- Journal of RNA and Genomics (2011) Research Report
RNA silencing of lactate dehydrogenase gene in Rhizopus oryzae
| Ali Hashemi Gheinani¥, Neda Haghayegh Jahromi¥, Elisabeth Feuk-Lagerstedt*, Mohammad J Taherzadeh University of Borås, School of Engineering, SE-501 90 Borås, Sweden ¥These authors contributed equally to this work |
| *Correspondence to: Elisabeth Feuk-Lagerstedt, Email: elisabeth.feuk@hb.se, Tel: +46 33 435 4710, Fax: +46 33 435 4008 |
| Received; 08 April 2011, Revised; 20 June; 2011, Accepted; 20 June 2011, Published online; 20 June 2011 |
| © Copyright The Authors: This is an open access article, published under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/2.0/uk/). This license permits noncommercial use, distribution and reproduction of the article, provided the original work is appropriately acknowledged with correct citation details. |
Abstract
Rhizopus oryzae is a filamentous fungus, belonging to the order Mucorales. It can ferment a wide range of carbohydrates hydrolyzed from lignocellulosic materials and even cellobiose to produce ethanol. However, R. oryzae also produces lactic acid as a major metabolite, which reduces the yield of ethanol. In this study, we show that significant reduction of lactic acid production could be achieved by short (25nt) synthetic siRNAs targeting the ldhA gene. The average yield of lactic acid production by R. oryzae during the batch fermentation process, where glucose had been used as a sole carbon source, diminished from 0.07gm/gm in wild type to 0.01gm/gm in silenced samples. In contrast, the average yield of ethanol production increased from 0.39gm/gm in wild type to 0.45gm/gm in silenced samples. These results show 85.7% (gm/gm) reduction in lactic acid production as compared with the wild type R. oryzae, while an increase of 15.4% (gm/gm) in ethanol yield.
Keywords |
| siRNA, delivery, Rhizopus oryzae, ldhA gene, lactate dehydrogenase, lactic acid, ethanol production |
Introducation |
| Lignocellulosic biomass is considered as the prime component for use as carbohydrate feedstock for industrial fermentation of chemicals. Currently, ethanol is the largest source of biofuel in the global market, which is produced either from sugar-based materials, such as sugarcane, or from starch-based materials, such as corn. However, there is an intensive global research effort to develop the process of ethanol production from lignocellulosic materials. This process involves pre-treatment of lingocelluloses, acid or enzymatic hydrolysis to extract sugars from cellulose and hemicelluloses, followed by their fermentation into ethanol. Enzymatic processes require two classes of enzymes, cellulase and β-glucosidase, as well as the microorganisms to produce ethanol: |
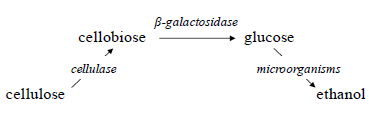 |
| Since optimum temperatures for enzymatic hydrolysis and fermenting organisms are generally different (45oC vs 30oC, respectively), the process is sometimes carried out by separate hydrolysis and fermentation (SHF).However, SHF suffers from the end-product (glucose) inhibition; therefore, another approach known as simultaneous saccharification and fermentation (SSF) carried out at a compromising temperature (e.g., 37oC) is preferred. |
| There are several microorganisms that can utilize lignocellulosic sugars and ferment them into numerous valuable products. R. oryzae is one such microbe; it is a saprophytic filamentous fungus and can up take a wide range of sugars such as, mannose, xylose, glucose, and galactose (Edebo, 2000), and the cellulose unit monomer cellobiose (Karimi et al, 2006). It has been largely used as a host microorganism to produce lactic acid (Skory, 2004a), but it can also produce a variety of other valuable metabolites, such as, gallic acid (Misro et al, 1997), lipase (Salah et al, 1994), protease (Tunga et al, 1999), cellulolytic enzymes (Amadioha, 1993), and ethanol (Abedinifar et al, 2009). It is a good alternative for ethanol production because of its tolerance to inhibitors in lignocellulose acid hydrolyzates (Karimi et al, 2006), its valuable material contents in the biomass, and its ability to grow at higher temperatures lowering the risk of contamination (Abedinifar et al, 2009). Thus, R. oryzae can utilize cellobiose quite well in the SSF processes: |
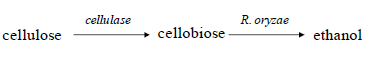 |
| However, ethanol yield by R. oryzae fermentation is relatively low because the fungus converts the sugar into lactic acid as a major by-product (Abedinifar et al, 2009). Therefore, it is conceivable to silence genes involved in lactic acid production to increase ethanol yield. |
| In RNA interference (RNAi), dsRNA facilitates degradation of the homologous mRNA, thereby diminishing or abolishing gene expression (Fire et al, 1998). However, despite numerous reports on RNA silencing in a variety of organisms, only a few species among filamentous fungi, such as Neurospora crassa and Cryptococcus neoformans have been shown to have RNAi machinery (Cogoni and Macino, 1999). Furthermore, RNA silencing has also been shown in Magnaporthe oryzae, the rice blast fungus by Naoki Kadotani and colleagues, using enhanced green fluorescence protein (eGFP) as a model (Kadotani et al, 2003). Skory (Skory, 2004b) tried several types of gene expression systems for R. oryzae to produce altered transcript that can form dsRNA of the ldhA gene. However, he was not able to achieve any significant reduction in lactic acid production for isolates containing short (20-25nt) synthetic ldhA RNAi in the expression plasmids. |
| The aim of this study was to address two main questions: (i) Can direct delivery of siRNAs results in silencing of the ldhA gene in R. oryzae? (ii) Can silencing of ldhA gene favorably increase ethanol production in the fermentation process considering the biochemical pathway of ethanol production in R. oryzae? |
Marerials and Methods |
| Fungal strain, growth media and spore solution |
| R. oryzae CCUG 28958 (Culture Collection, University of Gothenburg, Sweden) was used in all experiments. The strain was maintained at 4oC on potato dextrose agar (PDA) slants (potato extract 4gm/l, glucose 20gm/l, agar 15gm/l). For spore formation, a swab from the slant was taken and spread onto PDA agar plates. The plates were incubated at 30oC for 4 days. To prepare the inoculum, the agar plates containing sporulated fungi were washed with 20ml sterile water, and 20ml/l spore solution used in fermentation. |
| siRNA design |
| We targeted the region of ldhA with the highest homology with ldhB to silence both genes simultaneously in order to reduce chances of any bypass activity of ldhB. We used BLASTN 2.2.24+ (Zhang et al, 2000) to determine the regions of ldhA gene with high homology to ldhB gene. SiRNAs were then designed using these sequences with BLOCK-iT™ RNAi Designer (Invitrogen, Carlsbad, CA). The following sequences were used: |
| Sense: 5’GGAGGCAGGGCAGGCAGAUAUUGUU |
| Antisense: 5’AACAAUAUCUGCCUGCCCUGCCUCC |
| The target sequence was screened against the Broad Institute database (Ustilago Maydis Sequencing Project, Broad Institute of MIT and Harvard) to ensure that the sequences were specific for the target genes. |
| siRNA oligonucleotides (Invitrogen, Carlsbad, CA) were stored at -20oC until used. Duplex siRNAs (80nmol) were resuspended in 800μl DEPC treated water (Invitrogen, Carlsbad, CA), and then diluted 1:5 in TE buffer (10mM Tris-HCl, 20mM NaCl, 1mM EDTA, pH 8). |
| Fungal protoplast formation and transformation |
| Protoplast preparation and transformation in fungi has many limitations due to the wide variety of cell composition in different fungal species. Every so often the lytic enzymes used in the process do not digest the cell properly (Nakayashiki, 2008). We used the CaCl2/PEG (polyethylene glycol) method to transform R. oryzae with siRNAs, which has been used for some fungal species, such as Magnaporthe oryzae, Neurospora, Aspergillus, and Fusarium (Nakayashiki et al, 2005). This method was originally reported by Vollmer and colleagues (Vollmer and Yanofsky, 1986). Briefly, 150ml of semi-synthetic growth media was inoculated with 3ml of spore solution and incubated at 30oC and with shaking at 140rpm, for 3-4 days. Then R. oryzae mycelia were harvested by filtration through a Buchner funnel containing filter paper (Whatman No. 1). The mycelia pad was transferred to a plastic tube and 10ml of digestion buffer (10mM Na2HPO4, 1.2M MgSO4, 10mg/ml Lysozyme) (Sigma- Aldrich) was added per gram, wet weight of mycelia. The tube was incubated for 3hr at room temperature with gentle inversion on a horizontal shaker (New Brunswick Scientific, USA). The contents of the tube were passed through four layers of sterile gauze. The filtrate was centrifuged at 800xg for 5min (Megafuge 1.0R, UK), and the collected protoplasts were resuspended in 50ml STC buffer (1M Sorbitol, 50mM CaCl2, 50mM Tris-HCl, pH 8) (Nakayashiki, 2008). After another round of centrifugation as above the supernatant was discarded and the protoplasts were prepared. |
| Fungal transformation was carried out by mixing 100μl protoplasts and 100μl of 20μM siRNA solution. The mixture was incubated on ice for 15min. 1.2ml of PEG solution (60% (w/v) PEG 4000 (Sigma) in 50mM Tris- HCl, pH 8 and 50mM CaCl2) was added to the tube and mixed gently, followed by incubation on ice for 10min and centrifugation at 2000xg for 3min (Megafuge 1.0R). The supernatant was discarded and 1ml of STC buffer (1M Sorbitol, 50mM CaCl2, 50mM Tris-HCl, pH 8) was added to the protoplasts, which were poured onto the surface of a prepared PDA plate. The plates were incubated for at 30oC for 20hr. |
| Batch culture fermentation |
| In order to examine the efficiency of knockdown, fermentation was carried out with both wild type and transformed R. oryzae. Eight batch cultures were set up in 500ml Erlenmeyer flasks containing 150ml of semisynthetic growth media: 30gm/l glucose, 5gm/l yeast extract, 20gm/l (NH4)2SO4, 20gm/l KH2PO4, 5gm/l MgSO4.7H2O, 5gm/l CaCl2.2H2O, 10ml/l trace metal solution (3gm/l EDTA (C10H14N2Na2O8·2H2O), 0.9gm/l CaCl2·2H2O, 0.9gm/l ZnSO4·7H2O, 0.6 gm/l FeSO4·7H2O, 0.2gm/l H3BO3, 0.19gm/l MnCl2·4H2O, 0.08gm/l Na2MoO4·2H2O, 0.06gm/l CoCl2·2H2O, 0.06gm/l CuSO4·5H2O, 0.02gm/l KI), and 1ml/l vitamin solution (50mg/l d-Biotin, 200mg/l p-amino benzoic acid (PABA), 1gm/l Nicotinic acid, 1gm/l Ca-Panthothenate, 1gm/l Pyroxidine HCl, 1gm/l Thiamine HCl, 25gm/l m- Inositol, pH 5.5). The flasks were inoculated with 20ml/l spore solution and cultivated in a shaker incubator at 30oC with 140rpm for 62hr. |
| HPLC |
| The measurement of ethanol, lactic acid, and glucose was performed by High-Performance Liquid Chromatography (alliance/waters 2695, Waters, USA). The separation system consisted of a solvent delivery system equipped with an autosampler, a refractive index detector (410 differential refractometer, Waters), a dual λ absorbance detector (2487 Millipore, Milford, USA, Waters), and computer software (Waters Empower). A hydrogen-based ion-exchange column (Aminex HPX- 87H, Bio-Rad, USA) was used, and maintained at 65oC with 5mM H2SO4 eluent at a flow rate of 0.6ml/min. Refractive index (RI) detector and UV absorbance detector at 210nm were used in series. Concentrations of ethanol and glucose were determined by using the refractive index (RI) detector, while the lactic acid was quantified both with RI and UV absorbance detectors. |
Results and Discussion |
| Rationale behind efficient siRNA design |
| optimum region of the target gene is critical. It has already been reported that among two different genes (ldhA and ldhB), coding for NAD+-dependent L-lactate dehydrogenases (EC 1.1.1.27) in R. oryzae, ldhA gene plays the key role (Skory, 2000). Therefore, ldhA (Accession number AB111547) was chosen as the prime target for silencing. However, as we decided to inhibited both IdhA and IdhB to curtail any bypass activity of IdhB, we set out to discover regions of extensive homology between the two genes using the method of Zhang and Colleagues (Zhang et al, 2000). ldhA and ldhB exhibit more than 89% overall nucleotide sequence identity and contain no introns. The key sequence variation between the two genes lies at position 922-960 in ldhB, which ldhA lacks. Using these set of data, the siRNA was designed to target nucleotide region 198-222 in the ldhA gene, which targeted both ldhA and ldhB mRNAs. |
| Efficient R. oryzae protoplast formation |
| The cell wall of R. oryzae is mainly composed of chitin fibers cemented by chitosan and protein or peptides scattered in a mosaic manner (Zamani et al, 2008). Therefore, purified lytic enzymes, protease chitosanase, and chitinase were necessary for digestion of the cell wall. In this study, two different enzymes were investigated for degrading the cell wall of the R. oryzae; one based on Lysozyme from chicken egg white (Sigma-Aldrich, Fluka Biochemika, Switzerland) and another one based on Yatalase from culture supernatants of Corynebacterium (Takara Bio, Japan). The protoplast was then prepared, and the amount of protoplast and formation of mycelia were examined accordingly. The results revealed that Lysozyme from chicken egg white (Sigma-Aldrich, Fluka Biochemika, Switzerland) yielded less protoplast, however, growth of R. oryzae on the agar plate was faster in those lysed by Lysozyme (data not shown). Formation of R. oryzae protoplast by Lysozyme suggests that Lysozyme can break down the bond between N-acetyl-D-glucosamine residues in chitin present in the cell wall of R. oryzae. |
| Effect of ldhA gene silencing on lactic acid and ethanol production |
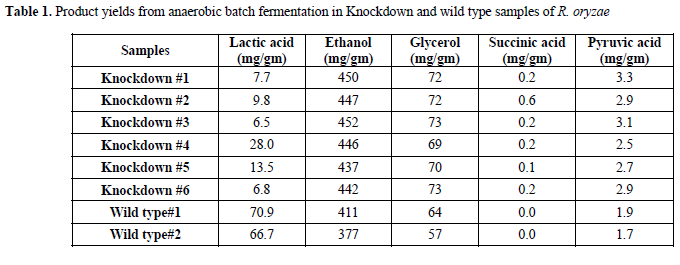
| After protoplast preparation by using Lysozyme (10mg/ml in digestion buffer), R. oryzae transformation with siRNA was carried out using the calcium chloride/polyethylene glycol (CaCl2/PEG) method. Different transformation systems, such as, electroporation, lithium acetate method, microparticle bombardment, and Agrobacterium tumefaciens-mediated transformation (ATMT) (Nakayashiki, 2008), have been tested in a variety of fungi. However in this experiment, the CaCl2/PEG transformation method was used with minor modifications in mucorales fungus R. oryzae. This method was examined previously for ascomycetes fungus Neurospora crassa (Vollmer and Yanofsky, 1986), Magnaporthe oryzae, and Colletotrichum lagenarium (Nakayashiki et al, 2005). The results of knockdown were investigated by running eight batch cultivations. The product yields obtained for all experiments are summarized in Table 1. |
| In six siRNA-inhibited samples, lactic acid production was reduced in comparison with the wild type strain. The average yield of lactic acid production was 0.01gm/gm in inhibited samples, while lactic acid production by wild type R. oryzae was 0.07gm/gm. It represents 85.7% (gm/gm) reduction in lactic acid production (Table 2). These results |
 |
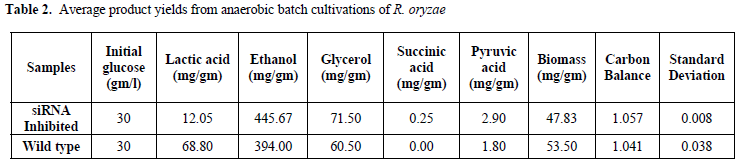 |
| indicate noticeable inhibition of lactic acid genes. The HPLC results also show that ethanol production increased after siRNA-mediated silencing of the ldh genes. The average ethanol production increased from 11.82gm/l in wild types to 13.37gm/l in knockdown samples (Table 1, Figure 1). It corresponds to an average increase in the yield of ethanol from glucose from 0.39gm/gm in wild types to 0.45gm/gm in the siRNA-inhibited samples (15.4% (gm/gm)). |
| Effect of RNA silencing on other metabolites of glucose metabolism pathway |
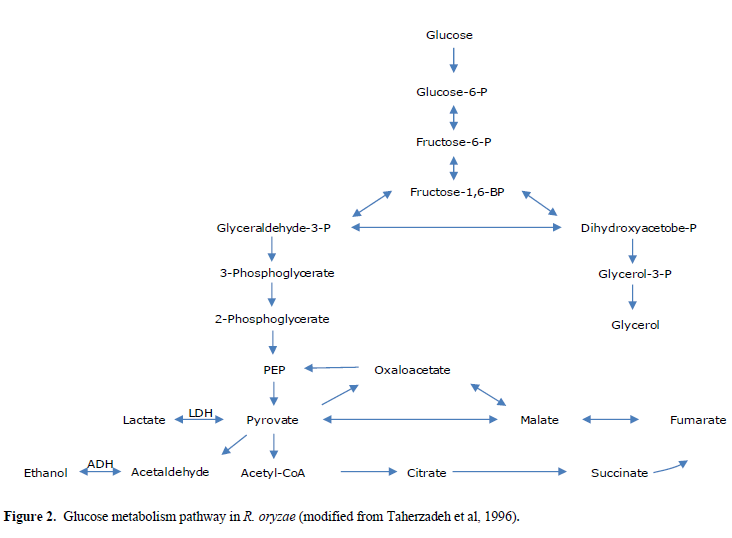
| Lactate dehydrogenase (LDH) catalyzes conversion of pyruvate to lactate. Consequently, the silencing of ldhA, i.e., the corresponding gene in lactic acid production, can affect the pathway of glucose metabolism in R. oryzae (Figure 2). Suppression of lactic acid production in this pathway triggers pyruvate to mainly go through the production of other metabolites. Moreover, after RNA silencing of the lactic acid and ethanol, the yields of glycerol, succinic acid, pyruvate, and biomass were affected too (Table 1). The yields of glycerol, succinic acid, and pyruvate increased after the silencing of ldhA gene. The biomass concentration was decreased after the silencing of the ldhA gene. The product yields obtained for anaerobic batch cultivation of R. oryzae with initial concentration of 30gm/l of glucose as a sole carbon source are shown in Table 2. |
| Wilcoxon rank sum test |
| In order to investigate whether the silencing of the ldhA gene resulted in significant differences between the production of lactic acid and ethanol in wild type and knockdown, respectively, the samples were tested using the R statistical program (Version 2.11.1) for one-sided Wilcoxon rank sum test (Mann-Whitney U-test), for two independent samples as they could not be considered to be normally distributed. Wilcoxon rank sum test showed that the silenced ldhA gene in R. oryzae could with statistical significance, on the 5% level, both reduce the lactic acid production (p-value=0.033) and increase the ethanol production (p-value=0.036) in knockdowns compared with the wild types. |
| It is evident that chemically-synthesised siRNAs cannot be used on industrial scale to produce ethanol and other metabolites due to the prohibitive costs of their synthesis. Therefore, the aim of this study was to demonstrate that gene silencing by RNAi is indeed effective in R. oryzae and can lead to improved ethanol yields. We now aim to develop genetically-modified R. oryzae strains expressing endogenous shRNAs targeting ldh genes for industrial applications. We anticipate that the data obtained this study will be of considerable value to this future work. |
Acknowledgements |
| This work was funded by the Swedish Energy Agency. |
Completing Interests |
| None declared. |
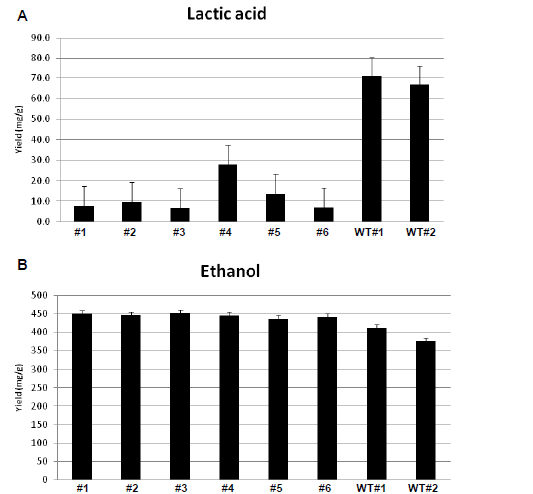 |
| Figure 1. Obtained results from HPLC. A. Lactic acid production in Knockdown and wild types Rhizopus oryzae detected by UV in HPLC. B. Ethanol production in Knockdown and wild types Rhizopus oryzae detected by RI in HPLC. (WT: Wild type; #1-#1: siRNA-treated sample) |
 |
References
|