Research Article - Biomedical Research (2017) Volume 28, Issue 4
Morphological and molecular imaging of skin samples
Edgar Guevara1*, Jose Manuel Gutierrez-Hernandez1, Alexandre Castonguay2,3, Frederic Lesage2,3, Benjamín Moncada4 and Francisco Javier González11Coordinacion para la Innovacion y Aplicacion de la Ciencia y la Tecnología, Universidad Autonoma de San Luis Potosi, San Luis Potosi, Mexico
2Department of Electrical Engineering, Ecole Polytechnique de Montreal, Montreal, Canada
3Montreal Heart Institute, Montreal, Canada
4Department of Dermatology, Hospital Ignacio Morones Prieto, San Luis Potosi, Mexico
- *Corresponding Author:
- Edgar Guevara
CONACYT Research Fellow, Coordinacion para la Innovacion y Aplicacion de la Ciencia y la Tecnología
Universidad Autonoma de San Luis Potosi, Mexico
Accepted date: September 14, 2016
Abstract
An exhaustive characterization of skin samples is a necessary step in investigating dermatological disorders. By combining depth sensitive morphological imaging with molecular imaging, we aim to provide multiple pieces of information about the skin. In this work, we present tridimensional, highresolution mosaic images of skin biopsies using Optical Coherence Tomography (OCT) co-registered with standard microscopy, bi-dimensional Raman spectral mapping, and fluorescence imaging. A paraffin-embedded skin sample of a healthy human breast (female, age=49), was imaged with a sweptsource OCT system at 1310 nm. Individual OCT volumes were acquired in a fully automated fashion in order to obtain a relatively large field-of-view (~ 2 × 2 cm) at high resolution (~ 10 μm). Using anatomical landmarks, the other three modalities were manually co-registered to the surface of the OCT volume, using an affine transformation. The main limitation of our method is manual co-registration, which may hamper its effectiveness. Nonetheless, the results suggest that the concomitant use of several imaging modalities may provide morphological and molecular information about the sample. Our results suggest the use of multi-modal microscopy as a tool in skin biopsies characterization.
Keywords
Biomedical optical imaging, Raman spectroscopy, Skin, Microscopy.
Introduction
A useful tool in evaluating skin biopsies is Optical Coherence Tomography (OCT), which is a non-contact optical imaging modality based on the measured echo delay of backscattered light [1]. Being a depth-sensitive technique, it provides information beneath the epidermis with a resolution around ten micrometers. Diagnosis of non-pigmented cancer tumors has been investigated with the aid of OCT, showing promising results [2]. This technique has also proved useful in the definition of the excision area prior to surgery of basal cell carcinoma [3]. Raman Spectroscopy (RS) has been shown to discriminate between benign and malignant tumors, determining the biochemical differences among them with high specificity [4,5]. RS has been previously applied as an optical biopsy technique in filaggrin-related atopic dermatitis [6], skin cancer, in-vivo classification of skin cancers, pre-cancers, and benign lesions [7], as well as a complementary technique for diagnosis of tissues during cancer surgery [8]. A detailed characterization of skin lesions in a non-invasive way requires the development of a multi-modal instrument with the potential to provide anatomical and biochemical information, using imaging techniques suited for each modality. Previous demonstrations of combined RS and OCT show promising results [9-15]. A limitation of these previous efforts is the small field-of-view of approximately 5 × 5 mm2. We present an alternative to overcome this drawback with the stitching of multiple pictures on a mosaic image, covering large areas of the skin biopsy. This technique permits a larger Field of View (FOV) while preserving the native resolution of individual scans.
Methods
Tissue preparation
This pilot study was approved by the institutional ethics committee and carried out at the hospital “Ignacio Morones Prieto” on a patient (female, age=49) who underwent excisional biopsy for breast cancer diagnosis. Informed consent was obtained prior to her participation in this research. All procedures in this pilot study were carried out on a single sample, comprised of two sections of human breast tissue obtained and preserved in a paraffin block for pathological examination [16]. The fixative for histology was 10% neutral buffered formalin. These two skin sections were oriented at an angle perpendicular to the epidermis. These specimens were diagnosed as healthy by an expert histopathologist.
Optical coherence tomography
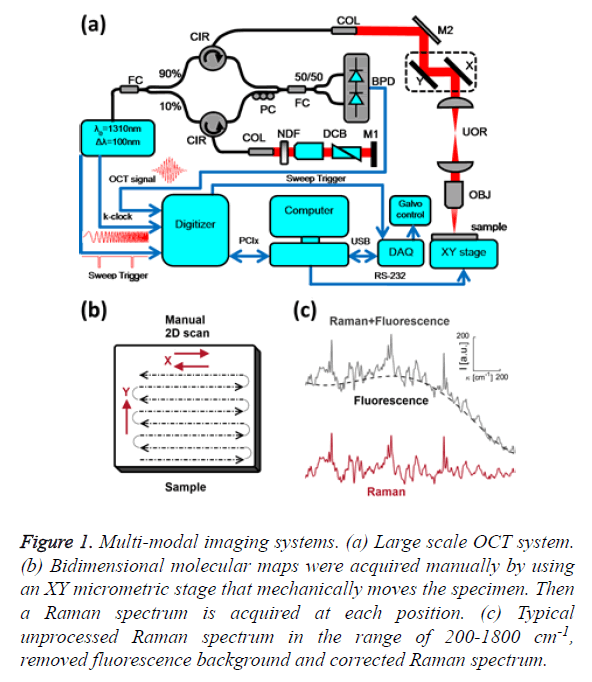
OCT is a recent biomedical imaging technique. It fills the void between confocal microscopy and high-resolution ultrasound in terms of resolution and penetration depth. The structural information of the skin specimen was provided by a Swept- Source OCT setup, as depicted in Figure 1A. This custom made system employs a 1310 Swept Source Engine (Axsun Billerica, MA) as light source. This laser has a central wavelength of λ0=1310 nm with a tuning bandwidth Δλ=100 nm. The average power at the laser output was about 18 mW. The output of the source is split into a reference arm, which shines light on a stationary mirror and a sample arm, which illuminates and receives the light reflected from within the sample. Reflectance measurements of tissue at each depth are recorded simultaneously with a Balanced Photo Detector (BPD). All fibers are single-mode, 125 μm cladding, terminated with FC/APC (8˚ angle physical contact) connectors that minimize back-reflections in the OCT system.
Figure 1: Multi-modal imaging systems. (a) Large scale OCT system. (b) Bidimensional molecular maps were acquired manually by using an XY micrometric stage that mechanically moves the specimen. Then a Raman spectrum is acquired at each position. (c) Typical unprocessed Raman spectrum in the range of 200-1800 cm-1, removed fluorescence background and corrected Raman spectrum.
The core of the imaging system is a Michelson-type interferometer built with two fiber couplers and two circulators. The incoming light from the swept-source passes through a broadband 90/10 fused fiber optic coupler (FC, Thorlabs, FC1310-70-10-APC). The 90% portion goes into a circulator (CIR, Thorlabs, CIR-1310-50-APC) that sends light to the sample arm and collects the backscattered light from the sample. The 10% portion is passed into another circulator that sends light to the reference arm and collects it again. Backscattered light from both the sample and reference arms are combined in the 50/50 fused fiber optic coupler (FC, Thorlabs, FC1310-70-50-APC), producing an interference pattern that goes to the inputs of the BPD (Thorlabs, PDB120C). The polarization controller (PC, Thorlabs, PLC-900) is used to optimize the contrast of the interference pattern.
The light from the sample interferes with light reflected by the silver-coated mirror M1 (Thorlabs, PF10-03-P01); interference can occur only when the optical path lengths of the two arms match to within the coherence length of the source. The intensity of the light from the reference arm is adjusted with an adjustable neutral density filter (NDF, Thorlabs, NDC-50C-2M) in order to avoid saturation of the detected signal. The raw output of the detected signal is defined by Equation 1,
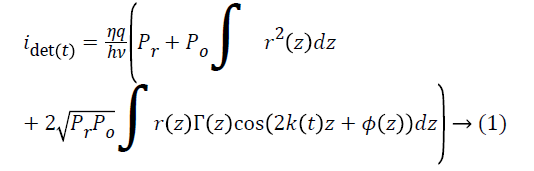
Where η is the photo detector sensitivity, q is the elementary charge and hν is the energy of a single photon. The first term Pr is the optical power reflected from the reference arm. The second term depends on the optical power illuminating the sample Po. Γ (z) is the coherence function of the instantaneous laser output and k (t) is the wavenumber, which is linearly swept by the laser source. The third term represents the interferometric signal, which is related to the sample reflection profile via the Fourier transform relation.
The main advantage of a swept-source system with respect to Fourier-domain OCT is that the intensity of the light across the bandwidth, distributed in high peak powers at each discrete wavelength, allows for a greater sensitivity with little risk of optical damage [17], while the 50 kHz sweeping rate enables high-speed imaging. In practice, the detected signal is digitized and a FFT is performed to obtain an axial scan, called an A-line. The signal acquisition was done with an ATS9350 digitizer (DAQ, 500 MS/s, 12 bit, PCIx. Alazar Technologies Inc. Pointe-Claire, QC). Data sampling was done directly in k-space, triggered by the k-clock from the swept-source, eluding interpolation inaccuracies and thus avoiding degraded resolution. The sample was imaged through an infinity-corrected 10x objective (OBJ, Olympus UMPLFLN 10 XW, Markham, ON). The axial and lateral resolutions were 13 and 10 μm, respectively. Cross-sectional images were generated by performing a series of axial scans at different transverse positions to generate a two-dimensional data set (B-scan). Three-dimensional data sets were then generated by raster scanning a series of two-dimensional B-scans. Scanning was performed by a pair of galvanometer mirrors (X,Y, Thorlabs, GVSM002/M) mounted on a cage (GCM002/M). After the galvanometer mirrors, there is a Unitary Optical Relay (UOR) consisting of two identical lenses arranged in an afocal (infinite focus) assembly (Thorlabs, LA1131-C, f1=f2=50 mm, 1050<λ<1620 nm). This UOR is used to image the mirror’s pivot point at the entrance pupil of the objective. Volumes consisting of 512 B-scans with 512 A-lines per B-scan were acquired over 1 mm × 1 mm. The k-clock from the swept source triggered the acquisition of the 1128 samples along an A-line. The volumes were cropped at a depth of 520 μm, because the SNR and image contrast are very weak at deeper locations and the morphological information was difficult to discern. Due to the large bandwidth of the swept source, different wavelengths will travel at different speeds when propagating through dispersive media such as the glass in the fiber optics and the lenses. A dispersion mismatch between the interferometer arms will affect the axial resolution of the system. Therefore, it is necessary to balance the dispersion mismatch between the sample and reference arms to achieve optimal resolution. Thus, the same amount of glass that exists in the sample arm is placed in the reference arm. The objective glass and the glass of the optical relay are partially compensated by a dispersion compensation block (DCB, Thorlabs, LSM04DC) and by a prism pair (Thorlabs, PS908L-C), which are coupled with refractive index matching oil for BK7 glass (Cargille, BK-7 Matching Liquid 11510).
A dispersion compensation algorithm was implemented according to a procedure described elsewhere [18]. Data was high-pass filtered to remove the DC scattering components from the image.
To acquire multiple volumes in an automated fashion, a motorized XY stage was used. The XY stage was comprised of two motorized linear slides (T-LSR150B and T-LSR300B, Zaber Technologies Inc., Vancouver, Canada) controlled by RS-232 serial commands. A large mosaic field-of-view (FOV 14.6 mm × 22.1 mm) was comprised of hundreds of individual volumes, aligned and stitched in an automated fashion [19,20].
Individual volumes were flat-field corrected to minimize shading artifacts in the resulting mosaic. A memory-mapped file was deemed necessary to deal with physical memory limitations. All acquisition and processing routines were custom written in MATLAB (The Math Works, Natick, MA), except where noted.
Visual microscopy
The 10X microscope objective used for OCT was coupled to a digital single lens reflex camera (Canon EOS 7D, Canon Inc., Tokyo, Japan), using a RMS-to-EF mount adapter. Several photographs were taken in order to cover the whole sample. Stitching of the individual images was done with image composite editor (Microsoft Corporation, Redmond, WA). Co-registration of the visual images and the projection of the tomographic volume were performed via manual alignment of three anatomical features, which defines the affine transformation.
Raman spectroscopy and fluorescence imaging
Raman spectroscopy was performed at 785 nm yielding 90 mW to the sample (R-3000 Raman Systems, Woburn, MA). This spectrometer works in the range of 200-1800 cm-1 with a spectral resolution <10 cm-1. A bi-dimensional map was created by taking one spectrum every 500 μm over a 5 mm × 5 mm FOV, with the aid of a micrometric XY stage, as depicted in Figure 1B. The integration time was 10 seconds per pixel. All spectra were processed by a fluorescence removal algorithm, known as the Vancouver algorithm [21], defined by Equation 2 and illustrated by Figure 1C.
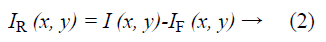
Where IR (x, y) is the spatially varying intensity of the pure Raman spectra, I (x, y) is the original Raman signal plus the autofluorescence background and IF (x, y) is the multi-polynomial fitting representing the fluorescence signal.
This intrinsic fluorescence background removal has been shown to yield reproducible and consistent Raman spectra, relatively free of fluorescence contributions, thus minimizing the influence of paraffin fluorescence in our measurements. The first 2-D map encodes the intensity IR (x, y) of the polysaccharides Raman band, at 848 cm-1 [22]. The second map shows the integrated fluorescence of the sample IF (x, y) from 5.55 μm to 50 μm. In order to reduce systematic errors in Raman measurements, all spectra were min/max normalized to the amide-III band at 1268 cm-1, i.e. IR (x, y) = (IR (x, y)-Imin)/ Imax [23]. Both maps were upsampled in order to reduce pixelization, and coregistered to the OCT volume using the edges of the plastic container of the sample as points of reference. This was done because the Raman spectral map and fluorescence image offer little anatomical features, while the spectra coming from the plastic were very distinct from the sample.
Results and Discussion
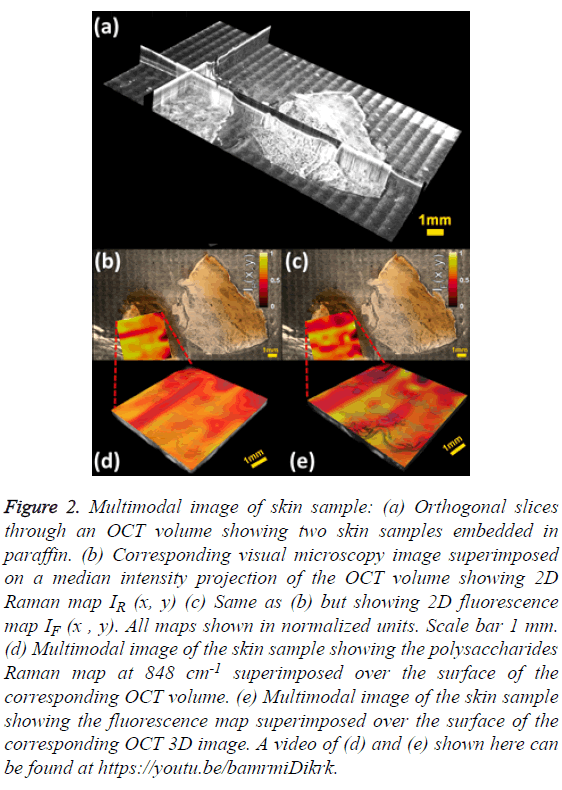
The mosaic volume provides depth-sensitive structural information about the skin sample, revealed by orthogonal cross sections visualized with Fiji [24] as shown in Figure 2A. The visible image overlaid on the median intensity-projected OCT volume shows some structures not accessible by standard microscopy, as displayed in Figures 2B and 2C. The visual microscopic image of the sample may be used as a tool in diagnosis of pigmented lesions, for instance, assessing color variegation or border non-uniformity. Molecular maps IR (x, y) and IF (x, y) are also displayed. One limitation of stitching multiple images is the shading artifact present at every individual volume, which persisted even after flat-field correction. Moreover, this pattern changes in every slice of the volume, making its correction difficult, as it is done in standard microscopy [25]. The intensity map of the polysaccharides band IR (x, y) shows low values for the epidermis region and mixed values in the dermis zone, as depicted in Figure 2B. A similar behaviour is observed in the fluorescence map IF (x, y) as shown in Figure 2C. The advantage of multimodal imaging relies on the fact that conventional imaging can only detect anatomical abnormalities, which might not indicate diseased tissue. In these cases, a biopsy must be performed in order to find out if the tissue is healthy or not. When molecular information is available, it can determine whether the tissue is healthy or not. Molecular information can help to get a diagnosis non-invasively, which is why it is often called “optical biopsy.” In order to supply as much information as possible in a single image, the 2D molecular maps were superimposed on the top surface of the OCT volume using the novel method of Jo et al. [26].
Figure 2: Multimodal image of skin sample: (a) Orthogonal slices through an OCT volume showing two skin samples embedded in paraffin. (b) Corresponding visual microscopy image superimposed on a median intensity projection of the OCT volume showing 2D Raman map IR (x, y) (c) Same as (b) but showing 2D fluorescence map IF (x , y). All maps shown in normalized units. Scale bar 1 mm. (d) Multimodal image of the skin sample showing the polysaccharides Raman map at 848 cm-1 superimposed over the surface of the corresponding OCT volume. (e) Multimodal image of the skin sample showing the fluorescence map superimposed over the surface of the corresponding OCT 3D image. A video of (d) and (e) shown here can be found at https://youtu.be/bamrmiDikrk.
Discussion
Conclusion
This pilot study indicates that multi-modal microscopy may be a valuable tool in the characterization of skin biopsies, allowing the potential extraction of several biomarkers. Furthermore, our results suggest that it would be very useful to integrate a Raman probe into the OCT setup, in order to obtain high-resolution molecular and structural images over a large FOV, in a fully automated fashion.
Several limitations need to be mentioned. First, and most crucial, is the lack of automatic co-registration between the different modalities. Despite the careful choice of control points based on anatomy, there may remain errors that would hamper the usefulness of the method. This will be alleviated in future work where the Raman spectrometer probe will be integrated onto the OCT sample arm, through a dichroic mirror replacing M2 in Figure 1A. Alternating acquisition of OCT and Raman data would then be performed, automatically solving the problem of positioning concordance. Although acquisition of Raman spectra is several orders of magnitude slower than OCT, it does not limit its potential as a chemical technique if it is applied ex-vivo. For in-vivo applications, the use of the faster Raman spectrometers, capable of acquiring hundreds of spectra per second, would be mandatory.
Although not demonstrated here, the concurrent use of these imaging modalities makes it possible to extract several biomarkers of the skin sample. For instance, the optical density of the OCT images has been shown to correlate with systemic sclerosis [27] and average skin attenuation has been explored as a predictor of solar elastosis score [28], while psoriasis has been shown to affect skin fluorescence [29]. Although the mechanism is not completely clear, polysaccharides have been shown to exert antitumor action [30]. Thus, the authors hypothesized the value of using their distribution as a marker of tumor prognosis.
Moreover, Raman imaging is not restricted to a single compound, but can be performed to identify the spatial distribution of multiple molecules. For example, it has been demonstrated that Raman spectroscopy is useful in both carcinoma and melanoma diagnosis, characterized by the 788 and 1093 cm-1 bands corresponding to the O-P-O phosphodiester and PO2 vibrations in DNA, respectively [31].
Further examples of multi-modal Raman and OCT imaging can be found in the literature, where visual defects of pork skin, [32] ex-vivo retina [33], or basal cell carcinoma [10] have been characterized.
Acknowledgements
This work was supported in part (F. L. and F. J. G.) by the second stage of the Groupe de Travail Quebec/Mexique/XIIIe reunion/Biennum 2011-2013 and by the "Cátedras CONACYT" program, project 528. The authors cordially thank Dr. Javier A. Jo and Paritosh Pande for advice on image visualization and Samuel Belanger for allowing us the use of his photographic equipment.
References
- Huang D, Swanson EA, Lin CP, Schuman JS, Stinson WG. Optical coherence tomography. Science 1991; 254: 1178-1181.
- Forsea AM, Carstea EM, Ghervase L, Giurcaneanu C, Pavelescu G. Clinical application of optical coherence tomography for the imaging of non-melanocytic cutaneous tumors: A pilot multi-modal study. J Med Life 2010; 3: 381-389.
- Wang KX, Meekings A, Fluhr JW, McKenzie G, Lee DA, Fisher J. Optical coherence tomography-based optimization of mohs micrographic surgery of basal cell carcinoma: A pilot study. Dermatol Surg Off Publ Am Soc Dermatol Surg Al 2013; 39: 627-633.
- Calin MA, Parasca SV, Savastru R, Calin MR, Dontu S. Optical techniques for the noninvasive diagnosis of skin cancer. J Cancer Res Clin Oncol 2013; 139: 1083-1104.
- Drakaki E, Vergou T, Dessinioti C, Stratigos AJ, Salavastru C, Antoniou C. Spectroscopic methods for the photodiagnosis of nonmelanoma skin cancer. J Biomed Opt 2013; 18: 61221.
- Gonzalez FJ, Alda J, Moreno-Cruz B, Martinez-Escaname M, Ramirez-Elias MG, Torres-Alvarez B. Use of Raman spectroscopy for the early detection of filaggrin-related atopic dermatitis. Skin Res Technol Off J Int Soc Bioeng Skin ISBS Int Soc Digit Imaging Skin ISDIS Int Soc Skin Imaging 2011; 17: 45-50.
- Lui H, Zhao J, McLean D, Zeng H. Real-time Raman spectroscopy for in vivo skin cancer diagnosis. Cancer Res 2012; 72: 2491-2500.
- Kong K, Rowlands CJ, Varma S, Perkins W, Leach IH, Koloydenko AA. Diagnosis of tumors during tissue-conserving surgery with integrated autofluorescence and Raman scattering microscopy. Proc Natl Acad Sci USA 2013; 110: 15189-15194.
- Patil CA, Bosschaart N, Keller MD, van Leeuwen TG, Mahadevan-Jansen A. Combined Raman spectroscopy and optical coherence tomography device for tissue characterization. Opt Lett 2008; 33: 1135-1137.
- Patil CA, Kirshnamoorthi H, Ellis DL, van Leeuwen TG, Mahadevan-Jansen A. A clinical instrument for combined Raman spectroscopy-optical coherence tomography of skin cancers. Lasers Surg Med 2011; 43: 143-151.
- Patil CA, Krishnamoorthi H, Ellis DL, van Leeuwen TG, Mahadevan-Jansen A. A clinical probe for combined Raman spectroscopy-optical coherence tomography (RS-OCT) of the skin cancers. Proc SPIE San Francisco California USA SPIE 2010.
- Patil CA, Kalkman J, Faber D, Penn JS, van Leeuwen TG, Mahadevan-Jansen A. Structural and biochemical characterization of the rat retina with combined Raman Spectroscopy-Spectral Domain Optical Coherence Tomography (RS-SDOCT). Proc SPIE San Francisco California USA SPIE 2010; 75501-75501.
- Patil CA, Kalkman J, Faber DJ, Nyman JS, van Leeuwen TG, Mahadevan-Jansen A. Integrated system for combined Raman spectroscopy-spectral domain optical coherence tomography. J Biomed Opt 2011; 16: 11007.
- Liu CH, Qi J, Lu J, Wang S, Wu C, Shih WC. Improvement of tissue analysis and classification using optical coherence tomography combined with Raman spectroscopy. Spi Org 2014.
- Liu CH, Qi J, Lu J, Wang S, Wu C, Shih WC. Improvement of tissue analysis and classification using optical coherence tomography combined with Raman spectroscopy. J Innov Opt Health Sci 2014 13; 1550006.
- Al-Refu K. General methods in preparation of skin biopsies for haematoxylin and eosin stain and immunohistochemistry. Skin Biopsy Persp Tech 2011.
- Liu B, Brezinski ME. Theoretical and practical considerations on detection performance of time domain, Fourier domain, and swept source optical coherence tomography. J Biomed Opt 2007; 12: 44007.
- Wojtkowski M, Srinivasan V, Ko T, Fujimoto J, Kowalczyk A, Duker J. Ultrahigh-resolution, high-speed, Fourier domain optical coherence tomography and methods for dispersion compensation. Opt Express 2004; 12: 2404-2422.
- Guevara E, Gutierrez-Hernandez JM, Castonguay A, Lesage F, Gonzalez FJ. Ex-vivo multi-modal microscopy of healthy skin. Photonic Sensors and Biomedical Optics Montreal Canada 2014; 92880-9288020.
- Castonguay A, Avti PK, Moeini M, Pouliot P, Tabatabaei MS, Belanger S. Investigating the correlation between white matter and microvasculature changes in aging using large scale optical coherence tomography and confocal fluorescence imaging combined with tissue sectioning. Proc Spid Lib 2015; 93281.
- Zhao J, Lui H, McLean DI, Zeng H. Automated autofluorescence background subtraction algorithm for biomedical Raman spectroscopy. Appl Spectrosc 2007; 61: 1225-1232.
- Parker FS. Applications of infrared, Raman, and resonance Raman spectroscopy in biochemistry. New York Spr 1983; 582.
- Hseu TH, Chang H. Laser Raman studies on the conformation of Pro-Leu-Gly-NH2. Biochim Biophys Acta 1980; 624: 340-345.
- Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M. Fiji: an open-source platform for biological image analysis. Nat Methods 2012; 9: 676-682.
- Marty GD. Blank-field correction for achieving a uniform white background in brightfield digital photomicrographs. BioTechniques 2007; 42: 716-720.
- Jo JA, Applegate BE, Jesung Park, Shrestha S, Pande P, Gimenez-Conti IB. In vivo simultaneous morphological and biochemical optical imaging of oral epithelial cancer. IEEE Trans Biomed Eng 2010; 57: 2596-2599.
- Abignano G, Aydin SZ, Castillo-Gallego C, Liakouli V, Woods D, Meekings A. Virtual skin biopsy by optical coherence tomography: the first quantitative imaging biomarker for scleroderma. Ann Rheum Dis 2013; 72: 1845-1851.
- Korde VR, Bonnema GT, Xu W, Krishnamurthy C, Ranger-Moore J. Using optical coherence tomography to evaluate skin sun damage and precancer. Lasers Surg Med 2007; 39: 687-695.
- Portugal-Cohen M, Horev L, Ruffer C, Schlippe G, Voss W, Maor Z evi. Non-invasive skin biomarkers quantification of psoriasis and atopic dermatitis: cytokines, antioxidants and psoriatic skin auto-fluorescence. Biomed Pharmacother Biomed Pharmacotherapie 2012; 66: 293-299.
- Ooi VE, Liu F. Immunomodulation and anti-cancer activity of polysaccharide-protein complexes. Curr Med Chem 2000; 7: 715-729.
- Larraona-Puy M, Ghita A, Zoladek A, Perkins W, Varma S. Development of Raman microspectroscopy for automated detection and imaging of basal cell carcinoma. J Biomed Opt 2009; 14: 054031.
- Egodage K, Dochow S, Bocklitz T, Chernavskaia O, Matthaeus C, Schmitt M. The combination of optical coherence tomography and Raman spectroscopy for tissue characterization. J Biomed Photonics Eng 2015; 1: 169-177.
- Evans JW, Zawadzki RJ, Liu R, Chan JW, Lane SM. Optical coherence tomography and Raman spectroscopy of the ex-vivo retina. J Biophotonics 2009; 2: 398-406.