Research Article - Journal of RNA and Genomics (2017) Volume 13, Issue 1
Isolation and identification of Wolbachia from Hemiscorpius lepturus scorpion in Khuzestan province with PCR method
Neda Ashtian1*, Sara Rajaie1, Ahmad Taghavi Moghadam2 and Mohammad Ali Bayatzadeh3
1Islamic Azad University of Hamedan
2Agricultural Research, Education and Extension Organization (AREEO), Tehran, Iran
3Razi Vaccine and Serum Research Institute-Ahvaz Branch, Iran
Received: 22 September 2017; Revised: 23 October 2017; Accepted: 25 October 2017; Published: 01 November 2017
© Copyright The Author(s). First Published by Allied Academies. This is an open access article, published under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0). This license permits noncommercial use, distribution and reproduction of the article, provided the original work is appropriately acknowledged with correct citation details
Abstract
Wolbachia is the most frequent gram negative Endosymbiont bacteria belonging to α-proteobacter group which can effect sexual and somatic tissues of arthropods. Due to manipulate the host’s reproductive system can be used as a new method for biological control of insects, bacteria important veterinary disease vectors used. In this study, after collection of scorpion species from field with the help of UV lamp at night, their DNA was isolated and two special primers against 16srDNA and WSP were designed. After setting up PCR conditions with positive control samples, PCR unknown samples were marked. The amplified fragment by agarose gel electrophoresis have shown. Then we have the positive samples sequenced. During our study on 40 Hemiscorpius lepturus scorpions, Twenty-eight of the specimens were positive for Wolbachia bacteria, all of which were positive for both primers WSP and 16srDNA primers. The presence of Wolbachia bacteria in the Hemiscorpius Lepturus scorpion has been proven. This detection was performed by two 16srDNA and WSP genes. The prevalence of the bacterium Wolbachia sp. in Khuzestan province can be due to effects of temperature, which is an inhibitory factor for growth of the bacterium in the arthropods. The presence of Wolbachia in H. lepturus scorpions was approved. The accessions number of 16Sr DNA sequences is KY554756 and wsp gene is KY554757 that deposit in Gen Bank.
Keywords
Wolbachia, Hemiscorpius lepturus, PCR
Introduction
Wolbachia is a gram negative, obligate, intracellular symbiont, that is classified in α-proteobacter (Inaki et al., 2011). Wolbachia is a cytoplasmical inherited rickettsia that was found in a wide range of arthropods (Jeyaprakash and Hoy, 2000; Werren and Windsor, 2000; Hilgenboecker et al., 2008). They are widespread symbionts detected in all arthropod classes (infecting between 20 and 70% of insect species) (Bourtzis and Miller, 2003; Zug and Hammerstein, 2012) as well as in a nematode family, the Onchocercidae, commonly known as filarial (Bandi et al., 1998; Sironi et al., 1995). Thus, Wolbachia are present in millions of species.
Intracellular bacterium Wolbachia first time within the context of reproductive culex pipiens mosquitoes was reported in 1924 by Wolbach & Hertig. Infections were detected in all of the major insect orders, including Coleoptera, Diptera, Hemiptera/Homoptera, Hymenoptera, Lepidoptera, and Orthoptera (Werren and Windsor, 2000). Depending on both bacterial lineage and host, they may have different effects on host reproduction, such as cytoplasmic incompatibility (Breeuwer and Werren, 1990; O’Neill and Karr, 1990), male killing (Hurst et al., 1999), parthenogenetic reproduction (Stouthamer, 1997; Zchori-Fein et al., 2001) or feminization of genetic males (Bouchon et al., 1998; Juchault et al., 1994). Wolbachia cannot be cultured outside the host cells (O’Neill et al., 1992), and its potential application to the bio control of insect pests (Ravikumar et al., 2011) and vector borne diseases such as dengue fever (Inaki et al., 2011), malaria and filariasis (Chaoyang et al., 2009) have been studied.
Classification of strains of Wolbachia is based on the incompatibility of the cytoplasm, the sequence of genes 16srDNA (Breeuwer et al., 1992; O’Neill et al., 1992; Rousset et al., 1992; Stouthamer et al., 1993) ftsZ (the cell cycle belongs) (186), WSP (the gene encoding the protein regions vary wildly) (Bourtzis et al., 1994; Braig et al., 1998; Zhou et al., 1998), coxa, glta and groEL done that 16sr DNA sequence-based classification and WSP among them the most important factors are high (Glowska et al., 2015; Vera et al., 2008).
The initial report for the presence of Wolbachia in scorpion is related to southern African genus, Opistophthalmus (Laura et al., 2007). Lincoln Suesdek-Rocha et al in 2007 reported the Wolbachia bacteria isolated from scorpion Tityus serrulatus (Lincoln Suesdek-Rocha et al., 2007).
The main purpose of this study was to search for Wolbachia 16S rDNA and WSP gene in the genomic DNA preparation made from the dangerous scorpions H. lepturus (Hemiscorpiidae) as the most lethal and stinging a great number of people annually one with major concern (Moghadam et al., 2011) Among different types of available scorpions in Khuzestan.
Materials and Methods
Scorpion’s samples
40 Iranian scorpions H. lepturus were collected from Khuzestan province Cities include: Masjed Soleiman
(10 sample), Izeh
(10 sample), Ramhormoz
(10 sample), Ahvaz
(10 sample). All scorpions were identified by trained healthcaring personnel of the laboratory reference of Razi Institute in Ahvaz. Scorpions are individual and have maintained separately. Samples of both sexes were used.
Scorpions were washed with 70% alcohol (Roche, Germany). After catching the scorpion tail end zone (stinger), tail (mentasoma), pendipalp, the head, the chest (shoulder sex), abdomen and legs separated and extracted DNA and kept at -80°C.
DNA extraction
All DNA extraction using High Pure PCR Product Purification Kit (Roche, Germany) with the instructions of kit. The extracted DNA was dissolved in DEPCddH2O (Roche, Germany) solution and frozen at -20°C and kept until it was used for rDNA and WSP amplification by PCR.
16S rDNA and WSP amplification by PCR
The PCR amplification reactions were carried out by Bio- Rad thermocycler (Bio-Rad Laboratories, Inc. USA). In 25 μl reaction mixtures consisting of 0.6 μl of dNTP, 0.6 μl of each primer, 5 μl of the crude DNA extract, 2 μl MgCl2, 2.5 μl of 10X PCR buffer, 0.4 μl of Taq polymerase DNA and 13.3 μl of D.D.W for primer 16S rDNA. In 25 μl reaction mixtures consisting of 0.6 μl of dNTP, 0.5 μl of each primer, 5 μl of the crude DNA extract, 2 μl MgCl2, 2.5 μl of 10X PCR buffer, 0.5 μl of Taq polymerase DNA and 13.4 μl of D.D.W for primer wsp. Specific primers used in this work were Wsp-R (5/AAA AAT TAA ACG CTA CTC CA) and Wsp-F (5/TGG TCC AAT AAG TGA TGA AGA AAC)( Zhou et al., 1998) and rDNA-R (5/GAA TAG GTA TGA TTT TCA TGT) and rDNA- F (5/TTG TAG CCT GCT TAT GGT ATA ACT)(O’Neill et al., 1992). 16Sr DNA amplifies a 890 bp fragment and wsp amplify a 620 bp fragment.
16S rDNA sequence are designed from the moderately variable regions of 16S rDNA and are relatively conserved among Wolbachia. WSP primers are the gene encoding a surface protein.
The PCR Wsp primer conditions for initial denaturation were carried out for 5 min at 95°C, followed by 35 cycles consisting of 94°C for 45 s, 49°C for 1 min and 72°C for 45 s, and a final extension for 7 min at 72°C.
The PCR rDNA primer conditions for initial denaturation were carried out for 5 min at 95°C, followed by 35 cycles consisting of 94°C for 45 s, 50°C for 1 min and 72°C for 1 min, and a final extension for 7 min at 72°C as described.
10 μl of the amplified product with a 100 bp DNA ladder were separated by 1% agarose gel (Roche, Germany) electrophoresis including ethidium bromide and visualized by UV transilluminator (UVItec Limited, United Kingdom) to determine the presence and size of the amplified DNA.
DNA sequence analysis
Wolbachia positive samples then had their 16S rDNA and WSP gene fragment amplified and sequenced with the forward (5/TTG TAG CCT GCT TAT GGT ATA ACT) and reverse (5/GAA TAG GTA TGA TTT TCA TGT) PCR primers, and WSP gene fragment amplified and sequenced with the forward (5/TGG TCC AAT AAG TGA TGA AGA AAC) and reverse (5/AAA AAT TAA ACG CTA CTC CA) PCR primers. Amplicons were subjected to electrophoresis to confirm that they consisted of a single product and then purified using GF-1 gel DNA recovery Kit (vivantis ind.). After purification, sequencing was performed sent to Bioneer Company (Munpyeongseo-ro, Daedeok-gu, Daejeon 306-220, Republic of Korea) for nucleotide sequencing in both directions.
Phylogenetic and sequence alignment analyses
Alignment and Phylogenetic analysis of these sequences were performed using BioEdit 7.2 (http://www.mbio. ncsu.edu/bioedit/bioedit.html) and MEGA7 (http://www. megasoftware.net/) software respectively. The phylogenetic analysis was based on the Neighbor-Joining method with bootstrap test (1000 replicates) method. In addition to the isolate obtained in the present study, Sequence was compared with Gen Bank database using the BLAST software from NCBI site (http://www.ncbi.nlm.nih.gov) and the sequences of isolates from GenBank were also added to the analysis.
Results
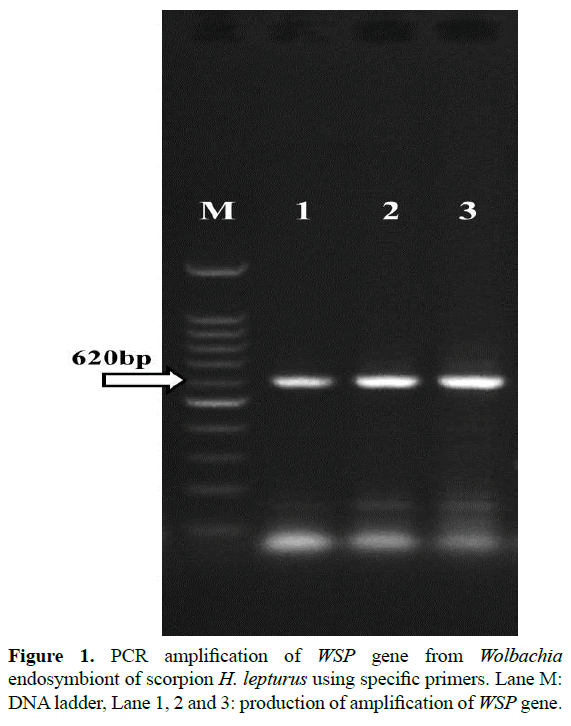
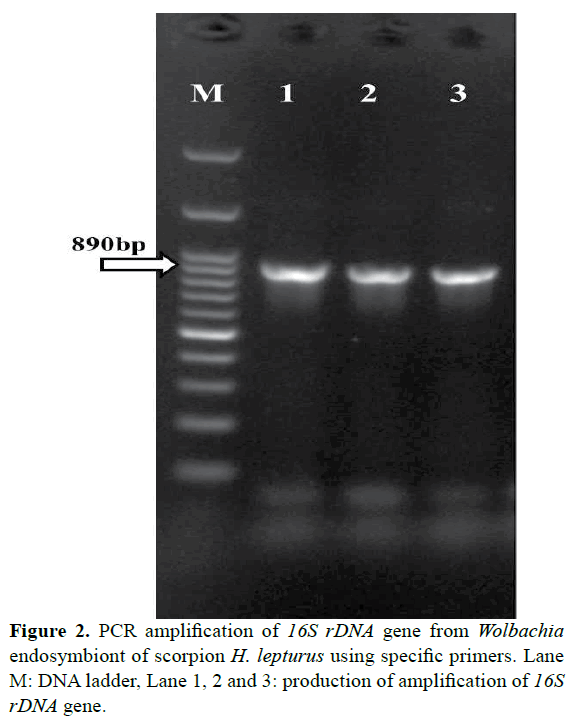
All scorpions were morphologically Hemiscorpius lepturus. Total of 40 scorpions H. lepturus were screened for Wolbachia by PCR method. Wolbachia positive by both 16S rDNA and WSP gene PCR were found in 28 of them. 8 from Masjed Soleiman, 9 from Izeh, 7 from Ramhormoz, 4 from Ahvaz. PCR reaction amplified fragments in the range of about 620 bp for wsp of Wolbachia endosymbiont (Figure 1). PCR reaction amplified fragments in the range of about 890 bp for 16S rDNA of Wolbachia endosymbiont (Figure 2). Therefore, the presence of Wolbachia in H. lepturus scorpions was approved.
The accessions number of 16Sr DNA sequences is KY554756 and WSP gene is KY554757 that deposit in Gen Bank.
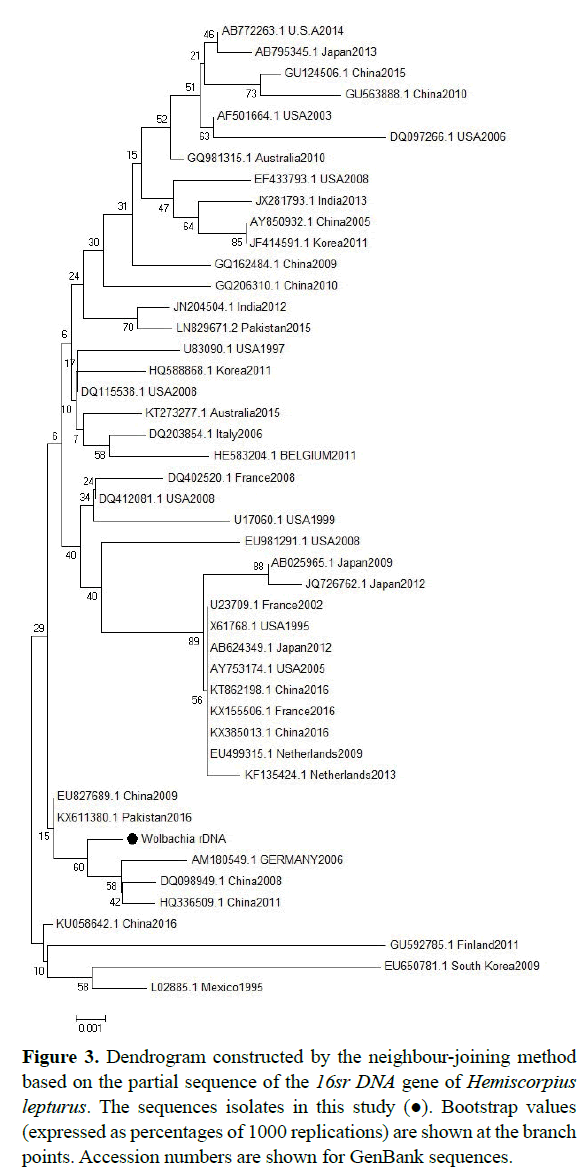
According to the phylogenetic tree, sequences derived from the sequence of the Wolbachia 16srDNA sequences isolated from China, Pakistan and Germany have similar (accession numbers are shown in the dendrogram), but the sequences isolated from Netherland, America, Australia and India is different (Figure 3). According to the phylogenetic tree, sequences derived from the sequence of the Wolbachia WSP gene sequences isolated from China and America and have very similar (accession numbers are shown in the dendrogram), but the sequences isolated from America, France, china and japan is very different (Figure 4).
Figure 3: Dendrogram constructed by the neighbour-joining method based on the partial sequence of the 16sr DNA gene of Hemiscorpius lepturus. The sequences isolates in this study (?). Bootstrap values (expressed as percentages of 1000 replications) are shown at the branch points. Accession numbers are shown for GenBank sequences.
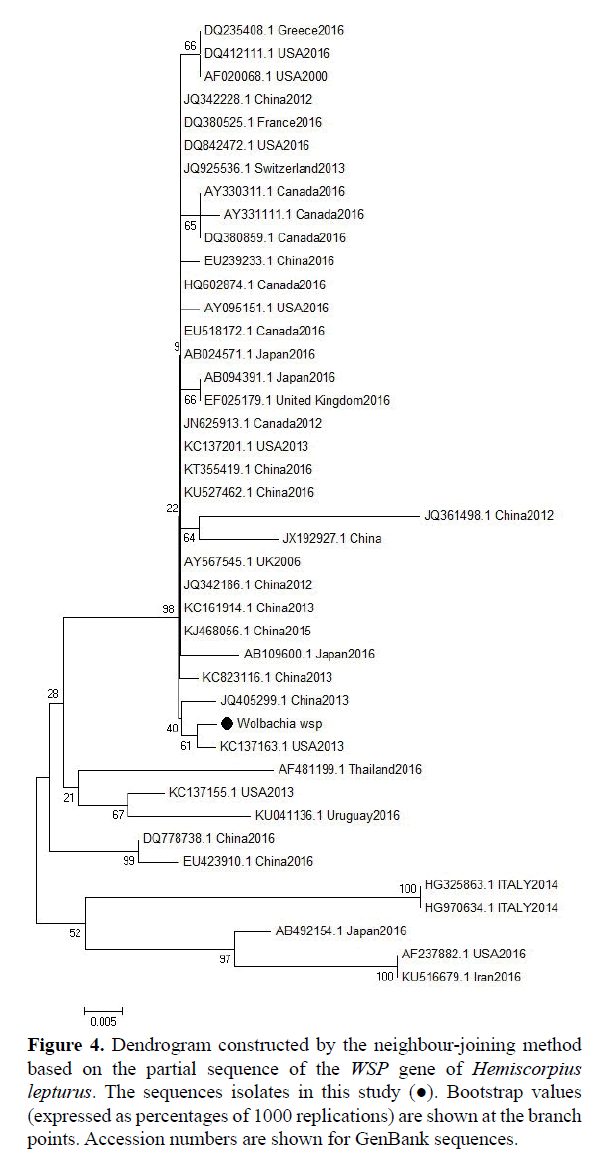
Figure 4: Dendrogram constructed by the neighbour-joining method based on the partial sequence of the WSP gene of Hemiscorpius lepturus. The sequences isolates in this study (?). Bootstrap values (expressed as percentages of 1000 replications) are shown at the branch points. Accession numbers are shown for GenBank sequences.
The most similar rDNA sequences with sequences (Pakistan, 2016) and (China, 2009) 99.6% and Differences with (Korea, 2011) and (Netherlands, 2013) was 97%. The most similar wsp sequences with sequences (japan, 2016) 99.3% and (Canada, 2009) 98% and Differences with (China, 2015) 79% and (France, 2016) was 72%.
Discussion
In this study, Hemiscorpius lepturus scorpion in Khuzestan, Iran could test based on PCR for 16S rDNA and WSP genes was isolated in the bacterium Wolbachia. Scorpions coming from Izeh were collected, have more infections than scorpions were collected from more temperate regions (Ramhormoz and Ahvaz). According to this study, Wolbachia infection in scorpion Hemiscorpius lepturus was detected by PCR method.
Sylvia Joanne et al. (2014), in Malaysia studied on Aedes albopictus been, a total of 286 samples based on the sequence wsp tested (67 male and 219 female), which (91.60% ) for both Wolbachia (A, B) were infected, (3.15 %) only had type A, (1.05) percent were infected with only one of two types or (2.05%) of the unknown and (5.25) percent were without infection. (Joanne et al., 2015).
Didier Bouchon et al. (1998) in a study carried out on the 689 number of insects that includes (231 male and 458 female) of arthropod species reported 22 cases of insects Wolbachia bacteria were extracted.
O'Neill et al. in 1992 studied on different genes and 16s rDNA sequences on Culex pipiens and several other species of insects showed, Their DNA sequences and insect colony PCR and investment in different species, different sequences of these genes from Wolbachia were found. (O’Neill et al., 1992).
According to our findings, Wolbachia bacteria isolated from scorpion Hemiscorpius lepturus by PCR method in both WSP gene and 16S rDNA gene. The method used in this study is similar to the studies mentioned.
Bradaran et al. (2011), examined the 20 scorpion Hemiscorpius lepturus were collected from Khuzestan (Iran). Wolbachia bacteria are present in 10 samples patented Hemi scorpion. This study was conducted by PCR method and 16Sr DNA primers. (Baradaran et al., 2011).
Comparison of the 16S rDNA fragment and wsp fragment with the GenBank database revealed that the nucleotide sequence of 16S rDNA of Wolbachia endosymbiont of H. lepturus was highly similar to that of Drosophila melanogaster.
Some horizontal transmission has nevertheless occurred between scorpions, Wolbachia 16S rRNA and WSP gene sequences are identical in infected species of the H. lepturus. Similarly, the 16S rRNA gene sequence of Wolbachia from Drosophila melanogaster, an insect (Arthropoda), is identical to that of scorpion H. lepturus were examined.
The present study is the first report of the presence of Wolbachia in H. lepturus. Since the sampling were performed through Khuzestan, the prevalence of the bacterium Wolbachia sp. in these area can be due to effects of temperature, which is an inhibitory factor for growth of the bacterium in the arthropods.
In this study, 40 samples from scorpion H. lepturus were examined to detect presence of bacteria Wolbachia; we were able to isolate the bacteria from scorpion H. lepturus. The survey has correlation with the similar studies. In 40 samples, 28 samples were positive for the WSP gene and the 16S rDNA gene.
References
- Baldo L, Prendini L, Corthals A, et al. 2007. Wolbachia Are Present in Southern African Scorpions and Cluster with Supergroup F. Current Microbiology. 55, 367-373.
- Bandi C, Anderson TJ, Genchi C, et al. 1998. Phylogeny of Wolbachia in filarial nematodes. P ROY SOC B-BIOL SCI. 265, 2407-2413
- Baradaran M, Jalali A and Jolodar A. 2011. Molecular diagnosis of Wolbachia endosymbiont from Iranian scorpion Hemiscorpius lepturus using polymerase chain reaction (PCR) amplification of 16S rDNA gene. African Journal of Biotechnology. 10, 19802-19206.
- Bouchon D, Rigaud T and Juchault P. 1998. Evidence for widespread Wolbachia infection in isopod crustaceans: molecular identification and host feminization. Proceedings of the Royal Society B: Biological Sciences. 265, 1081–1090.
- Bourtzis K and Miller T, editors. 2003. Insect Symbiosis. Contemporary Topics in Entomology. CRC Press.
- Bourtzis K, Nirgianaki A, Onyango P, et al. 1994. A prokaryotic dnaA sequence in Drosophila melanogasten Wolbachia infection and cytoplasmic incompatibility among laboratory strains. Insect Molecular Biology. 3, 131–142.
- Braig HR, Zhou W, Dobson SL, et al. 1998. Cloning and Characterization of a Gene Encoding the Major Surface Protein of the Bacterial EndosymbiontWolbachia pipientis. Journal of bacteriology. 180, 2373-2378.
- Breeuwer JA and Werren JH. 1990. Microorganisms associated with chromosome destruction and reproductive isolation between two insect species. Nature. 346, 558-560.
- Breeuwer JA, Stouthamer R, Barns SM, et al. 1992. Phylogeny of cytoplasmic incompatibility microorganisms in the parasitoid wasp genus Nasonia (Hymenoptera: Pteromalidae) based on 16S ribosomal DNA sequences. Insect molecular biology. 1, 25-36.
- Cordaux R, Michel-Salzat A, Frelon-Raimond M, et al. 2004. Evidence for a new feminizing Wolbachia strain in the isopod Armadillidium vulgare: evolutionary implications. Heredity. 93, 78-84.
- Glowska E, Dragun-Damian A, Dabert M, et al. 2015. New Wolbachia supergroups detected in quill mites (Acari: Syringophilidae). Infection, Genetics and Evolution. 30, 140-146.
- Hilgenboecker K, Hammerstein P, Schlattmann P, et al. 2008. How many species are infected with Wolbachia? A statistical analysis of current data. FEMS Microbiology Letters. 281, 215-220.
- Hurst GDD, Jiggins FM, Hinrich Graf von der Schulenburg J, et al. 1999. Male-killing Wolbachia in two species of insect. Proceedings of the Royal Society B: Biological Sciences. 266, 735-740.
- Iturbe-Ormaetxe I, Woolfit M, Rancès E, et al. 2011. A simple protocol to obtain highly pure Wolbachia endosymbiont DNA for genome sequencing. Journal of Microbiological Methods. 84, 134-136.
- Jeyaprakash A and Hoy MA. 2000. Long PCR improves Wolbachia DNA amplification: wsp sequences found in 76% of sixty-three arthropod species. Insect Molecular Biology. 9, 393-405.
- Jin C, Ren X and Rasgon JL. 2009. The virulent Wolbachia strain wMelPop efficiently establishes somatic infections in the malaria vector Anopheles gambiae. Applied and environmental microbiology. 75, 3373-3376.
- Joanne S, Vythilingam I, Yugavathy N, et al. 2015. Distribution and dynamics of Wolbachia infection in Malaysian Aedes albopictus. Acta Tropica. 148, 38-45.
- O’Neill SL and Karr TL. 1990. Bidirectional incompatibility between conspecific populations of Drosophila simulans. Nature. 348, 178-180.
- O’Neill SL, Giordano R, Colbert AM, et al. 1992. 16S rRNA phylogenetic analysis of the bacterial endosymbionts associated with cytoplasmic incompatibility in insects. Proceedings of the National Academy of Sciences. 89, 2699-2702.
- Ravikumar H, Ramachandraswamy N and Puttaraju H. 2011. Molecular strain typing of Wolbachia infection from Indian mosquitoes using wsp gene. Asian Pacific Journal of Tropical Disease. 1, 106-109.
- Ros VID, Fleming VM, Feil EJ, et al. 2008. How Diverse Is the Genus Wolbachia? Multiple-Gene Sequencing Reveals a Putatively New Wolbachia Supergroup Recovered from Spider Mites (Acari: Tetranychidae). Applied and Environmental Microbiology. 75, 1036-1043.
- Rousset F, Vautrin D and Solignac M. 1992. Molecular Identification of Wolbachia, the Agent of Cytoplasmic Incompatibility in Drosophila simulans, and Variability in Relation with Host Mitochondrial Types. Proceedings of the Royal Society B: Biological Sciences. 247, 163-168.
- Sironi M, Bandi C, Sacchi L, et al. 1995. Molecular evidence for a close relative of the arthropod endosymbiont Wolbachia in a filarial worm. Molecular and Biochemical Parasitology. 74, 223-227.
- Stouthamer R, Breeuwer JAJ, Luck RF, et al. 1993. Molecular identification of microorganisms associated with parthenogenesis. Nature. 361, 66-68.
- Stouthamer R. 1997. Wolbachia-induced parthenogenesis. In: O’Neill SL, Hoffman AA and Werren JH (eds) Influential Passengers: Inherited Microorganisms and Arthropod Reproduction. Oxford Univerity Press, New York, pp 102-124.
- Suesdek-Rocha L, Bertani R, DaSilva P, et al. 2006. The first record for Wolbachia in a scorpion: the parthenogenetic yellow scorpion Tityus serrulatus (Scorpiones, Buthidae). Rev. Iber. Arachnol. 14, 183-184.
- Taghavi Moghadam A. 2011. Evaluation of scorpion hemolymph (blood) proteinic pattern by SDS-PAGE. Clinical Biochemistry. 44, S241.
- Werren JH and Windsor DM. 2000. Wolbachia infection frequencies in insects: evidence of a global equilibrium? Proceedings of the Royal Society B: Biological Sciences. 267, 1277-1285.
- Werren JH, Zhang W and Guo LR. 1995. Evolution and Phylogeny of Wolbachia: Reproductive Parasites of Arthropods. Proceedings of the Royal Society B: Biological Sciences. 261, 55-63.
- Zchori-Fein E, Gottlieb Y, Kelly SE, et al. 2001. A newly discovered bacterium associated with parthenogenesis and a change in host selection behavior in parasitoid wasps. Proceedings of the National Academy of Sciences. 98, 12555-12560.
- Zhou W, Rousset F and O’Neill S. 1998. Phylogeny and PCR-based classification of Wolbachia strains using wsp gene sequences. Proceedings of the Royal Society B: Biological Sciences. 265, 509-515.
- Zug R and Hammerstein P. 2012. Still a Host of Hosts for Wolbachia: Analysis of Recent Data Suggests That 40% of Terrestrial Arthropod Species Are Infected. Cordaux R, editor. PLoS ONE. 7, e38544.