Research Article - Biomedical Research (2018) Volume 29, Issue 3
Correlation analysis between polymorphism of GSTP1, XPD and XRCC1 gene in esophageal carcinoma patients and treatment efficiency of cisplatin chemo-therapy
Xu Li*, Huiyan Xiao, Rongqi He, Shaogeng Chen and Heshan Chen
Department of Surgical Oncology, the Affiliated First Hospital of Quanzhou, Fujian Medical University, Quanzhou, PR China
- *Corresponding Author:
- Xu Li
Department of Surgical Oncology
The Affiliated First Hospital of Quanzhou
Fujian Medical University, PR China
Accepted date: November 13, 2017
DOI: 10.4066/biomedicalresearch.29-17-1223
Visit for more related articles at Biomedical ResearchAbstract
Objective: As a common malignant tumor, esophageal carcinoma lacked specific clinical symptoms, leading to distal metastasis at terminal stage. Cisplatin is the first-line chemotherapy drug for the treatment of esophageal cancer. Identification of sensitive molecular marker is thus of critical importance for individualized treatment of cancer. This study analysed polymorphism of GSTP1, XPD and XRCC1 genes, aiming to illustrate its correlation with treatment efficacy of cisplatin on esophageal cancer, in order to provide evidences for individualized treatment effect.
Methods: A total of 104 esophageal carcinoma patients were recruited for follow-ups. Using Response Evaluation Criteria in Solid Tumor (RECIST), treatment efficacy was evaluated. PCR-RFLP was employed for detecting gene polymorphism.
Results: Frequency distribution of XRCC1 Arg399Gln, XPD Lys751Gln and GSTP1 Ile105Val all fitted Hardy-Weinberg equilibrium (p>0.05). Both one-factor and multi-factor analysis showed higher effective rate and survival time of patients with XRCC1 Arg399Gln GA+AA genotype after receiving cisplatin treatment compared to those patients with GG genotype (p<0.05). Patients with GSTA1 Ile105Val AG+GG had higher effective rate and survival time than individuals with AA genotype (p<0.05). No significant difference was observed among patients with different genotypes at XPD Lys751Gln locus (p>0.05).
Conclusions: Esophageal carcinoma patients with XRCC1 Arg399Gln GA+AA genotype and GSTP1 Il3105Val AG+GG genotype are more sensitive to cisplatin treatment.
Keywords
XRCC1, GSTP1, XPD, Esophageal carcinoma, Cisplatin, Gene polymorphism
Introduction
Esophageal Cancer (EC) is a common malignant tumor ranked as the sixth cause of morality worldwide. Recent survey showed that EC had the fourth high incidence and mortality among all malignant tumors in China [1]. In certain areas of China such as Henan Province, the incidence and mortality of EC are even higher than liver cancer, and ranked as the third deadly cancer [2], thus severely affecting public health.
Surgery is the major approach for treating EC. However, due to the infiltrative growth toward peripheral tissues/organs, the post-op survival rate maintains at a relatively low level. Meanwhile due to the lack of specific clinical manifestation, EC patients are usually at terminal stage or with metastasis at the time of confirmed diagnosis, making radical surgery impossible. Chemotherapy thus has become the major treatment way for EC. Combined drug chemotherapy mainly using platinum-based drug is the primary plan for EC treatment [3]. Cisplatin combined with 5-fluorouracil has become the classical plan for treating EC patients [4]. Individual difference in treatment efficacy is a common phenomenon in clinic, as mainly caused by unique genetic characteristics of patients [5]. Therefore, searching for molecular index in predicting drug sensitivity is of critical importance for treating EC.
Platinum-based drug can bind to tumor cell DNA to form complex, thus impeding DNA replication and blocking cancer progression. The self-repair mechanism of DNA by tumor cells, however, can weaken treatment efficiency of platinumbased drugs. Therefore, DNA repair potency is correlated with treatment efficacy [6]. Repair function related with platinum drug resistance is mainly consisted of nucleotide incision and base removal/repairmen [7-9]. Current findings revealed that the repair potency of human X-ray Repair Cross- Complementing gene 1 (XRCC1) was correlated with Single Nucleotide Polymorphism (SNP) at 399th locus, thus affecting chemotherapy efficiency of platinum based drugs [10-14]. Xeroderma pigmentosum group D (XPD) gene facilitates DNA repair by denaturing damaged site. Previous findings showed the close relationship between SNP of XPD gene at 751st locus and the sensitivity to platinum based drugs [15-17]. Moreover, study has found that Glutathione S-Transferase P1 (GSTP1) activity can be down-regulated by SNP of GSTP1 gene at 105th locus, thus weakening platinum drug sensitivity [18,19]. Currently few studies have been performed regarding the molecular markers guiding treatment efficiency of platinum based drugs in EC patients. This study thus examined gene polymorphism at these three SNP loci, in order to elucidate their correlations with EC treatment efficacy using platinum drugs, thus paving grounds for individualized treatment of EC.
Materials and Methods
Major equipment and reagent
Gel imaging analyzer (Alpha Innotech, Japan). Model 2720 PCR cycler (ABI, US). Model 22331 Hamburg protein/nucleic acid analyzer (Eppendorf, Germany). Electrophoresis apparatus, electrophoresis tank, model DYY-6C power (Liuyi Instrument, China). DNA extraction kit (TaKaRa, Japan). PCR amplification kit (TaKaRa, Japan). MspI, MboII and BsmAI restriction enzymes (TaKaRa, Japan).
Research objects
A total of 104 EC patients who were diagnosed in the Affiliated First Hospital of Quanzhou from January 1st 2011 to December 31st 2013 were recruited. The study was approved by the Ethic Committee of the Affiliated First Hospital of Quanzhou, Fujian Medical University, China. All participants had signed the informed consent form before enrolment.
Inclusive criteria: (1) EC diagnosis by pathology examination; (2) Not received platinum based drugs treatment previously; (3) Received EC treatment by combined chemotherapy mainly using platinum based drugs; (4) With complete clinical data; (5) Normal kidney and liver functions. After obtaining informed consent of patients, 5 ml fasted venous blood samples were collected before chemotherapy and were stored at -80°C fridge for further experiments.
Follow-up and treatment efficacy evaluation
Survival time was calculated since confirmation of the diagnosis. Follow-up was performed until December 31st, 2015 with 6~60 months length. No individual was lost during the follow-up. Basic information, treatment plan, treatment time period and efficacy were all collected. Treatment efficacy was evaluated based on Response Evaluation Criteria in Solid Tumor (RECIST) stipulated by European Cancer Research Institute and Treatment Society, National Cancer Institute, and Canadian National Cancer Research Institute. Treatment efficacy can be divided by complete response (CR): disappearance of all lesions and <10 mm dimension of all pathology lymph nodes; Partial Response (PR): ≥ 30% decrease of the summation of diameters of all lesions; Stable Disease (SD), no response or progression of disease; and Progression Disease (PD): ≥ 20% of lesion diameter summation, or >5 mm increase of absolute value of all diameters, or development of new lesions. Overall effective rate was calculated as (CR+PR)/(CR+PR+SD+PD).
Genetic polymorphism assay
Whole genomic DNA was extracted from peripheral blood. DNA purity and content was measured by protein-nucleic acid analyzer. Those samples with A260/A280 values between 1.8 and 2.0 were used for further experiments. Gene polymorphism was examined by PCR and PCR-restriction fragment length polymorphism (RFLP). Primers were designed and synthesized by Jierui (China) and were shown in Table 1. After amplifying targeted fragments by PCR, 5 μL PCR product was used for agarose gel electrophoresis to determine the successful amplification of target genes. MspI, MboII, BsmAI enzymes were then used to digest PCR products at polymorphism loci including XRCC1 Arg399Gln, XPD Lys751Gln and GSTP1 Ile105Val. 2% agarose gel electrophoresis was used for genotyping after enzymatic digestion.
| Polymorphism loci | Primer sequence (5′-3′) |
|---|---|
| XRCC1 Arg399Gln | F-TTGTGCTTTCTCTGTCTCCA |
| R-TCCTCCAGCCTTTCTGATA | |
| XPD Lys751Gln | F-CTCTCCCTTTCCTCTGTTC |
| R-CAGGTGAGGGGGATCTG | |
| GSTP1 Ile105Val | F-ACCCCAGGGCTCTATGGGAA |
| R-TGAGGGCACAAGAAGCCCCT |
Table 1. PCR primer sequence of target fragments.
Statistical analysis
SPSS19.0 software was used for statistical analysis. Chi-square test was employed to test if allele frequency fitted Hardy-Weinberg equilibrium and to compare parameters between two groups. Enumeration data were analysed by Logistic regression by multi-variate analysis. Survival analysis was assessed by Kaplan-Meier method. The comparison between survival rates was performed by Log-rank test. A statistical significance was defined when p<0.05. Test level α=0.05.
Results
Basic information of patients
The age of enrolled 104 patients age ranged from 35 to 73 y old with an average age of 58.89 ± 8.25 y old, including 72 cases older than 60 y (69.23%), and 32 patients (30.77%) younger than 60 y. There were 65 males (62.50%) and 39 females (37.50%). In an ECOG behavioral score, there were 17 cases (16.35%), 58 cases (55.77%) and 29 cases (27.88%) with 0, 1 and 2 scores. All patients belonged to squamous carcinoma, with 82 cases (78.85%) at moderate to high differentiation and 22 cases (21.15%) at low differentiation.
Genotype distribution
Table 2 showed genotyping analysis tested by PCR-RFLP. At XRCC1 Arg399Gln locus, there were 64 cases with GG genotype (61.54%), 32 cases with GA genotype (30.77%) and 8 cases with AA genotype (7.69%). At XPD Lys751Gln locus, there were 82 cases (78.85%) with AA genotype, 19 cases (18.27%) with AC genotype, 3 cases (2.88%) with CC genotype. At GSTP1 Ile105Val locus, there were 66 cases (63.64%) with GG genotype, 33 cases (31.73%) with GA genotype and 5 cases (4.81%) with AA genotype. By chisquare test, frequency distribution of alleles at all three SNP locus fitted Hardy-Weinberg equilibrium (p>0.05), indicating that enrolled patients had representativeness of the population.
| Polymorphism locus | Allele | Genotype | Allele | χ2 | P | |||
|---|---|---|---|---|---|---|---|---|
| AA (%) | AB (%) | BB (%) | A (%) | B (%) | ||||
| XRCC1 Arg399Gln | G/A | 64 (61.54) | 32 (30.77) | 8 (7.69) | 160 (76.92) | 48 (23.08) | 1.85 | 0.17 |
| XPD Lys751Gln | A/C | 82 (78.85) | 19 (18.27) | 3 (2.88) | 183 (87.98) | 25 (12.02) | 1.93 | 0.16 |
| GSTP1 Ile105Val | A/G | 66 (63.46) | 33 (31.73) | 5 (4.81) | 165 (79.33) | 43 (20.67) | 0.11 | 0.74 |
Table 2. Hardy-Weinberg equilibrium test for genotype frequency at polymorphism locus.
Correlation between gene polymorphism and treatment efficacy of chemotherapy
After combined therapy mainly using platinum-based drugs, treatment efficiency of all 104 patients were evaluated based on RECIST standard. There were 4 cases achieving CR, 58 PR, 31 SD and 11 PD, with overall effective rate of 59.61%.
Chi-square analysis was used to analyse the correlation between clinical pathology indexes of patients and gene polymorphism with treatment efficacy of platinum based drugs. As shown in Table 3, no significant correlations existed between sex, age, body status, Eastern Cooperative Oncology Group (ECOG) score and XPD Lys751Gln locus with effective rate of platinum treatment (p>0.05). Differentiation grade of tumors, polymorphism at XRCC1 Arg399Gln and GSTP1 Il3105Val locus was significantly correlated with the effective rate of platinum treatment (p<0.05). Those patients with higher differentiation grade of tumors, and those with XRCC1 Arg399Gln GA+AA and GSTP1 Ile105Val AG+GG carriers had remarkably higher effective rate than those with lower differentiated tumors, as well as those carrying XRCC1 Arg399Gln GG and GSTP1 Ile105Val AA (p<0.05).
| Index | n | PR+CR (n=62) | SD+PD (n=42) | Effective rate (%) | χ2 | P |
|---|---|---|---|---|---|---|
| (PR+CR)/n | ||||||
| Sex | ||||||
| Male | 65 | 37 | 28 | 56.92 | 0.52 | 0.47 |
| Female | 39 | 25 | 14 | 64.1 | ||
| Age | ||||||
| ≥ 60 y | 72 | 47 | 25 | 65.28 | 3.12 | 0.08 |
| <60 y | 32 | 15 | 17 | 46.88 | ||
| Differentiation | ||||||
| High | 82 | 56 | 26 | 68.29 | 12.12 | <0.001* |
| Low/middle | 22 | 6 | 16 | 27.27 | ||
| ECOG score | ||||||
| 0 | 17 | 12 | 5 | 70.59 | 1.04 | 0.59 |
| 1 | 58 | 33 | 25 | 56.9 | ||
| 2 | 29 | 17 | 12 | 58.62 | ||
| XRCC1Arg399Gln | ||||||
| GG | 64 | 32 | 32 | 50 | 6.39 | 0.01* |
| GA+AA | 40 | 30 | 10 | 75 | ||
| XPD Lys751Gln | ||||||
| AA | 82 | 50 | 32 | 60.98 | 0.3 | 0.59 |
| AC+CC | 22 | 12 | 10 | 54.55 | ||
| GSTP1 Ile105Val | ||||||
| AA | 66 | 34 | 32 | 51.52 | 4.92 | 0.03* |
| AG+GG | 38 | 28 | 10 | 73.68 |
Table 3. Correlation between clinical pathology indexes and gene polymorphism and efficacy of chemotherapy.
To invade possible confounding factors, unconditional Logistic regression was used for multi-variant analysis. As shown in Table 4, except sex, age, ECOG score and XPD Lys751Gln locus polymorphism, those patients with moderate or low differentiated tumor tissues, CRCC1Arg399Gln GG and GSTP1 Ile105Val AA carriers had lower possibility of beneficial effects (p<0.05).
| Parameter | P | OR | 95% CI |
|---|---|---|---|
| Sex | 0.76 | 1.17 | 0.44~31.2 |
| Age | 0.31 | 1.03 | 0.98~1.08 |
| Differentiation grade | 0.001* | 0.14 | 0.04~0.46 |
| ECOG score | 0.64 | 0.84 | 0.41~1.73 |
| XRCC1Arg399Gln GG/ GA+AA | 0.005* | 0.21 | 0.07~0.63 |
| XPD Lys751Gln AA/AC+CC | 0.42 | 1.62 | 0.51~5.22 |
| GSTP1 Ile105Val AA/AG+GG | 0.007* | 0.23 | 0.08~0.67 |
Table 4. Unconditional logistic regression analysis of factors for chemotherapy efficiency.
Relationship between post-chemotherapy survival time and genotype
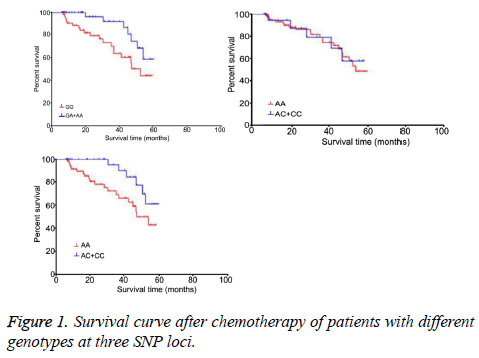
Kaplan-Meier method was used to analyse the correlation between survival time of patients after chemotherapy and genotypes. Setting the survival time as variable, and survival status as 0 whilst survive as 1. Analysis was then performed using genotypes at three loci as grouping variables. Log-rank test was employed for statistical analysis, in parallel with survival curve plotting. As shown in Figure 1, significant difference existed for survival time among different genotypes at XRCC1 Arg399Gln locus (χ2=4.17, p=0.04). Patients with GA+AA genotype had higher survival time than individuals with GG genotype. No significant difference existed for survival time among different genotypes at XPD Lys751Gl locus (χ2=0.03, p=0.86). Patients with AG+GG genotypes at GSTP1 Ile105Val loci had significantly longer survival time than those carrying AA genotype (χ2=4.35, p=0.03).
Screening of independent factors affecting patient survival by Cox regression analysis
In Cox multi-variant analysis, sex, age, differentiation grade, ECOG score and expression of three SNP loci were analysed. Results showed that differentiation grade, polymorphism at XRCC1 Arg399Gln and GSTP1 Ile105Val locus were independent risk factors affecting patient survival (Table 5).
| Affecting factor | Regression coefficient (B) | Standard error (S.E) | Wald | P | OR | 95% CI |
|---|---|---|---|---|---|---|
| Sex | 1.36 | 0.98 | 2.31 | 0.32 | 2.01 | 0.66-12.26 |
| Age | 1.6 | 1.11 | 2.85 | 0.17 | 2.43 | 1.46-11.22 |
| Differentiation | 1.11 | 0.28 | 15.58 | 0.025* | 3.06 | 1.69-5.56 |
| ECOG score | 1.17 | 1.26 | 1.22 | 0.3 | 3.15 | 0.41-13.65 |
| XRCC1Arg399Gln | 2.19 | 1.23 | 3.18 | 0.04* | 1.86 | 1.12-8.67 |
| XPD Lys751Gln | -0.94 | 1.31 | 0.67 | 0.4 | 0.3 | 0.06-3.13 |
| GSTP1Ile105Val | 2.18 | 0.3 | 3.78 | 0.02* | 3.34 | 1.20-4.33 |
Table 5. Analysis of independent risk factors affecting patient survival by Cox regression analysis.
Discussion
Most of EC patients already lost the chance of radical surgery, making chemo- or radio-therapy as major treatment approaches. Previous study revealed similar efficacy of synchronized chemo-and radio-therapy as those of surgery [20]. As the frontline medication for EC chemotherapy, platinum based drug is of critical importance. Therefore, identification of molecular index predicting drug sensitivity may help to guide the application of EC. Polymorphism of Excision Repair Cross-Complementing 1 (ERCC1) gene is correlated with treatment efficacy using platinum drug in EC patients [21]. However, few studies have been performed regarding the correlation between XRCC1, XPD and GSTP1 gene and platinum drug efficacy against EC. This study thus investigated the polymorphism of these three genes and their correlations with treatment efficacy using platinum in the treatment of EC.
Results showed higher effective rate and longer survival time in EC patients with XRCC1 Arg399Gln GA+AA and GSTP1 Ile105Val AG+GG genotypes, compared to those carrying XRCC1 Arg399Gln GG and GSTP1 Ile105Val AA patients, suggesting the possible correlation between gene polymorphism of XRCC1 Arg399Gln and GSTP1 Ile105Val and treatment efficacy using platinum drug for EC patients. Those patients with XRCC1 Arg399Gln GA+AA and GSTP1 Ile105Val AG+GG genotypes are more likely to be satisfied in platinum based chemotherapy. No significant difference was found for effective rate or survival time among different genotypes of XPD Lys751Gln, indicating the lack of significant correlation between XPD Lys751Gln polymorphism and treatment efficacy of cisplatin in EC patients. Similar reports can be found in a meta-analysis conducted by Gu et al. [10], who showed the correlation between XRCC1 Arg399Gln polymorphism and efficacy of platinum drug in treating Non- Small Cell Lung Cancer (NSCLC). Similar observations have been found in the treatment of colon cancer [14,22] and ovarian carcinoma [23] using platinum drugs. Kim et al. also found the correlation between polymorphism of Arg399Gln and GSTP1 Ile105Val locus with treatment efficacy of platinum on colorectal carcinoma [24]. However, the correlation between XPD Lys751Gln polymorphism and treatment efficacy of platinum treating tumors is still inconclusive. For example, Li et al. suggested a correlation between XPD Lys751Gln polymorphism and overall survival rate or progression-free survival time of NSCLC patients who received platinum drug treatment [15]. Kalikaki et al. however, reported the absence of such correlation [25]. Currently no correlation study has been reported regarding the polymorphism of XRCC1 Arg399Gln, GSTP1 Ile105Val and XPD Lys751Gln and treatment efficacy of platinum drugs against EC. Results from this study showed the correlation between XRCC1 Arg399Gln, GSTP1 Ile105Val polymorphism and efficacy of platinum drugs in treating various tumors, making them possible markers for individualized marker in treatment. The debate on XPD Lys751Gln loci, however, requires a large cohort clinical study to investigate.
In summary, this study showed better effective rate and longer survival time in EC patients carrying XRCC1 Arg399Gln GA +AA genotype after treatment using platinum drugs, and higher effective rate and longer survival time in those patients with GSTP1 Ile105Val AG+GG genotype compared to those with AA genotype. These results provided certain evidences for selecting chemotherapy treatment plan for EC patients. Large-scale clinical trial, however, is required for the application of using these two locus for guiding clinical treatment and drug selection.
Conclusion
EC patients with XRCC1 Arg399Gln GA+AA genotype and GSTP1 Ile105Val AG+GG genotype are more sensitive to platinum treatment.
References
- Chen W, Zheng R, Zeng H, Zhang S. The incidence and mortality of major cancers in China, 2012. Chin J Cancer 2016; 35: 73.
- Liu S, Chen Q, Quan P, Zhang M, Zhang S. Cancer incidence and mortality in Henan province, 2012. Chin J Cancer Res 2016; 28: 275-285.
- Ajani J, DAmico TA, Hayman JA, Meropol NJ, and Minsky B, Esophageal cancer. Clinical practice guidelines in oncology. J Natl Compr Canc Netw 2003; 1: 14-27.
- Oettle H, Arnold D, Kern M, Hoepffner N, Settmacher U, Neuhaus P, and Riess H, Phase I study of gemcitabine in combination with cisplatin, 5-fluorouracil and folinic acid in patients with advanced esophageal cancer. Anticancer Drugs 2002; 13: 833-838.
- Rodriguez-Vicente AE, Lumbreras E, Hernandez JM, Martín M, Calles A. Pharmacogenetics and pharmacogenomics as tools in cancer therapy. Drug Metab Pers Ther 2016; 31: 25-34.
- Rosell R, Cobo M, Isla D, Camps C, Massuti B. Pharmacogenomics and gemcitabine. Ann Oncol 2006; 17: 13-16.
- Jiao JW, Wen F. Tanshinone IIA acts via p38 MAPK to induce apoptosis and the down-regulation of ERCC1 and lung-resistance protein in cisplatin-resistant ovarian cancer cells. Oncol Rep 2011; 25: 781-788.
- Aloyz R, Xu ZY, Bello V, Bergeron J, Han FY, Yan Y, Malapetsa A, Alaoui-Jamali MA, Duncan AM, Panasci L. Regulation of cisplatin resistance and homologous recombinational repair by the TFIIH subunit XPD. Cancer Res 2002; 62: 5457-5462.
- Reed E. Platinum-DNA adduct, nucleotide excision repair and platinum based anti-cancer chemotherapy. Cancer Treat Rev 1998; 24: 331-344.
- Gu AQ, Wang WM, Chen WY, Shi CL, Lu JH. XRCC1 genetic polymorphisms and sensitivity to platinum-based drugs in non-small cell lung cancer: an update meta-analysis based on 4708 subjects. Int J Clin Exp Med 2015; 8: 145-154.
- Peng Y, Li Z, Zhang S, Xiong Y, Cun Y, Qian C, Li M, Ren T, Xia L, Cheng Y, Wang D. Association of DNA base excision repair genes (OGG1, APE1 and XRCC1) polymorphisms with outcome to platinum-based chemotherapy in advanced nonsmall-cell lung cancer patients. Int J Cancer 2014; 135: 2687-2696.
- Zhao W, Hu L, Xu J, Shen H, Hu Z, Ma H, Shu Y, Shao Y, Yin Y. Polymorphisms in the base excision repair pathway modulate prognosis of platinum-based chemotherapy in advanced non-small cell lung cancer. Cancer Chemother Pharmacol 2013; 71: 1287-1295.
- Zhang L, Ma W, Li Y, Wu J, Shi GY, Pharmacogenetics of DNA repair gene polymorphisms in non-small-cell lung carcinoma patients on platinum-based chemotherapy. Genet Mol Res 2014; 13: 228-236.
- Zaanan A, Dalban C, Emile JF, Blons H, Flejou JF, Goumard C, Istanbullu M, Calmel C, Alhazmi K, Validire P, Louvet C, de Gramont A, Laurent-Puig P, Taieb J, Praz F. ERCC1, XRCC1 and GSTP1 single nucleotide polymorphisms and survival of patients with colon cancer receiving oxaliplatin-based adjuvant chemotherapy. J Cancer 2014; 5: 425-432.
- Li P, Wang YD, Cheng J, Chen JC, Ha MW. Association between polymorphisms of BAG-1 and XPD and chemotherapy sensitivity in advanced non-small-cell lung cancer patients treated with vinorelbine combined cisplatin regimen. Tumour Biol 2015; 36: 9465-9473.
- Qin Q, Zhang C, Yang X, Zhu H, Yang B, Cai J, Cheng H, Ma J, Lu J, Zhan L, Liu J, Liu Z, Xu L, Sun X. Polymorphisms in XPD gene could predict clinical outcome of platinum-based chemotherapy for non-small cell lung cancer patients: a meta-analysis of 24 studies. PLoS One 2013; 8: 79864.
- Qian YY, Liu XY, Pei D, Xu JL, Shen H, Chen XF, Liu YQ, Shen LZ, Shu YQ. The XPD Lys751Gln polymorphism has predictive value in colorectal cancer patients receiving oxaliplatin-based chemotherapy: a systemic review and meta-analysis. Asian Pac J Cancer Prev 2014; 15: 9699-9706.
- Chen YC, Tzeng CH, Chen PM, Lin JK, Lin TC, Chen WS, Jiang JK, Wang HS, Wang WS. Influence of GSTP1 I105V polymorphism on cumulative neuropathy and outcome of FOLFOX-4 treatment in Asian patients with colorectal carcinoma. Cancer Sci 2010; 101: 530-535.
- Kumamoto K, Ishibashi K, Okada N, Tajima Y, Kuwabara K, Kumagai Y, Baba H, Haga N, Ishida H. Polymorphisms of GSTP1, ERCC2 and TS-3UTR are associated with the clinical outcome of mFOLFOX6 in colorectal cancer patients. Oncol Lett 2013; 6: 648-654.
- Diaz-Rubio E, Tabernero J, Gomez-Espana A, Massuti B, Sastre J, Chaves M, Abad A, Carrato A, Queralt B, Reina JJ, Maurel J, Gonzalez-Flores E, Aparicio J, Rivera F, Losa F, Aranda E. Phase III study of capecitabine plus oxaliplatin compared with continuous-infusion fluorouracil plus oxaliplatin as first-line therapy in metastatic colorectal cancer: final report of the Spanish Cooperative Group for the Treatment of Digestive Tumors Trial. J Clin Oncol 2007; 25: 4224-4230.
- Chen WH, Xin PL, Pan QX, Chen YY, Wang CR, Zhang ZS, Chen YF, Zhang CY, Cai WJ. ERCC1 single nucleotide polymorphism C8092A, but not its expression is associated with survival of esophageal squamous cell carcinoma patients from Fujian province, China. PLoS One 2014; 9: 106600.
- Gan Y, Li XR, Chen DJ, Wu JH. Association between polymorphisms of XRCC1 Arg399Gln and XPD Lys751Gln genes and prognosis of colorectal cancer in a Chinese population. Asian Pac J Cancer Prev 2012; 13: 5721-5724.
- Li K, Li W. Association between polymorphisms of XRCC1 and ADPRT genes and ovarian cancer survival with platinum-based chemotherapy in Chinese population. Mol Cell Biochem 2013; 372: 27-33.
- Kim DY, Paek TY, Oh SY, Kim YB, Lee JH, Lee MY, Choi ZS, Suh KW. Pretreatment selection of regimen according to genetic analysis improves the efficacy of chemotherapy in the first line treatment of metastatic colorectal cancer. J Surg Oncol 2014; 109: 250-254.
- Kalikaki A, Voutsina A, Koutsopoulos A, Papadaki C, Sfakianaki M, Yachnakis E, Xyrafas A, Kotsakis A, Agelaki S, Souglakos J, Mavroudis D, Georgoulias V. ERCC1 SNPs as potential predictive biomarkers in non-small cell lung cancer patients treated with platinum-based chemotherapy. Cancer Invest 2015; 33: 107-113.