Research Article - Biomedical Research (2017) Volume 28, Issue 14
High urine miR-126 level predicts bladder cancer in hematuria patients
Xiangqian Xu, Hui Yuan, Jinyan Zhong, Baotong Wu, Liang Zhang and Youbao Wang*
Department of Urology, Qingdao Hiser Hospital, Qingdao, PR China
Accepted on June 8, 2017
Abstract
Objective: Bladder cancer is one of the commonest genitourinary tumors all over the world. The early detection rate was still very low due to the lack of effective diagnostic biomarkers. In this study, we examined the feasibility and efficacy of urine miR-126 detection in the diagnosis of bladder cancer, which could help identify a new diagnostic biomarker for such deadly disease.
Methods: Patients with bladder cancer (bladder cancer group) and urinary tract infection (non-bladder cancer group) admitted in our department from January 2013 to December 2016 were enrolled. The two groups were age and gender matched. Demographic and clinical data were retrieved, and urine samples were collected. Real-time RT-PCR was used to detect urine miR-126 relative expression.
Results: A total of 134 bladder cancer patients were included in bladder cancer group, while there were 268 patients with urinary tract infection in non-bladder cancer patients. Real-time RT-PCR results proved that urine miR-126 level in bladder cancer patients were greatly higher than that in non-bladder cancer patients (P=0.03). In our study, there were a total of 181 patients with hematuria. High miR-126 could predict the diagnosis of bladder cancer in patients with hematuria (P=0.01).
Conclusions: Increased urine miR-126 level could predict the diagnosis of bladder cancer, especially in patients with hematuria. Our findings supported that the detection of urine miR-126 could serve as a diagnostic biomarker for bladder cancer.
Keywords
Bladder cancer, microRNAs, miR-126, Diagnosis, Hematuria.
Introduction
Bladder cancer is one of the commonest malignant diseases in urinary system, which has the highest incidence among genitourinary tumors in China [1,2]. In recent years, the prevalence of bladder cancer has been greatly on the increase. Epidemiological investigations proved that a series of etiological factors were associated the development and progression of bladder cancer, including smoking, drinking [3]. Painless hematuria is the main clinical manifestation in such patients, but the symptoms at early stage are nonspecific. Thus, the identification of early diagnostic biomarkers for bladder cancer is of special significance.
Micro-RNAs (miRNAs) are small RNA molecules that regulate the downstream genes [4]. It has been reported that the dysregulations of miRNAs could contribute the pathogenesis of multiple human diseases [5]. The oncogenic roles of miR-126 have been well validated in bladder cancer, which could promote tumor growth and progression both in vitro and in vivo experiments [6,7]. Previous studies demonstrated that aberrant miRNAs levels may be introduced as new biomarkers for early diagnosis and prognosis in bladder cancer [8]. However, most researchers detected the miRNAs level in tissue and blood samples [9]. Considering that bladder cancer as a genitourinary tumor and the convenience of urine collections,we further examined whether urinary miRNAs detection can serve as a diagnostic markers for bladder cancer, especially in patients with hematuria. Our findings on the detection of urine miR-126 level for predicting bladder cancer may provide new insights into the developing a new effective diagnostic and prognostic markers for bladder cancer, which can enlighten the future researches and optimize the current management.
Patients and Methods
Patients
Patients who were initially diagnosed as bladder cancer in our department from January 2013 to December 2016 (bladder cancer group) were enrolled, and all the patients were pathologically diagnosed. Patients with urinary tract infection who were 1: 2 matched with age and gender admitted in the same period (non-bladder cancer group) were also included. Demographic and clinical data were retrieved from the computerized database, and the urine samples were collected.
This study was approved by the Ethic Committee of our hospital and all the patients gave their written informed consent.
Urine preparation and total RNA extraction
3.54 g Guanidinium Thiocyanate (GTC) and salt solutions were added into the urine samples to make a final concentrations of 6 mol/l GTC, 0.025 mol/l sodium acetate, 0.25% N-lauroylsarcosin, 0.5 mol/l HEPES at pH 7, and then the samples were stored at -80°C for RNA preparation.
The pH value of stabilized urine samples was adjusted to pH 7.0 by adding 1 mol/l HEPES at pH 7.0. An equal volume of acidic phenol: chloroform was added to the urine samples to extract RNA. The silica membrane and 100% (vol/vol) ethanol were applied to purify RNA. The miRNeasy kit (Qiagen, Hilden, Germany) was used to extract miRNAs from the urine samples. RNA extracts were quantified by NanoDrop 2000 (Thermo Scientific, Massachusetts, the USA) and stored at -80°C.
Real-time RT-PCR and miR-126 detection
The reverse transcription reaction of 5 μl RNA extract was carried out using TaqMan MicroRNA Reverse Transcription Kit (Applied Biosystems, Darmstadt, Germany), containing 5 μl RNA, 1 mmol/l of each deoxyribonucleotide triphosphate, 50 units of Multicribe Reverse Transcriptase, 1 μl reaction buffer, 4 units RNase inhibitor and 1 μl gene-specific primers, and nuclease-free H2O was added to a final volume of 15 μl. The mixture was incubated at 16°C for 30 min and then at 42°C for 30 min to synthesize cDNA. cDNA was quantified using NanoDrop 2000 (Thermo Scientific, Massachusetts, the USA).
miR-126 and the small nuclear RNA U6 (internal control) were detected in triplicates via qRT-PCR using human TaqMan MicroRNA Assay Kits (Applied Biosystems, Darmstadt, Germany). 1.33 μl cDNA solution was amplified using 1 μl TaqMan Universal PCR Master Mix 656 M, 1 μl gene-specific primers, and nuclease-free H2O in a final volume of 20 μl. The quantitative PCR was run on a 7900 HT quantitative PCR machine (Applied Biosystems, Darmstadt, Germany) with an initial denaturation at 95°C for 10 min, followed by 40 cycles with a denaturation at 95°C for 15 s and an annealing/ elongation at 60°C for 60 s. The Cycle threshold (Ct) values were calculated. The relative expression of miR-126 was normalized to the internal control. The primers for miR-126 and RNU6B were purchased from Takara Bio Inc.
Statistical analysis
All the statistical analysis was conducted by SPSS 17.0 software (SPSS Inc., Chicago, IL, USA). Continuous and categorical data were presented as mean (range) and percentage (%), and the differences on continuous and categorical data were tested by student t-test and chi-square test when appropriate, respectively. A two-tailed P value less than 0.05 was considered to be statistically significant.
Results
Demographic and clinical characteristics
A total of 134 bladder cancer patients were included in bladder cancer group, while there were 268 patients with urinary tract infection in non-bladder cancer patients (Table 1). The two groups were age and gender matched. The mean age at admission was both 52.8, ranging 38-68 (P=1.00). The female: male ratio was both around 1:1.6 in both two groups (P=1.00). More patients in bladder cancer group had a family history of genitourinary tumors (n=89, 66.4% vs. n=92, 34.3%, P=0.03). In bladder cancer patients, the commonest clinical manifestation was hematuria (n=101, 75.4%), while urinary irritation symptoms including odynuria, urgent urination and frequent micturition (n=186, 69.4) were most commonly seen in non-bladder cancer group (P=0.04, Table 1). It was suggested that patients with hematuria, especially painless hematuria should be further examined to exclude the diagnosis of bladder cancer.
| Bladder cancer group (n=134) | Non bladder cancer group (n=268) | P value | |
|---|---|---|---|
| Gender, n (%) | 1 | ||
| Female | 52 (38.8) | 104 (38.8) | |
| Male | 82 (61.2) | 164 (61.2) | |
| Age at admission, years, mean (range) | 52.8 (38-68) | 52.8 (38-68) | 1 |
| Family history of genitourinary tumors, n (%) | 89 (66.4) | 92 (34.3) | 0.03 |
| Clinical manifestations, n (%) | 0.04 | ||
| Hematuria | 101 (75.4) | 80 (29.9) | |
| Urinary irritation symptoms | 21 (15.7) | 186 (69.4) | |
| Weight loss | 12 (8.9) | 2 (0.7) |
Table 1. Demographic and clinical characteristics.
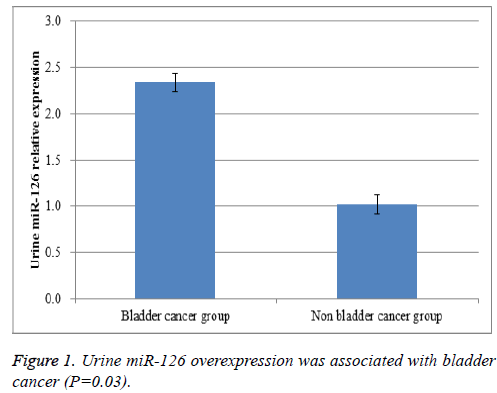
Increased urine miR-126 level was associated with bladder cancer
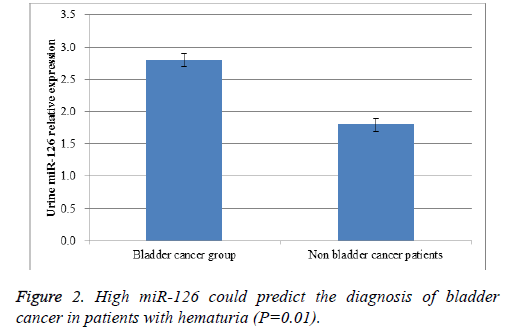
Urine samples were collected and total RNA was extracted. Real-time RT-PCR results proved that urine miR-126 level in bladder cancer patients were greatly higher than that in nonbladder cancer patients (P=0.03, Figure 1). Then, we further analysed whether urine miR-126 detection could be able to differentiate bladder cancer from urinary tract infection in patients with hematuria. In our study, there were a total of 181 patients with hematuria. High miR-126 could predict the diagnosis of bladder cancer in patients with hematuria (P=0.01, Figure 2). These findings could benefit the early diagnosis of bladder cancer in patients who had suspected clinical symptoms like painless hematuria.
Discussion
Bladder cancer remains a deadly disease with relatively poor survival and early detection rate. Although great efforts have been made to clarify the carcinogenesis and develop new effective treatments for bladder cancer, there is still no cure [10]. Bladder cancer patients at early stage could be treated by radical surgery, which may improve the patients’ life quality and prognosis. However, those patients at advanced stage may only be treated by chemotherapy, radiotherapy and targeted therapy, and the clinical efficacy is quite different among different individuals. Thus, the early diagnosis of bladder cancer is closely correlated with the clinical outcome, indicating that the validation of effective biomarkers for early diagnosis of bladder cancer is vital to the clinical management of such patients [11,12]. In this study, we reported that the detection of urine miR-126 could be feasible and effective in differentiating bladder cancer from urinary tract infection, especially in patients with hematuria.
So far, hundreds of molecules have been proved to be the potential biomarkers for early diagnosis of bladder cancer [13-15]. Among them, miRNAs are the most promising candidates, which could regulate the functions of downstream target genes, consequently mediating a series of biological and physiological processes including cell proliferation, cells apoptosis, cell metabolism and the like [4,16]. In bladder cancer, miR-126 was overexpressed in tumor tissue compared with that in normal tissue [17]. Both in vitro and in vivo experiments have demonstrated that miR-126 could enhance tumor cell proliferation, invasion and migration, but suppress tumor cell apoptosis in bladder cancer, suggesting that miR-126 could play an oncogenic role in the development and progression of bladder cancer [18]. In clinical practice, the acquisition of tissue specimens is difficult, which can be completed either by biopsy or by surgical resection, while urine collection is much more easy and convenient. In addition, the detection of urine miR-126 level in the diagnosis of genitourinary tumors has been rarely reported [19]. Thus, we proposed that urine miR-126 expression may be a very promising early diagnostic biomarkers for bladder cancer.
In this study, patients with bladder cancer and urinary tract infection were both enrolled, and the later served as the controls. The results proved that urine miR-126 level was higher in bladder cancer than that in non-bladder cancer, which was statistically significant. Subgroup analysis on the patients with hematuria reached a similar conclusion that increased miR-126 level in urine samples could differentiate bladder cancer from urinary infection tract, which was of very special clinical significance. There were still limitations in our study. First, all the patients were from one single center and a large scale multicenter investigation may provide more reliable results. Second, healthy controls were not enrolled, because all the patients enrolled were hospitalized in our department and there were no healthy controls. In future, eligible outpatients may also be included as healthy controls. Third, the prognostic value of urine miR-126 detection has not been evaluated in this study. In addition, Due to the limitation of our sample size, ROC curve was not drawn to determine the cut-off value of miR-126 for diagnosing bladder cancer. We are now enrolling more patients and the optimal cut-off value of miR-126 will be further examined.
Taken together, urine detection of miR-126 level was feasible and effective in the early diagnosis of bladder cancer, and this method could also be introduced to the differentiation of bladder cancer and non-bladder cancer in patients with hematuria. This conclusion that urine miR-126 is potential biomarker for early diagnosis and outcome prediction of bladder cancer still needs to be further examined in large scale studies.
References
- Soloway MS. Challenging Cases in Urothelial Cancer. Bladder Cancer 2016; 2: 465-466.
- Chen W, Zheng R, Baade PD, Zhang S, Zeng H. Cancer statistics in China, 2015. CA Cancer J Clin 2016; 66: 115-132.
- Masaoka H, Matsuo K, Ito H, Wakai K, Nagata C, Nakayama T, Sadakane A, Tanaka K, Tamakoshi A, Sugawara Y, Mizoue T, Sawada N, Inoue M. Cigarette smoking and bladder cancer risk: an evaluation based on a systematic review of epidemiologic evidence in the Japanese population. Japan J Clin Oncol 2016; 46: 273-283.
- Amir S, Mabjeesh NJ. microRNA expression profiles as decision-making biomarkers in the management of bladder cancer. Histol Histopathol 2017; 32: 107-119.
- Kurozumi A, Goto Y, Okato A, Ichikawa T, Seki N. Aberrantly expressed microRNAs in bladder cancer and renal cell carcinoma. J Hum Genet 2016; 62: 49-56.
- Xiao J, Lin HY, Zhu YY, Zhu YP, Chen LW. MiR-126 regulates proliferation and invasion in the bladder cancer BLS cell line by targeting the PIK3R2-mediated PI3K/Akt signaling pathway. Onco Targets Ther 2016; 9: 5181-5193.
- Enokida H, Yoshino H, Matsushita R, Nakagawa M. The role of microRNAs in bladder cancer. Investig Clin Urol 2016; 57: 60-76.
- Lee JY, Ryu DS, Kim WJ, Kim SJ. Aberrantly expressed microRNAs in the context of bladder tumorigenesis. Investig Clin Urol 2016; 57: 52-59.
- Mitash N, Agnihotri S, Mittal B, Tiwari S, Mandhani A. Molecular cystoscopy: Micro-RNAs could be a marker for identifying genotypic changes for transitional cell carcinoma of the urinary bladder. Indian J Urol 2016; 32: 149-153.
- Chou R, Selph S, Buckley DI, Fu R, Griffin JC, Grusing S, Gore JL. Intravesical therapy for the treatment of non-muscle-invasive bladder cancer: a systematic review and meta-analysis. J Urol 2016; 197: 1189-1199.
- Fendler A, Stephan C. The translational potential of microRNAs as biofluid markers of urological tumours. Nat Rev Urol 2016; 13: 734-752.
- Nagata M, Muto S, Horie S. Molecular biomarkers in bladder cancer: novel potential indicators of prognosis and treatment outcomes. Disease Markers 2016; 2016: 8205836.
- Deng S, Wang W, Li X, Zhang P. Common genetic polymorphisms in pre-microRNAs and risk of bladder cancer. World J Surg Oncol 2015; 13: 297.
- Armstrong DA, Green BB, Seigne JD, Schned AR, Marsit CJ. MicroRNA molecular profiling from matched tumor and bio-fluids in bladder cancer. Mol Cancer 2015; 14: 194.
- Piletic K, Kunej T. MicroRNA epigenetic signatures in human disease. Arch Toxicol 2016; 90: 2405-2419.
- Wu P, Cao Z, Wu S. New progress of epigenetic biomarkers in urological cancer. Dis Markers 2016; 2016: 9864047.
- Jia AY, Castillo-Martin M, Bonal DM, Sanchez-Carbayo M, Silva JM, Cordon-Cardo C. MicroRNA-126 inhibits invasion in bladder cancer via regulation of ADAM9. Br J Cancer 2014; 110: 2945-2954.
- Liu LY, Wang W, Zhao LY, Guo B, Yang J, Zhao XG, Hou N, Ni L, Wang AY, Song TS, Huang C, Xu JR. Mir-126 inhibits growth of SGC-7901 cells by synergistically targeting the oncogenes PI3KR2 and Crk, and the tumor suppressor PLK2. Int J Oncol 2014; 45: 1257-1265.
- Hanke M, Hoefig K, Merz H, Feller AC, Kausch I, Jocham D, Warnecke JM, Sczakiel G. A robust methodology to study urine microRNA as tumor marker: microRNA-126 and microRNA-182 are related to urinary bladder cancer. Urol Oncol 2010; 28: 655-661.