixture containing 10 mM of dNTPs, 0.4 mM of each forward and reverse primer and 3 U of Taq polymerase in 10X PCR buffer containing 1.5 mM MgCl2. Amplification was performed in a Corbett CGI-96 thermocycler with cycling parameters comprising the following steps, initial denaturation at 94°C for 5 min, sequentially followed by, denaturation 94°C for 30 s, annealing at 50°C for 30 s, extension at 72°C for 1 min and final extension at 72°C for 10 min for 35 cycles [22]. Finally, after accomplishing amplification, DNA ladder of 100 bp and 500 bp (Bangalore Genei Pvt Limited, India) were used to determine the size of amplicons.
Results
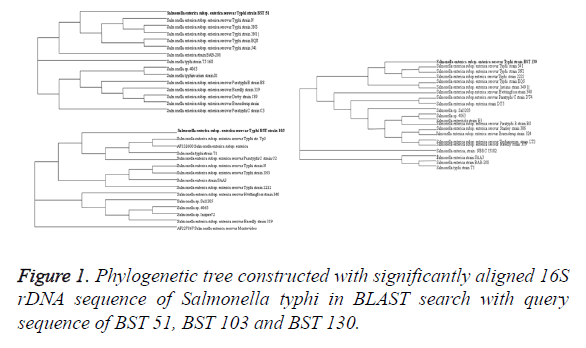
Of the total 650 samples, only 66 have shown culture positivity for S. enterica Serovar typhi with isolation rate of 10.15%. Results of 16S rDNA sequencing show 100% resemblance with S. typhi isolates in nucleotide BLAST analysis in comparison with existing databases. The sequences have been submitted to National Centre for Biotechnology Information through bankit and the accession number of three MDR isolates, namely, BST 51 (accession no. KR537431) BST 103 (accession no. KR537432) and BST 130 (accession no. KR537433) were obtained from the existing database through BLAST and phylogenetic tree constructed using bioinformatics tool (Figure 1).
Antimicrobial susceptibility testing
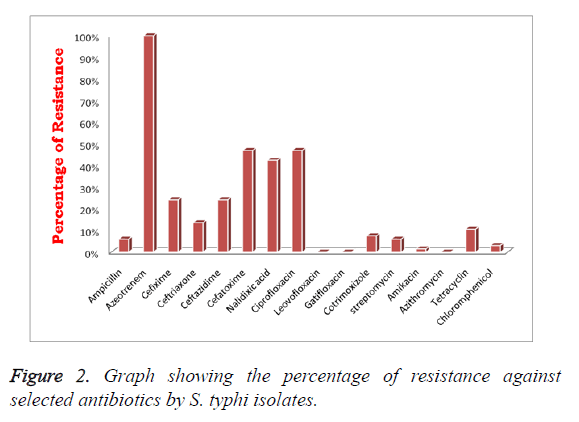
Sixty-six positive S. typhi isolates were chosen for this study. Among them, 06.06% (n=04) were ampicillin resistant but 100% (n=66) of isolates showed reduced susceptibility to aztreonem. In addition to that 24.25% (n=16) strains were resistant to cefexime, ceftazidime respectively, in contrast only 13.63% (n=09) to ceftriaxone. However, 46.97% (n=31) isolates were cefetoxime resistant, comparatively higher to other cephalosporin’s. Subsequently, 42.42% (n=28), 46.97% (n=31) of isolates shown resistance to nalidixic acid, ciprofloxacin respectively in contrast 100% (n=66) isolates were showed reduced resistance against higher-generation quinolones such as levofloxacin and gatifloxacin. In addition to that, all n=66 isolates were entirely sensitive to cotrimoxizole and azithromycin. We observed decreased resistance in S. typhi isolates to aminoglycoside antibiotics; streptomycin and amikacin, i.e., 06.06% (n=04), 1.5% (n=01) respectively (Figure 2). Only 10.60% (n=7) isolates were resistant to tetracycline and 03.03% (n=2) to chloramphenicol. However, standard MTCC S. typhi 734 strain showed resistance to ciprofloxacin, in contrast it was susceptible to cefetoxime, cefexime and ceftriaxone belongs to group of cephalosporin’s (Table 2).
| Sl. no | Antibiotic | No. of resistant strains | Percentage of resistance (%) | Percentage of intermediates (%) | Percentage of susceptibility (%) |
|---|---|---|---|---|---|
| 1 | β-lactams | ||||
| Ampicillin | 4 | 6.06 | 4.54 | 89.39 | |
| Aztreonem | 66 | 100 | 0 | 0 | |
| 2 | Cephalosporins | ||||
| Cefixime | 16 | 24.25 | 15.15 | 60.6 | |
| Ceftriaxone | 9 | 13.63 | 24.24 | 62.13 | |
| Ceftazidime | 16 | 24.25 | 28.78 | 46.97 | |
| Cefetoxime | 31 | 46.97 | 16.67 | 36.36 | |
| 3 | Quinolones | ||||
| Nalidixic acid | 28 | 42.42 | 28.79 | 28.79 | |
| Ciprofloxacin | 31 | 46.97 | 13.64 | 39.39 | |
| Levofloxacin | 0 | 0 | 1.5 | 98.5 | |
| Gatifloxacin | 0 | 0 | 1.5 | 98.5 | |
| 4 | Cotrimoxizole | 0 | 0 | 0 | 100 |
| 5 | Aminoglycosides | ||||
| Streptomycin | 4 | 6.06 | 21.2 | 72.73 | |
| Amikacin | 1 | 1.5 | 7.6 | 90.9 | |
| 6 | Macroloides | ||||
| Azithromycin | 0 | 0 | 9.09 | 90.91 | |
| 7 | Tetracycline | 7 | 10.6 | 7.6 | 81.8 |
| 8 | Chloromphenicol | 2 | 3.03 | 0 | 96.97 |
Table 2: Antibiotic resistance of Salmonella typhi isolates.
MIC of S. typhi isolates
Among n=31 ciprofloxacin-resistant strains; n=12 isolates showed a significantly increased MIC range of 128-256 μg/ml. Moreover, n=28 isolates have shown a remarkable increase in MIC range of 128-2048 μg/ml to nalidixic acid. However only one isolate S. typhi BST 174 has shown the maximum MIC of 512 μg/ml to chloramphenicol. Subsequently, strain S. typhi BST 43 has shown significantly increased MIC of 512 μg/ml to ampicillin and in contrast, S. typhi BST 45 and S. typhi BST 48 have shown MIC at 256 μg/ml. Consequently, n=4 strains, S. typhi BST 51, S. typhi BST 130, S. typhi BST 103, S. typhi BST 43, have shown a greater degree of MIC at 256 μg/ml to cefetoxime. In addition, the standard MTCC S. typhi 734 strain showed decreased MIC range to cefetoxime of 2 μg/ml and the high range of MIC of 128 μg/ml to ciprofloxacin (Table 3).
| S. no. | Antibiotics | MIC clinical breaking point (µg/ml) | MIC level (in µg/ml) | No. of resistant isolates |
|---|---|---|---|---|
| 1 | Ampicillin | 16-32 | 256-512 | 4 |
| 2 | Chloramphenicol | 8-32 | ≥ 512 | 2 |
| 3 | Cefotaxime | 8-32 | 16-256 | 31 |
| 4 | Nalidixic acid | 8-32 | 128-2048 | 28 |
| 5 | Ciprofloxacin | 1-4 | ≥ 128-256 | 31 |
Table 3: Minimum inhibitory concentration (MIC) level of Salmonella typhi isolates.
Phenotypic screening of ESBL production in S. typhi
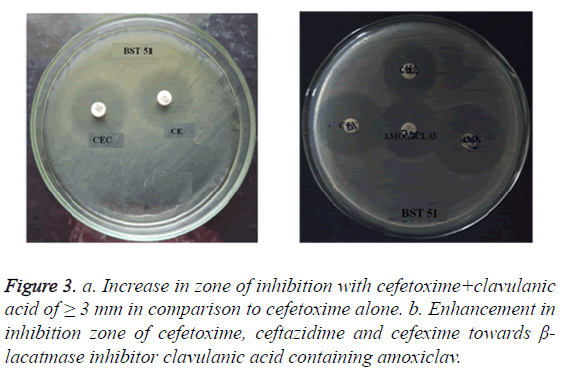
Of the 66 S. typhi isolates, 55% (n=36) showed resistance to third-generation cephalosporin’s such as ceftazidime, ceftriaxone and cefetoxime. Among 36 cephalosporin’s resistant S. typhi (CeRST) isolates, 25% (n=09) CeRST S. typhi isolates were positive ESBL producers. The ESBL production was evident in both combination disc method and double disc synergetic test. In addition to that entire n=9 S. typhi isolates were responsible for cefetoxime-mediated ESBL production (Figures 3a and 3b).
Detection of ESBL-resistant blaCTX-M2 and blaCTX-M9 genes
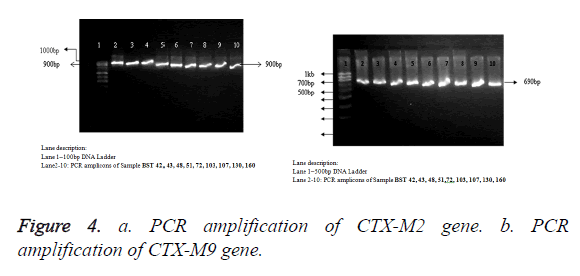
All the n=9 cefetoxime-mediated ESBL-producing S. typhi strains have been selected for genotypic study, i.e., PCR amplification of cefotaximase genes, blaCTX-M2 and blaCTXM9. The samples showed amplification of 884 bp and 692 bp, respectively (Figures 4a and 4b), with primers specific to blaCTX-M2 (accession no. KT277101) and blaCTX-M9 (accession no. KT277102). The blaCTX-M2 and blaCTX-M9 gene amplification was absent in ciprofloxacin resistant and cefetoxime sensitive standard MTCC S. typhi 734 and S. typhi BST 63.
Discussion
Enteric fever has caused a major concern to developing countries like India, Pakistan, Nepal and African countries. The incidence of typhoid fever is higher in developing countries compared to developed ones. The prevalence of MDR S. typhi and ESBL-producing S. typhi has now become a challenging task to the field of medicine. Earlier, chloramphenicol was used extensively as a drug of choice to treat typhoid fever. Its extensive prescription triggered the emergence of chloramphenicol-resistant S. typhi in Aden, Chile, and Kuwait [23]. The first epidemic was caused by chloramphenicol-resistant S. typhi in the year 1972 in Mexico [23,24] and subsequent outbreaks were reported from Kerala, India [25], Vietnam, Indonesia and Korea [23,26]. In 1989, the first outbreak, caused by multidrug-resistant S. typhi (MDRST), which is resistant to chloramphenicol, ampicillin, trimethoprim, sulphonamides, and tetracycline, was reported from developing countries like India [26-29], Pakistan [30,31] and Bangladesh [32]. In 1991, ciprofloxacin has been recommended for the treatment of typhoid fever caused by chloramphenicol-resistant S. typhi. In 1992, chromosomal encoded resistance to ciprofloxacin with MIC range of 0.3 mg/L was reported [33].
The present study evaluates the incidence of MDR S. typhi and cefetoxime-mediated ESBL production in S. typhi strains isolated from Kalaburagi City, Karnataka, India. Of the total of 650 Widal-positive samples collected, n=66 have shown positivity for S. typhi at the incidence rate of 10.15%, which is significantly higher in comparison with earlier studies conducted in this area [34]. Sixty-seven percent (n=44) of isolates were MDR and have shown resistance to more than three antibiotics. Antibiotic susceptibility test has shown 46.97% (n=31) strains having reduced susceptibility to ciprofloxacin; in contrast, to all n=66 highly susceptible to third- and fourth-generation antibiotics like levofloxacin and gatifloxacin. Decreased resistance of 7.15% (n=02) was observed in ampicillin, chloramphenicol, streptomycin and 10.6% (n=07) in tetracycline, respectively. Altogether, 66 isolates have shown 100% susceptibility to both cotrimoxizole and azithromycin. Interestingly, 54.54% (n=36) have shown remarkably reduced susceptibility to third-generation cephalosporin’s. The percentage of resistance is relatively higher in cefetoxime 86.11% (n=31), cefexime 44.44% (n=16), ceftriaxone 25% (n=9) and ceftazidime 44.44% (n=16) respectively.
Twelve out of 31 ciprofloxacin-resistant S. typhi isolates have shown an elevated level of MIC in the range of 128-256 μg/ml. There is a substantial increase in MIC level to cefetoxime with four (n=4) isolates in the range of 256 μg/ml; the result is relatively similar in comparison with the studies conducted in China by Yu et al. [35] on S. typhimurium. Four isolates have shown MIC to ampicillin in the range 256-512 μg/ml and also we found relatively higher range of MIC to chloramphenicol of 512 μg/ml.
Phenotypic detection of ESBL production was made against cefetoxime and ceftazidime by two standard phenotypic ESBL detection methods. Among n=36 CeRST isolates, 25% (n=9) strains have shown positivity for ESBL production; remarkably, all n=9 isolates were cefetoxime-mediated ESBL producers. Prevalence of ESBL production in India is far higher in comparison to the studies conducted in Nepal (0.5%) and Poland (0.3%) (36). ESBL production in S. typhi was genotypically confirmed by PCR amplification of blaCTX-M2 and blaCTX-M9 genes with amplicon size of 884 and 692 bp, respectively. The absence blaCTX-M2 and blaCTX-M9 amplification in cefetoxime sensitive standard MTCC S. typhi 733 and S. typhi BST 63 isolate indicates the significance of blaCTX-M2 and blaCTX-M9 genes in cefetoxime mediated ESBL production. The similar studies carried out by Xie et al. also showed the significance blaCTX-M genes in cefetoxime mediated drug resistance in E. coli isolates. Ryoo et al. [21] found nearly similar product length on Klebsiella pneumonia. Earlier studies already reported the presence of blaCTX-M2 gene in S. typhimurium and S. oranienberg [36-38].
Conclusion
Our study shows that the significant increase in the level of drug resistance among S. typhi isolates to higher-generation antibiotics like cephalosporin’s and higher-generation fluoroquinolones with higher level of MIC. Emergence of ESBL in S. typhi majorly in developing countries reflects poorly on sanitary conditions, inadequate medical attention, and contamination of water and food. Antibiotic susceptibility test and ESBL detection are recommended prior to treatment.
Conflict of Interest
The authors declare of no conflict of interest in conducting this study.
References
- Miller SI, Pegues DA. Salmonella species, including Salmonella typhi. Principles and Practice of Infectious Disease (5th Edn.). Churchill Livingstone New York 2000; 2344-2363.
- Bhan MK, Bahl R, Bhatnagar S. Typhoid and paratyphoid fever. Lancet 2005; 366: 749-762.
- Sur D, von Seidlein L, Manna B, Dutta S, Deb AK, Sarkar BL. The malaria and typhoid fever burden in the slums of Kolkata, India: data from a prospective community based study. Trans R Soc Trop Med Hyg 2006; 100: 725-733.
- Sur D1, Ali M, von Seidlein L, Manna B, Deen JL, Acosta CJ, Clemens JD, Bhattacharya SK. Comparisons of predictors for typhoid and paratyphoid fever in Kolkata, India. BMC Public Health 2007; 7: 289.
- Walia M, Gaind R, Paul P, Mehta R, Aggarwal P, Kalaivani M. Age related clinical and microbiological characteristics of enteric fever in India. Trans R Soc Trop Med Hyg 2006; 100: 942-948.
- Bauernfeind A, Stemplinger I, Jungwirth R, Ernst S, Casellas J M. Sequences of b-lactamase genes encoding CTX-M-1 (MEN-1) and CTX-M-2 and relationship of their amino acid sequences with those of other b-lactamases. Antimicrob Agents Chemother 1996; 40: 509-513.
- Al Naiemi N, Zwart B, Rijnsburger MC, Roosendaal R, Debets-Ossenko YJ, Mulder JA. Extended-spectrum-beta-lactamase production in a Salmonella enterica serotype typhi strain from the Philiines. J Clin Microbial 2008; 46: 2794-2795.
- Bonnet R. Growing group of extended-spectrum-lactamases: the CTX-M enzymes. Antimicrob Agents Chemother 2004; 48: 1-14.
- Rodriguez MM, Power P, Radice M, Vay C, Famiglietti A, Galleni M. Chromosome-encoded CTX-M-3 from Kluyvera ascorbata: a possible origin of plasmid borne CTX-M-1 derived cefotaximases. Antimicrob Agents Chemother 2004; 48: 4895-4897.
- Moland ES, Black JA, Hossain A, Hanson ND, Thomson KS, Pottumarthy S. Discovery of CTX-M-like extended-spectrum-lactamases in Escherichia coli isolates from five U.S. states. Antimicrob Agent Chemother 2003; 47: 2382-2383.
- Jin Y, Ling JM. CTX-M-producing Salmonella s. in Hong Kong: an emerging problem. J Med Microbiol 2006; 55: 1245-1250.
- Batchelor M, Hopkins K, Threlfall EJ, Clifton-Hadley FA, Stallwood AD, Davies RH. BlaCTX-M genes in clinical Salmonella isolates recovered from humans in England and Wales from 1992 to 2003. Antimicrob Agents Chemother 2005; 49: 1319-1322.
- Gazouli M, Tzelepi E, Sidorenko SV, Tzouvelekis, LS. Sequence of the gene encoding a plasmid-mediated cefotaxime-hydrolyzing class A beta-lactamase (CTX-M-4): involvement of serine 237 in cephalosporin hydrolysis. Antimicrob Agents Chemother 1998; 42: 1259-1262.
- Dutta S, Sur D, Manna B, Sen B, Bhattacharya M, Ochiai RL. Emergence of highly fluoroquinolone-resistant Salmonella enterica serovar typhi in community-based fever surveillance from Kolkata, India. Int J Antimicrob Agents 2008; 31: 387-389.
- Chau TT, Campbell JI, Galindo CM, Van Minh Hoang N, Diep TS, Nga TT. Antimicrobial drug resistance of Salmonella enterica serovar typhi in Asia and molecular mechanism of reduced susceptibility to the fluoroquinolones. Antimicrob Agents Chemother 2007; 51: 4315-4323.
- Farmer JJ, Davis BR, Hickman-Brenner FW, McWhorter A, Huntley-Carter GP, Asbury MA. Biochemical identification of new species and bio groups of Enterobacteriaceae isolated from clinical specimens. J Clin Microbiol 1985; 21: 46-76.
- Clinical and Laboratory Standards Institute Guidelines by CLSI/NCCLS-CLSI informational supplement. Aroved Standard M100-S15 Wayne PA 2005.
- Clinical and Laboratory Standards Institute. Performance standards for antimicrobial disk susceptibility tests; Aroved standard (11th Edn.). Clinical Laboratory Standards Institute Wayne PA 2012.
- Andrews JM. Determination of minimum inhibitory concentrations. J Antimicrob Chemother 2001; 48: 5-16.
- Drieux L, Brossier F, Sougakoff W, Jarlier V. Phenotypic detection of extended-spectrum b-lactamase production in Enterobacteriaceae: review and bench guide. Clin Microbiol Infect 2008; 14: 90-103.
- Ryoo NH, Kim EC, Hong SG. Dissemination of SHV-12 and CTX-M-type extended spectrum β-lactamases among clinical isolates of Escherichia coli and Klebsiella pneumoniae and emergence of GES-3 in Korea. J Antimicrob Chemother 2005; 56: 698-702.
- Jyoti S, Meera S, Pallab R. Detection of TEM and SHV genes in Escherichia coli and Klebsiella pneumoniae isolates in a tertiary care hospital from India. Indian J Med Res 2010; 132: 332-336.
- Anderson ES. The problem and implications of chloramphenicol resistance in the typhoid bacillus. J Hyg (Lond) 1975; 74: 289-299.
- Centers for Disease Control. Typhoid fever-Mexico. Morb Mortal Wkly Rep 1972; 21: 177-178.
- Paniker CKJ, Vilma KN. Transferable chloramphenicol resistance in Salmonella typhi. Nature 1972; 239: 109-110.
- Threlfall EJ, Rowe B, Ward LR. Occurrence and treatment of multi-resistant Salmonella typhi. Pub Health Lab Serv Microbiol Digest 1991; 8: 56-59.
- Anand AC, Kataria VK, Singh W, Chatterjee SK. Epidemic multi-resistant enteric fever in eastern India. Lancet 1990; 335: 352.
- Prakash K, Pillai PK. Multidrug resistant Salmonella typhi in India. Alliance Prudent Antibiot Newslett 1992; 10: 1-3.
- Threlfall EJ, Ward LR, Rowe B, Raghupathi S, Chandrasekaran V, Vandepitte J, Lemmens P. Widespread occurrence of multiple drug-resistant Salmonella typhi in India. Eur J Clin Microbiol Infect Dis 1992; 11: 990-993.
- Mandal BK. Treatment of multiresistant typhoid fever. Lancet 1990; 336: 1383.
- Karamat KA. Multiple drug resistant Salmonella typhi and ciprofloxacin. Proceedings of the 2nd Western Pacific Congress on Infectious Diseases and Chemotherapy. Jakarta, Indonesia: Western Pacific Society of Chemotherapy 1990; 480.
- Albert MJ, Haider K, Nahar S, Kibriya AK, Hossain MA. Multi-resistant Salmonella typhi in Bangladesh. J Antimicrob Chemother 1991; 27: 554-555.
- Rowe B, Ward LR, Threlfall EJ. Ciprofloxacin and typhoid fever. Lancet 1992; 339: 740.
- Kavita N, Shivannavar TC, Subhashchandra MG. Antimicrobial susceptibility of Salmonella typhi in India. J Infect Dev Ctries 2010; 4: 70-73.
- Yu F, Chen Q, Yu X, Li Q, Ding B. High prevalence of extended-spectrum beta lactamases among Salmonella enterica typhimurium Isolates from paediatric patients with diarrhoea in China. PLoS One 2011; 6: 16801.
- Szych J, Cieslik A, Paciorek J, Kaluzewski S. Antibiotic resistant in Salmonella enterica subspecies enterica strains isolated in Poland in the 1998-1999 periods. Med Dosw Microbial 2001; 53: 17-29.
- Maria AJ, Olga A, Ana T, Marta C. Extended-spectrum ß-lactamase-producing Salmonella enterica serovar oranienburg (CTX-M-2 group) in a pediatric hospital in Tucuman, Argentina. Rev Soc Bras Med Trop 2010; 43: 121-124.
- Xie M, Lin D, Chen K, chi Chan EW, Yao W, Chen S. Molecular characterization of E. coli strains isolated from retail meat that harbor blaCTX-M and fosA3 genes. Antimicrob Agents Chemother 2016.