Research Article - Biomedical Research (2018) Volume 29, Issue 2
MiR-100 inhibits invasion and migration of nasopharyngeal carcinoma cells by regulating ZBTB7A gene
Yi-Gang Zhao and Xian-Zhi Wang*
Department of Otorhinolaryngology, Linyi People’s Hospital, Linyi, Shandong, PR China
- *Corresponding Author:
- Xian-Zhi Wang
Department of Otorhinolaryngology
Linyi People’s Hospital, PR China
Accepted date: October 27, 2017
DOI: 10.4066/biomedicalresearch.29-17-2989
Visit for more related articles at Biomedical ResearchAbstract
Objective: This study aims to preliminarily investigate mechanisms of miR-100 in inhibiting invasion and migration of nasopharyngeal carcinoma cells by regulating ZBTB7A gene.
Methods: The nasopharyngeal carcinoma cell line CNE-2 was cultured and inoculated in a 12-well plate. Cells were divided into the overexpression group, interference group, and mock group, with four complex holes in each group. The medium was cultured for 24 h after every experiment. Invasive cells passing through Matrigel were detected using Transwell chamber, whereas cell migration rate of each group was recognized by conducting a 48 h scratch test. Meanwhile, expressions of miR-100 and ZBTB7A mRNA in each group were identified by using fluorescent quantitative polymerase chain reaction, whereas expression of ZBTB7A protein in each group was distinguished by administering Western blot.
Results: The number of invasive cells passing through Matrigel in the overexpression group was significantly lower than that of the interference and mock groups (P<0.05). Cell migration rate in the overexpression group was significantly lower (P<0.05) and showed no significant difference (P>0.05) with the remaining groups. Relative expression of miR-100 mRNA was the highest in the overexpression group (P<0.05), whereas that in the interference group was significantly lower than that of the mock group (P<0.05). Both expressions of ZBTB7A mRNA and protein in the overexpression group were significantly lower than those of the interference and mock groups (P<0.05). Expressions of ZBTB7A mRNA and protein in the overexpression group were significantly higher than those of the interference and mock groups (P<0.05).
Conclusion: We speculated that miR-100 inhibits invasion and migration of nasopharyngeal carcinoma cells through regulating ZBTB7A gene. However, the specific mechanism needs further study.
Keywords
Nasopharyngeal carcinoma, miR-100, ZBTB7A, Invasion, Metastasis
Introduction
Nasopharyngeal carcinoma is a highly malignant tumor in nasopharyngeal epithelium and is characterized by local and distant metastasis. This condition is common in South China and Southeast Asia [1]. However, the molecular mechanism of this condition has not been fully elucidated. Studies showed that microRNA (miRNA) plays an important regulatory role in occurrence, development, invasion, and metastasis of tumor [2]. Conversely, miR-100 is abnormally expressed in many tumors, mostly in the rectal and liver areas, and is closely related to occurrence and prognosis of cancer. Studies showed that ZBTB7A, which belongs to proto-oncogenes, is widely involved in tumor transformation, proliferation, and metastasis [3]. By regulating this gene, miR-100 can inhibit the spread of tumor cells. However, no study focused on its function in nasopharyngeal carcinoma. This experiment aimed to investigate movement of nasopharyngeal carcinoma cells through construction of overexpression miR-100 vector, interference miR-100 vector, and control vector. The present study also sought to detect expressions of ZBTB7A mRNA and protein and to preliminarily explore migration mechanisms of carcinoma cells in nasopharyngeal cavities.
Materials and Methods
Materials
The nasopharyngeal carcinoma cell line CNE-2 was obtained at Shanghai Institutes for Biological Sciences Cell Resource Center, China, whereas fetal bovine serum, Dulbecco’s Modified Eagle Medium (DMEM) and Roswell Park Memorial Institute (RPMI) 1640 culture medium, and cell digestive fluid were acquired in Gibco, USA. Total RNA extraction and reverse transcription kit were purchased in TAKARA, Japan; rabbit-anti-human ZBTB7A monoclonal antibody in Cell Signaling Technology, USA; horseradish peroxidase-labeled goat anti-mouse IgG in Santa Cruz, USA; and pre-miR-100, miR-100 inhibitor, and negative control at Shanghai Jikai Pharmaceutical Co. Ltd., China.
Cell groups and transfection
CNE-2 cells at logarithmic phase were digested and inoculated in a 12-well plate with 5 × 105 cell/hole. Cells were divided into three groups, the overexpression, interference, and mock groups, with four complex holes in each group. Cells in the overexpression group were transfected with vector pcDNA3.1- pre-miR-100, whereas those in the interference group were transfected with vector pcDNA3.1-IN-miR-100. Meanwhile, cells in the mock group were introduced with the empty plasmid pcDNA3.1. After reaching 50% cell fusion, 5 μL liposome and 100 pmol transfection vector were added in the culture plate. The process was indicated in kit instructions. The sample was incubated for 24 h for subsequent experiments.
TranswellTM was used to detect invasive cells passing through Matrigel
Matrigel was paved on the polycarbonate membrane and cultured in 5% CO2 incubator for 1 h at 37°C. Cells in each group were suspended with DMEM medium, concentration was adjusted to 5 × 105/hole, and cells were inoculated in the upper chamber. A 10% serum medium was added to the lower chamber as chemotactic agent. After 24 h incubation, cells in the upper chamber were cleaned, fixed with 4% paraformaldehyde for 30 min at room temperature, stained using 0.1% crystal violet for 20 min, and cleaned with phosphate-buffered saline (PBS) solution. Average value of cells passing through the membrane was calculated by five different fields of view under the microscope.
Scratch test was used to detect cell migration rate of each group
When cell fusion of each group reached 75% after transfection, a horizontal scratch was performed at the bottom of culture plate using a 200 μL pipette. The sample was washed using PBS and cultured using 5% fetal bovine serum culture medium. Width of scratch was measured at 0, 24, and 48 h under an inverted microscope.
Expressions of miR-100 and ZBTB7A mRNA in each group was detected by real-time quantitative polymerase chain reaction (PCR)
Following RNA kit instructions, CNE-2 cells in each group were collected for total RNA extraction using TRIzol method. RNA concentration was measured using a UV spectrophotometer. Total RNA was reverse-transcribed into cDNA. Amplification was performed by real-time fluorescent PCR. Primers for a) miR-100 amplification were as follows: upstream 5′-AGGAAGACGCCATACATGCG-3′, downstream 5′-GTCCACAGTGCGCACTAATG-3′; b) ZBTB7A amplification: upstream 5′-GGCGTCATGGACTACTACCT-3′, downstream 5′-CAGATGTTGCACTCGTAGGG-3′; and c) internal reference β-actin amplification: upstream 5′- TGGCATCCACGAAACTACCT-3′, downstream 5′- TGATCTTCATTGTGCTGGGTG-3′.
Statistical methods
The above results were analyzed using SPSS16.0 statistical software. Measurement data were expressed using mean and standard deviation. Pairwise comparison was performed using independent sample Dunnett’s t-test. P<0.05 indicated statistically significant difference.
Results
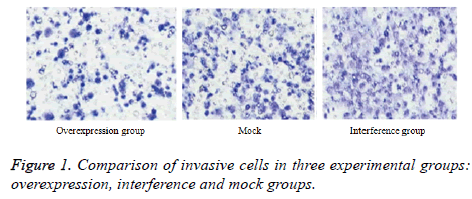
Invasive cells passing through Matrigel in each group
As shown in Figure 1, the lowest count of invasive cells was observed in the overexpression group (t=14.552, P=0.000; t=10.057, P=0.000). No significant difference was detected in the number of cells between the interference and mock groups (t=1.373, P=0.207).
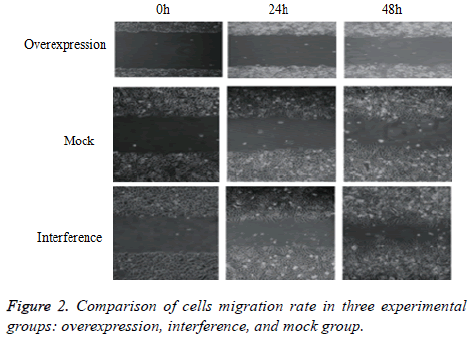
Cell migration rate in each group
As shown in Figure 2, cell migration rate in the overexpression group was significantly lower than those of the interference and mock groups in both 24 and 48 h intervals (24 h: t=6.677, P=0.000; t=4.401, P=0.002; 48 h: t=8.141, P=0.000; t=6.964, P=0.000). However, no significant difference was observed in cell migration rates between the interference and mock groups (24 h: t=2.193 P=0.060; 48 h: t=0.549, P=0.598).
Expressions of miR-100 and ZBTB7A mRNA in each group
Relative expression of miR-100 mRNA in the overexpression group was more significant than that of the interference and mock groups (t=10.025, P=0.000; t=12.935, P=0.000), whereas that of the interference group was less than that of the mock group (t=8.485, P=0.012). ZBTB7A mRNA in the overexpression group presented the lowest expression in comparison with other groups (t=7.765. P=0.000; t=13.868, P=0.000). Meanwhile, ZBTB7A mRNA expression was significantly higher in the interference group than that in the mock group (t=3.022, P=0.023).
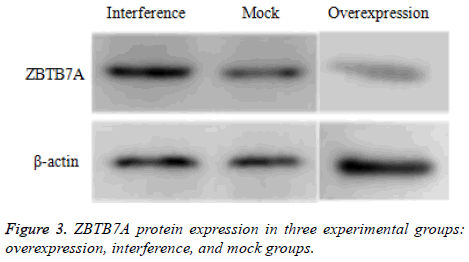
ZBTB7A protein expression in each group
As shown in Figure 3, the lowest expression of ZBTB7A protein was observed in the overexpression group (t=10.765, P=0.000; t=8.935, P=0.000). Meanwhile, expression of ZBTB7A protein was more significant in the interference group than the mock group (t=2.674, P=0.037).
Discussion
Transwell and cell scratch test revealed that after transfection of overexpressed miRNA-100, invasion and migration abilities of nasopharyngeal carcinoma cells decreased tremendously. While miRNA-100 was blocked, its invasion and migration abilities increased significantly, displaying its influence on metastasis of nasopharyngeal carcinoma [4]. Expressions of miR-100 and ZBTB7A mRNA in nasopharyngeal carcinoma cells were detected by fluorescence quantitative PCR. Results showed that up-regulation of miR-100 can decrease expression of ZBTB7A mRNA [5]. On the contrary, down-regulation of miR-100 can increase expression of ZBTB7A mRNA. ZBTB7A protein detection by Western blot indicated that overexpression of miR-100 can significantly down-regulate expression of ZBTB7A protein, whereas interference of miR-100 expression can significantly up-regulate expression of ZBTB7A [6]. These results suggest that ZBTB7A is a proto-oncogene in nasopharyngeal carcinoma, and up-regulation of miR-100 can inhibit expression of this proto-oncogene. Researchers performed a study on gastric cancer by using miR-100 in regulating ZBTB7A expression and discovered that overexpression of miR-100 can lower ZBTB7A protein expression [7].
Meanwhile, luciferase activity assay confirmed that ZBTB7A is the target gene of miR-100 in gastric carcinoma. ZBTB7A, also known as Pokemon, FBI-1, or osteoclast-derived zinc finger, belongs to the POK transcription factor family and is involved in gene transcriptional regulation of cell proliferation through protein ubiquitination in regulatory targets and signaling pathways [8]. Previous studies showed that ZBTB7A can also directly bind promoter regions and transcribe and inhibit key genes during glycolysis. For example, ZBTB7A can powerfully inhibit transcription of ADP-ribosylation factor (ARF) gene. ZBTB7A is also a proto-oncogene but functions as an antioncogene in prostate cancer [9,10]. Inactivation of ZBTB7A can activate the SOX9-dependent carcinogenic pathway to promote cancer cell proliferation and invasion. On the contrary, results of this study presented decreased expression of ZBTB7A, implying inhibition of invasion and migration of nasopharyngeal carcinoma cells. These findings also indicate that ZBTB7A gene can be regulated by different methods and can perform a variety of functions. Mechanisms of miR-100 in regulating ZBTB7A expression require further discussion.
Conclusion
To sum up, up-regulation of miR-100 can inhibit mobility of nasopharyngeal carcinoma cells. Expressions of both ZBTB7A mRNA and protein in experiments were minimized. Therefore, we speculated that decreased invasion and migration activities of nasopharyngeal carcinoma cells mainly depend on upregulation of miR-100 to inhibit ZBTB7A expression. However, related biological mechanism and signal pathway still require further studies.
References
- Chen P, Lin C, Quan J, Lai Y, He T, Zhou L, Pan X, Wu X, Wang Y, Ni L, Yang S, Wang T, Lai Y. Oncogenic miR-100-5p is associated with cellular viability, migration and apoptosis in renal cell carcinoma. Mol Med Rep 2017; 16: 5023-5030.
- Ruan FC, Dai J, Yan QH, Lu Y. Altered metabolic behavior of derivatives of herbal compounds used for treatment of carcinoma of mouth. Lat Am J Pharm 2017; 36: 1730-1735.
- Huang YL, Huang GY, Lv J, Pan LN, Luo X, Shen J. miR-100 promotes the proliferation of spermatogonial stem cells via regulating Stat3. Mol Reprod Dev 2017; 84: 693-701.
- Qin X, Yu S, Zhou L, Shi M, Hu Y, Xu X, Shen B, Liu S, Yan D, Feng J. Cisplatin-resistant lung cancer cell-derived exosomes increase cisplatin resistance of recipient cells in exosomal miR-100-5p-dependent manner. Int J Nanomedicine 2017; 12: 3721-3733.
- Latchoumi TP, Parthiban L. Abnormality detection using weighed particle swarm optimization and smooth support vector machine. Biomed Res-India 2017; 28: 4749-4751.
- Zhang XF, Chen Y, Wang JJ. Renal cell carcinoma treatment drug everolimus inhibited UDP-Glucuronosyltransferase (UGT) 1A6. Lat Am J Pharm 2017; 36: 1284-1288.
- Hsin CH, Huang CC, Chen PN, Hsieh YS, Yang SF, Ho YT, Lin CW. Rubus idaeus Inhibits Migration and Invasion of Human Nasopharyngeal Carcinoma Cells by Suppression of MMP-2 through Modulation of the ERK1/2 Pathway. Am J Chin Med 2017; 25: 1-16.
- Altin C, Orhan ER. Designing wearable joystick and performance comparison of EMG classification methods for thumb finger gestures of joystick control. Biomed Res-India 2017; 28: 4730-4736.
- Liu F, Lan J, Jiao W, Mo X, Huang Y, Ye H, Xiao R, Wang Y, Mo M, Shi L. Differences in Zbtb7a expression cause heterogeneous changes in human nasopharyngeal carcinoma CNE3 sublines. Oncol Lett 2017; 14: 2669-2676.
- Wang T, Dong XM, Zhang FL, Zhang JR. miR-206 enhances nasopharyngeal carcinoma radiosensitivity by targeting IGF1. Kaohsiung J Med Sci 2017; 33: 427-432.