Research Article - Biomedical Research (2017) Volume 28, Issue 11
Improved radiotherapy by survivin small hairpin RNA trigger apoptosis of human lung adenocarcinoma via caspase-3 upregulation in vitro and in vivo
Chang-Feng Li1, Yue Wang2, Xin-Ying Li3*, Jing-Peng Jin1, Bin Zhang1 and Dan-Dan Li11Endoscopy Center, China-Japan Union Hospital of Jilin University, Changchun, Jilin, PR China
2Department of Thoracic Surgery, China-Japan Union Hospital of Jilin University, Changchun, Jilin, PR China
3Department of Ultrasonography, China-Japan Union Hospital of Jilin University, Changchun, Jilin, PR China
- *Corresponding Author:
- Xin-Ying Li
Departments of Ultrasonography
China-Japan Union Hospital of Jilin University, PR China
Accepted date: March 23, 2017
Abstract
Purpose: RNA interference (RNAi) of survivin gene can induce cancer apoptosis. In the current study, combined survivin shRNA (small hairpin RNA) and radiotherapy is expected to enhance the efficacy of radiotherapy in human lung cancer in vitro and in vivo.
Methods: The pGenesil2 expression vectors containing survivin shRNAs was used to transfect A549 human lung adenocarcinoma cells and nude mouse grafts, followed by 5 Gy X-rays radiation. The MTT assay was performed to determine cell proliferation. Flow cytometry was used to detect the apoptosis of A549 cells. Tumor volumes were measured. Quantitative real-time PCR and Western blotting were performed to detect both mRNA and protein levels of survivin and caspase-3 in both A549 cells and nude mouse grafts.
Results: A549 cell proliferation and nude mouse grafts were inhibited in the shRNA alone, 5 Gy alone and shRNA+5 Gy groups, in particular the shRNA+5 Gy group. Apoptotic percentage increased in the 5 Gy, shRNA and shRNA+5 Gy groups, in particular the shRNA+5 Gy group. Survivin mRNA and protein levels in both A549 cells and nude mouse grafts decreased in the shRNA alone and shRNA+5 Gy groups. Caspase-3 protein in both A549 cells and nude mouse grafts increased in the shRNA alone, 5 Gy alone and shRNA+5 Gy groups, in particular the shRNA+5 Gy group.
Conclusion: Combined survivin shRNA and X-ray radiation can enhance the efficacy of radiotherapy by promoting the apoptosis of lung adenocarcinoma via upregulation of caspase-3 in vitro and in vivo. Combined survivin-targeted genotherapy and radiotherapy may be a potential strategy for treating lung cancer.
Keywords
Survivin, RNA interference, Small hairpin RNA, Ionizing radiation, Radiotherapy, Apoptosis, Molecular targeted therapy, Caspase-3, Lung cancer, Nude mouse.
Introduction
Radiotherapy is an alternative method for treatment of lung cancer in clinic. The efficacy of such method is limited due to either low radiosensitivity or radioresistence of cancer cells. Survivin, a member of the Inhibitor of Apoptosis Protein (IAP), is considered as a radioresistence factor [1,2]. Survivin has few expressions in normal mature tissues and is highly expressed in embryonic tissues and mitotic cells [3]. It is also detected in the majority of cancers and resists apoptosis by regulating cancer cell mitosis, where suppressing survivin expression leads to an anti-tumor effect [4].
RNA interference (RNAi) of survivin gene can knockdown the survivin expression to induce cancer apoptosis [5-7]. Survivin siRNA can induce the apoptosis of A549 human lung cancer cells by activating caspase-9 [8]. Use of survivin siRNA can even reverse the cisplatin-resistance of human lung adenocarcinoma in vitro and in vivo; meanwhile, downregulation of Lung Resistance-related Protein (LRP) and bcl-2 as well as upregulation of caspase-3 protein is observed [9].
In recent years, inhibition of survivin gene using siRNA is found to improve the radiosensitivity of human Non-Small Cell Lung Cancer (NSCLC) cells, thus survivin knockdown is a promising strategy for the targeted therapy with an improved radiosensitivity [10]. Inhibition of survivin is an effective way to enhance the radiosensitivity in glioblastoma and laryngeal squamous cell carcinoma and oral squamous cell carcinoma [11-13]. However, more studies are needed to investigate the function of combined survivin RNAi and ionizing irradiation therapy as well as the underlying mechanisms in vitro and in vivo.
The current study constructed the small hairpin RNA (shRNA) of survivin gene into the eukaryotic expression vector pGenesil2, and then treated human lung adenocarcinoma in vitro and nude mouse grafts by combining survivin shRNA with ionizing irradiation therapy. The survivin inhibitory effect of shRNA interference, an improved radiotherapy efficacy and the pro-apoptotic mechanism were investigated. The current study intended to provide experimental evidence for a new way to improve the effects of radiotherapy by combining with survivin shRNA interference in lung cancer.
Materials and Methods
Materials
A549 human lung adenocarcinoma cells were purchased from Peking Union Medical University Cell Bank (Beijing, China) and were cultured in high glucose Dulbecco's Modified Eagle's Medium (Gibco BRL, Grand Island, NY) containing 10% Fetal Bovine Serum (FBS; Gibco BRL, Grand Island, NY) at 37ºC in 5% CO2. Twenty female BALB/c nude mice (4-6 weeks, weight 20 ± 2 g) were purchased from Slac Laboratory Animal Company (Shanghai, China) and fed in Jilin University Animal Experimental Center (Changchun, China). RNA interference (RNAi) vector pGenesil2 was purchased from Cell Marker Biotech., Inc. (Wuhan, China). Mouse anti-human survivin, caspase-3 and β-actin monoclonal antibodies (diluted by 1:1000 for final use) and the Enhanced Chemical Luminescence (ECL) kits were from Santa Cruz Biotech., Inc. (Santa Cruz, CA). Secondary polyclonal antibodies (diluted by 1:2000 for final use) were from ZSGB Bio. Company (Beijing, China). Lipofectamine™ 2000 and TRIzol reagent kit were from Invitrogen Life Technologies (Carlsbad, CA).
Construction of pGenesil2-survivin shRNA
The hairpin shRNA of survivin was designed. Briefly, according to the cDNA sequences of survivin in the NCBI GenBank (NM_001168), a 19-nt sequence was designed: GAAGCGTCTGGCAGATACT, followed by BLAST to confirm the specific sequence homology. Afterwards, two single-strand 69-nt oligonucleotides to construct the hairpin shRNAs were synthesized by Sangon Bio. Inc. (Shanghai, China). The hairpin shRNAs were constructed as follows: BamHI+Sense sequence+Loop (9 nt+Antisense sequence +TTTTTT (terminator)+KpnI+HindIII; the forward sequence was 5'-GATCCGAAGCGTCTGGCAGATACT TTCAAGAGA AGTATCTGCCAGACGCTTC TTTTTTGGTACCGGAAA-3', and the reverse sequence was 5'-AGCTTTTCCGGTACCAAAAAA GAAGCGTCTGGCAGATACT TCTCTTGAA AGTATCTGCCAGACGCTTC G-3'. The BamH I and Hind III restriction sites were used for the ligation with the plasmid pGenesil2. The Kpn I site that pGenesil2 did not carry was added for identifying the success of constructs. The forward sequence and reverse sequence annealed to form a double strand, which was ligated with the linearized plasmid pGenesil2 by restriction sites BamH I and Hind III using the T4 DNA Ligase, forming the expression vector pGenesil2- Survivin shRNA. The pGenesil2-Survivin shRNA was transformed into E. coli DH5α, followed by sequencing (Sangon Bio. Inc. Shanghai, China). The sequences were certified to be the same as they were expected. Scrambled shRNA control is synthesized in Sangon Bio. Inc. (Shanghai, China).
Cell transfection and irradiation
A549 cells were seeded into 24-well culture plates (Corning Costar, Lowell, MA) at a density of 1 × 105 cells per well, 0.5 ml in each well. Cells were tested in five separate groups: normal control with no treatment, scrambled shRNA control, pGenesil2-survivin shRNA (survivin shRNA group), 5 Gy Xrays radiation (5 Gy group) and combined pGenesil2-survivin shRNA+5Gy X-rays radiation (Combined group), 4 wells for each group. When cells grew to 80% confluence, transfection was performed using LipofectamineTM 2000 according to the instructions, 10 μg of plasmids were used and the ratio between plasmid and liposome was 1: 2.5. High-glucose DMEM containing 10% FBS was refreshed 6 h after transfection, and then cells were cultured for 24 h. X-rays irradiation was administrated using the Philips X-ray Deep Therapy machine at 200 kV and 10 mA, using 0.5 mm Cu and 1 mm Al plates at 50 cm skin distance. A dose of 5 Gy was delivered at a rate of 0.287 Gy/min. The selection of dose and dose rate followed the report from the United Nation Scientific Committee on the Effects of Atomic Radiation (UNSCAR) in 1986 [14,15]. Cells were cultured for 24 h. Then cells were tested by the MTT assay and flow cytometry (BD FACSCalibur, Becton Dickinson Co., Franklin Lakes, NJ).
A549 cell grafting in nude mouse and treatment
Nude mice underwent aseptic injection with 0.2 ml of A549 cells at a concentration of 5 × 107/ml into the armpit. When tumors grew to 5-8 mm diameter, the day was defined as 0 d for the following experiment. Mice were divided into four separate groups: scrambled shRNA control, pGenesil2- Survivin shRNA (Survivin shRNA group), 5 Gy X-rays radiation (5 Gy group) and combined pGenesil2-Survivin shRNA+5Gy X-rays radiation (Combined group), 5 mice in each group. In as aseptic isolator, each nude mouse underwent in situ injection with 40 μg of expression vectors, which were coated with 100 μl of DMEM and 10 μl of liposome 20 min prior to injection. After 24 h, nude mice were anesthetized. The tumor parts were exposed to 5 Gy X-rays radiation. The irradiation condition was described above. The tumor sizes of Length (L) and Width (W) were measured with a Vernier calliper at the predetermined day, and the tumor volume (mm3) was calculated according to the formula L × W2/2. At the 18th day, mice were sacrificed, the tumor masses were removed by surgery and stored for the detection of mRNA and protein.
MTT assay and flow cytometry
A total of 50 μl MTT (5 mg/ml dissolved in DMEM; Sigma Aldrich) was added in each well of the 24-well plates to form a final concentration of 0.5 mg/ml. Cells were incubated for 4 h. The supernatant was discarded, and 200 μl dimethylsulfoxide (DMSO, Sigma Aldrich) were added for 10 min. The Optical Density (OD) value was measured at 570 nm using a microplate reader (Bio Rad, Hercules, CA). For flow cytometric assay, A549 cells were collected, centrifuged at 800X g for 5 min, and washed twice by 0.01 mol/L phosphate buffered saline (PBS, pH=7.0). BD FACSCalibur flow cytometer (Becton Dickinson Company, Franklin Lakes, NJ) was used to detect cell apoptosis using the Propidium Iodide (PI) and Annexin V Fluorescein Isothiocyanate (FITC) double staining kit (Nanjing KGI BioTech Inc., Nanjing, China). Briefly, A549 cells were suspended in 190 μl of PBS. Then 5 μl Annexin V FITC and 5 μl PI were added at room temperature in the darkness for 10 min. Cell samples (1 × 104 cells in each sample) were analysed by flow cytometry. The BD CellQuest™ software (equipped with BD FACSCalibur flow cytometer, Becton Dickinson Co., Franklin Lakes, NJ) was used to acquire and analyse the data.
Quantitative real-time PCR (qPCR) of survivin mRNA expression
Total RNA was extracted from A549 cells or 0.1 g of nude mouse tumor masses using the TRIzol Reagent according to the manufacturer's instructions. The cDNA was synthesized using a reverse transcription kit (Takara Bio., Inc., Dalian, China) according to the manufacturer's instructions (200 ng RNAs, 20 μl for the PCR reaction system). The reaction conditions included 42ºC for 60 min, followed by 70ºC for 2 min. The Beacon Designer 7 software (Premier Biosoft Inc., Palo Alto, CA) was used to design the primers of human GAPDH and survivin genes according to their cDNA sequences in the NCBI Genebank. The human GAPDH primers as inner reference were: 5’-tat tgg gcg cct ggt cac ca-3’ (forward) and 5’-cca cct tct tga tgt cat ca-3’ (reverse); the survivin primers were 5’-gga cca ccg cat ctc tac at-3’ (forward) and 5’-gac aga aag gaa agc gca ac-3’ (reverse). These primers were synthesized by Takara Bio, Inc. (Dalian, China). The 2X SYBR real-time qPCR kit was purchased from Roche (Shanghai, China). The qPCR conditions were as follows: 94ºC 30 s, 62ºC 30 s, 72ºC 45 s, 40 cycles. The Stratagene Mx3000P™ Real Time QPCR System (Agilent Technologies, Beijing, China) were used for detection. Relative mRNA expression was calculated as the interest gene/GAPDH ratio.
Western blotting of survivin and caspase-3 proteins
A549 cells and 0.1 g of nude mouse tumor masses were added to 500 μl of lysis buffer (10 mmol/L Tris HCl, pH 7.4; 1 mmol/L EDTA, pH 8.0; 0.1 mol/L NaCl; 1 μg/ml aprotinin; 100 μg/ml phenylmethanesulfonyl fluoride) to extract the total protein. The Coomassie Blue method was performed to quantify the total protein. Proteins were separated on 12% gel using the SDS PAGE with a loading of 50 μg per lane. Proteins were electroblot to a nitrocellulose membrane (Bio Rad, Hercules, CA). The membrane was blocked using 1X TBST buffer containing 5% non-fat milk overnight at 4ºC. The primary monoclonal antibodies against either survivin, caspase-3 or β-actin were added, and incubated at room temperature for 1 h, respectively. The membranes were washed three times by 1X TBST buffer, 10 min for each time. The horseradish peroxidase conjugated secondary antibodies were added at 37˚C for 1 h. After washed, the Enhanced Chemical Luminescence (ECL) method was used to visualize the blots. The X-Ray film was exposed and the pictures were photographed using a gel image processing system (UVP EC3 600 Imaging System, UVP LLC, Upland, CA). The protein levels were presented as the relative interest protein/β-actin grayscale ratio, which was analysed using the software Image- Pro® Plus. Version 6.0. (Media Cybernetics, Inc., Silver Spring, MD).
Statistical analysis
Data were stated as the mean ± Standard Deviation (SD). The statistical software SPSS 12.0 (SPSS, Inc., Chicago, IL) was used for data analysis. One way analysis of variance and Duncan’ ad hoc test was performed. P<0.05 indicated statistically significant difference.
Results
A549 cell proliferation
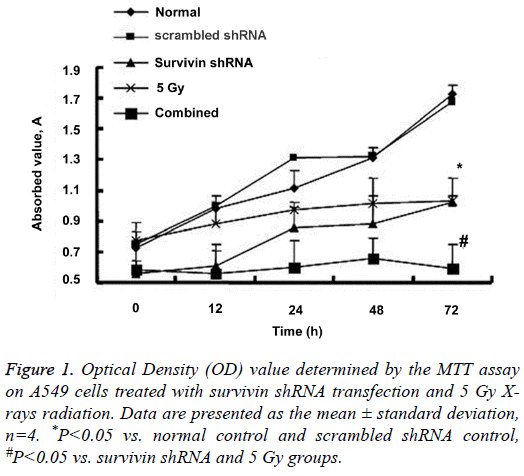
The MTT assay was used to assess A549 cell proliferation. Figure 1 shows the results for each group at time points of 1, 12, 24, 48 and 72 h. The OD values at 72 h (the final time point of cell culture) in the Survivin shRNA group and the 5 Gy group were significantly lower than the normal cell control and the scrambled shRNA control (P<0.05 for each). The OD value at 72 h in the combined group remained a very low level as low as that at 0 h, and was significantly lower than the Survivin shRNA group and the 5 Gy group (P<0.05 for each). No significant differences were observed between the Survivin shRNA group and the 5 Gy group or between the normal control and scrambled shRNA control (P>0.05). Compared with the normal control and scrambled shRNA control, Survivin shRNA and/or 5 Gy radiation treatment inhibited A549 cell proliferation markedly. In particular, in the combined group, use of Survivin shRNA enhanced the effects of radiotherapy on killing cancer cells.
Figure 1: Optical Density (OD) value determined by the MTT assay on A549 cells treated with survivin shRNA transfection and 5 Gy Xrays radiation. Data are presented as the mean ± standard deviation, n=4. *P<0.05 vs. normal control and scrambled shRNA control, #P<0.05 vs. survivin shRNA and 5 Gy groups.
Inhibition of nude mouse tumor growth
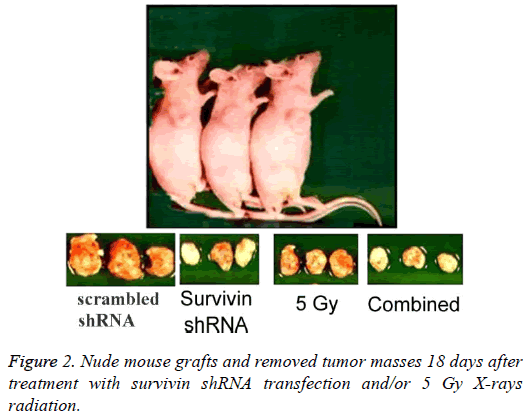
Figure 2 and Table 1 show the results of tumor growth. The tumors in all groups were enlarged in a time-dependent manner. The tumors in the scrambled shRNA control grew up to about 50-fold at 18 days versus those at 0 day. By comparison, each therapeutic treatment lowered the tumor growth markedly, and the order for the effects of lowering tumor growth was: the combined>Survivin shRNA>5 Gy>scrambled shRNA control, and the differences were statistically significant at the time points of 6, 12 and 18 days (P<0.05 or P<0.001). In particular, the tumor volume at 18 days in the combined group was only about 5.7-fold versus that at 0 day. Use of Survivin shRNA enhanced the effects of radiotherapy on lowering tumor growth.
| Days | Tumor volume (mm3) | |||
|---|---|---|---|---|
| Scrambled shRNA | Survivin shRNA | 5 Gy radiation | Survivin shRNA +5 Gy | |
| 0 | 87.52 ± 23.22 | 81.34 ± 21.57 | 88.50 ± 29.42 | 101.32 ± 13.2 |
| 2 | 200.52 ± 43.32 | 100.26 ± 46.38 | 167.12 ± 67.39 | 106.97 ± 32.67 |
| 6 | 823.14 ± 123.67 | 211.38 ± 87.32** | 500.83 ± 90.28* | 212.37 ± 98.34**△ |
| 12 | 2671.17 ± 515.08 | 611.79 ± 87.34** | 991.23 ± 198.37* | 315.26± 100.34**△ |
| 18 | 4040.81 ± 447.95 | 813.08 ± 161.15** | 3024.18 ± 455.95* | 579.64 ± 106.34△△ |
Table 1: Nude mouse tumor volumes after treatment with shRNA transfection and/or 5 Gy radiation.
A549 cell apoptosis
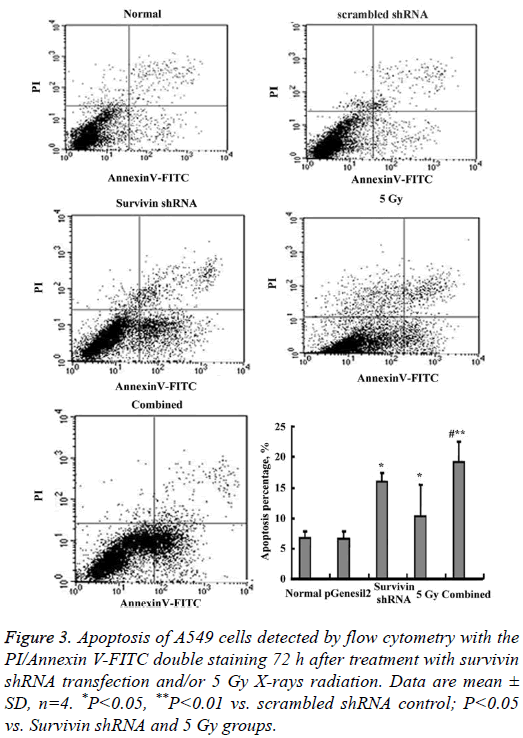
Figure 3 shows the results of A549 cell apoptosis detected by flow cytometry with the PI/AnnexinV-FITC double staining. The order for the apoptotic percentage in each group was: the combined>Survivin shRNA>5 Gy>scrambled shRNA control=normal control and the differences were statistically significant (P<0.05 or P<0.01). Particularly, the apoptotic percentage in the combined group (19.12 ± 3.48%) was about 3-fold versus the scrambled shRNA control (6.77 ± 1.04%).
Figure 3: Apoptosis of A549 cells detected by flow cytometry with the PI/Annexin V-FITC double staining 72 h after treatment with survivin shRNA transfection and/or 5 Gy X-rays radiation. Data are mean ± SD, n=4. *P<0.05, **P<0.01 vs. scrambled shRNA control; P<0.05 vs. Survivin shRNA and 5 Gy groups.
The mRNA and protein levels of survivin and/or caspase-3 in A549 cells
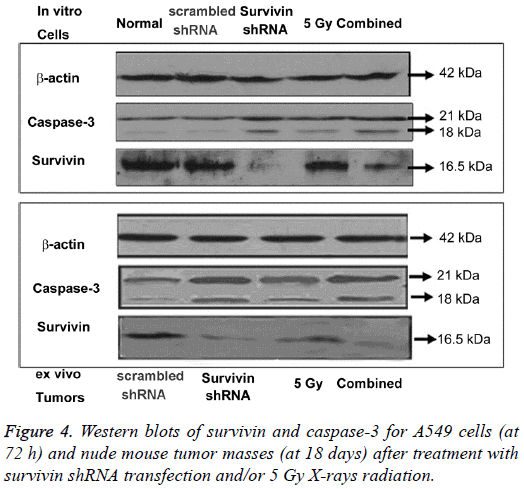
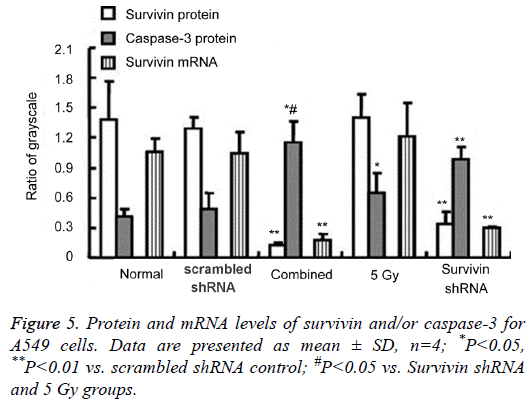
The protein and mRNA levels at 72 h for A549 cells are shown in Figures 4 and 5. Western blotting was performed to detect the survivin and caspase 3 protein expressions. The blots of housekeeping protein β actin (molecular weight, 42 kDa) were consistent in each group. The protein expression of survivin (16.5 kDa) decreased in both the survivin shRNA and combined groups, with the lowest expression observed in the combined group. The protein expression of survivin in the 5 Gy group increased, but no significant difference compared with the scrambled shRNA control. The protein expression of caspase-3 (21 kDa, 18 kDa) increased in either the survivin shRNA, 5 Gy or combined groups, with the highest levels for the combined group. The qPCR was performed to detect the survivin mRNA expression. Survivin mRNA levels in the survivin shRNA and combined groups were significantly decreased compared with those in the normal control and scrambled shRNA control (P<0.05 or P<0.01). The survivin shRNA group and the combined group had an apparent knockdown of survivin mRNA. Survivin mRNA levels in the 5 Gy group increased but no significant differences compared with the scrambled shRNA control. When compared with the 5 Gy group, the levels of pro-apoptotic caspase 3 protein increased in the combined group.
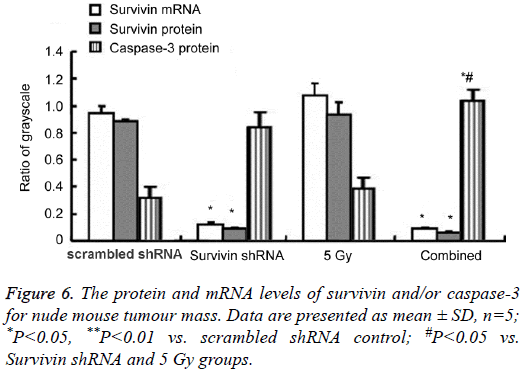
The mRNA and protein levels of survivin and/or caspase-3 in nude mouse tumor
The protein and mRNA levels at 18 days for nude mouse tumor are shown in Figures 4 and 6. The survivin protein (16.5 kDa) levels were downregulated in the survivin shRNA and combined groups, with the lowest expression for the combined group. The survivin protein in the 5 Gy group increased but no significant differences compared with the scrambled shRNA control. Survivin mRNA levels in the survivin shRNA and combined groups were significantly decreased compared with those in the scrambled shRNA control (P<0.05 or P<0.01), with the lowest mRNA levels for both the survivin shRNA group and the combined group. The caspase-3 (21 kDa, 18 kDa) protein expression increased in both the survivin shRNA, 5 Gy and combined groups, with the highest levels for the combined group. When compared with the 5 Gy group, the pro-apoptotic caspase 3 protein levels increased in the combined group.
Discussions
Tumor mass is a hypoxic focus, resulting in a therapy resistance. How to enhance the radiosensitivity of tumor mass is the interest of research. Survivin plays a role in promoting cell proliferation [16,17] and is highly expressed in tumor mass. Its inhibitor to knockdown the survivin is sufficient to induce the apoptosis of tumor cells [6,7]. Thus, survivin has become a hotspot target for anticancer therapy [18-24].
In the current study, the results of the MTT assay and the tumor volume measurement showed that 5 Gy X-rays alone can inhibit the growth of tumor cells and nude mouse grafts. Survivin shRNA had a stronger inhibitory effect than 5 Gy Xrays alone. Combined Survivin shRNA and 5 Gy X-rays had an apparently enhanced inhibitory effect versus each administration alone. These results indicate that combined Survivin shRNA and radiotherapy can improve the efficacy of treatment. Flow cytometry results showed that combined 5 Gy X-rays alone can promote A549 cell apoptosis. The Survivin shRNA alone increased the apoptotic percentage obviously, and the effect was superior to 5 Gy X-rays alone. Combined Survivin shRNA and 5 Gy X-rays had an enhanced apoptotic percentage, versus each administration alone. These results indicate that the combined therapy can elevate the effect of radiotherapy on inducing cell apoptosis.
To find the molecular evidence upon the ability to induce apoptosis and inhibit proliferation, we detected survivin and caspase-3. The 5 Gy X-rays alone led to an increased level of survivin in vitro and in vivo. This is consistent with the previous results [19]. The reason for this may be that the cells or tumor mass trigger the radioresistant signaling pathways long time after irradiation to increase the expression of survivin that repairs the radiation damage and promotes cancer survival. The Survivin shRNA alone and the combined therapy were observed to reduce the survivin gene expression in both cells and tumor masses. The use of survivin shRNA contributes to such effects because the survivin shRNA expression can act for a long term in cells or tumor masses. These results suggest that the ability of survivin to inhibit apoptosis and promote cell proliferation can be knockdown by survivin-targeted RNA interference, in particular for the combined therapy [11,12].
Survivin can directly inhibit the activity of caspase-3 the apoptosis effector and thus the blocking of survivin gene by a targeting inhibitor can induce cancer cell apoptosis [20-24]. In the current study, the expression levels of caspase-3 increased in both the 5 Gy, survivin shRNA and combined groups, in particular the combined therapy. This finding can exactly explain the apoptosis results of A549 cells detected by flow cytometry. Upregulation of caspase-3 resulting from use of survivin siRNA is involved in the reversion of cisplatinresistance of human lung adenocarcinoma [9]. In addition, survivin siRNA activates caspase-9 that induces the apoptosis of A549 human lung cancer cells [8]. Considering the cascade of caspases, the findings in the current study are consistent with these previous reports.
In conclusion, survivin-targeted shRNA can knockdown effectively the survivin gene in A549 human lung adenocarcinoma cell lines and the nude mouse grafts. Combined genotherapy and radiotherapy can enhance the anticancer effect of 5 Gy X-rays by inducing cancer cell apoptosis via activation of caspase-3. Combined survivintargeted genotherapy and radiotherapy is potential to become one of the effective methods for the treatment of lung cancer.
Declaration of Interests
I state the content and authorship of the submitted manuscript has been approved by all authors. All authors have seen and approved the final version of the manuscript. The manuscript is original work of author. All data, tables, figures, etc. used in the manuscript are prepared originally by authors. The manuscript has not been and will not be published elsewhere or submitted elsewhere for publication. There are no Conflicts of interests to be declared. There no issues pertinent to Ethics of experimentation/Informed consent.
References
- Rodel F, Hoffmann J, Distel L, Herrmann M, Noisternig T, Papadopoulos T, Sauer R, Rodel C. Survivin as a radioresistance factor, and prognostic and therapeutic target for radiotherapy in rectal cancer. Cancer Res 2005; 65: 4881-4887.
- Lei Y, Geng Z, Guo-Jun W, He W, Jian-Lin Y. Prognostic significance of survivin expression in renal cell cancer and its correlation with radioresistance. Mol Cell Biochem 2010; 344: 23-31.
- Colnaghi R, Connell CM, Barrett RM, Wheatley SP. Separating the anti-apoptotic and mitotic roles of survivin. J BiolChem 2006; 281: 33450-33456.
- Zaffaroni N, Pennati M, Daidone MG. Survivin as a target for new anticancer interventions. J Cell Mol Med 2005; 9: 360-372.
- Zhang YC, Taylor MM, Samson WK, Phillips MI. Antisense inhibition: oligonucleotides, ribozymes, and siRNAs. Methods Mol Med 2005; 106: 11-34.
- Ning S, Fuessel S, Kotzsch M, Kraemer K, Kappler M. siRNA-mediated down-regulation of survivin inhibits bladder cancer cell growth. Int J Oncol 2004; 25: 1065-1071.
- Uchida H, Tanaka T, Sasaki K, Kato K, Dehari H, Ito Y, Kobune M, Miyagishi M, Taira K, Tahara H, Hamada H. Adenovirus-mediated transfer of siRNA against survivin induced apoptosis and attenuated tumor cell growth in vitro and in vivo. MolTher 2004; 10: 162-171.
- Chen XQ, Yang S, Li ZY, Lu HS, Kang MQ. Effects and mechanism of downregulation of survivin expression by RNA interference on proliferation and apoptosis of lung cancer cells. Mol Med Rep 2012; 5: 917-922.
- Liu JL, Wang Y, Jiang J, Kong R, Yang YM. Inhibition of survivin expression and mechanisms of reversing drug-resistance of human lung adenocarcinoma cells by siRNA. Chin Med J (Engl) 2010; 123: 2901-2907.
- Yang CT, Li JM, Weng HH, Li YC, Chen HC, Chen MF. Adenovirus-mediated transfer of siRNA against survivin enhances the radiosensitivity of human non-small cell lung cancer cells. Cancer Gene Ther 2010; 17: 120-130.
- Reichert S, Rodel C, Mirsch J, Harter PN, Tomicic MT, Mittelbronn M, Kaina B, Rodel F. Survivin inhibition and DNA double-strand break repair: a molecular mechanism to overcome radioresistance in glioblastoma. RadiotherOncol 2011; 101: 51-58.
- Hu J, Pan J, Luo Z, Tao Z. Downregulation of survivin by shRNA inhibits invasion and enhances the radiosensitivity of laryngeal squamous cell carcinoma. Cell BiochemBiophys 2015; 72: 251-257.
- Sun HB, Zheng HY, Yan X. Survivin silencing enhances radiosensitivity in oral squamous cell carcinoma cell. Eur Rev Med PharmacolSci 2014; 18: 2678-2686.
- United Nations Scientific Committee on the Effects of Atomic Radiation. 1986 Report to the General Assembly, with annexes. General Assembly Official Records: Forty-first session, Supplement No. 16 (A/41/16). United Nations New York NY 1986.
- Liu G, Gong P, Zhao H, Wang Z, Gong S. Effect of low-level radiation on the death of male germ cells. Radiat Res 2006; 165: 379-389.
- Castedo M, Perfettini JL, Roumier T, Andreau K, Medema R, Kroemer G. Cell death by mitotic catastrophe: a molecular definition. Oncogene 2004; 23: 2825-2837.
- Wheatley SP, Henzing AJ, Dodson H, Khaled W, Earnshaw WC. Aurora-B phosphorylation in vitro identifies a residue of survivin that is essential for its localization and binding toinner centromere protein (INCENP) in vivo. J BiolChem 2004; 279: 5655-5660.
- Nam NH, Parang K. Current targets for anticancer drug discovery. Curr Drug Targets 2003; 4: 159-179.
- Asanuma K, Moriai R, Yajima T, Yagihashi A, Yamada M. Survivin as a radioresistance factor in pancreatic cancer. Jpn J Cancer Res 2000; 91: 1204-1209.
- Villapol S, Acarin L, Faiz M, Castellano B, Gonzalez B. Survivin and heat shock protein 25/27 colocalize with cleaved Caspase-3 in surviving reactive astrocytes following excitotoxicity to the immature brain. Neuroscience 2008; 153: 108-119.
- Hung HS, Wu WJ, Cheng YW, Wu TC, Chang KL, Lee H. Association of cooking oil fumes exposure with lung cancer: Involvement of inhibitor of apoptosis proteins in cell survival and proliferation in vitro. Mutat Res 2007; 628: 107-116.
- Shin S, Sung BJ, Cho YS, Kim HJ, Ha NC, Hwang JI, Chung CW, Jung YK, Oh BH. An anti-apoptotic protein human survivin is a direct inhibitor of Caspase-3 and -7. Biochemistry 2001; 40: 1117-1123.
- Zhang K, Li Y, Liu W, Gao X, Zhang K. Silencing survivin expression inhibits the tumor growth of non-small-cell lung cancer cells in vitro and in vivo. Mol Med Rep 2015; 11: 639-644.
- Li G, Xie B, Li X, Chen Y, Wang Q, Xu Y, Xu-Welliver M, Zou L. Down-regulation of survivin and hypoxia-inducible factor-1a by β-elemene enhances the radiosensitivity of lung adenocarcinoma xenograft. Cancer BiotherRadiopharm 2012; 27: 56-64.