Research Article - Biomedical Research (2018) Artificial Intelligent Techniques for Bio Medical Signal Processing: Edition-II
Detecting the ovarian cancer using big data analysis with effective model
Yasodha P1* and Ananthanarayanan NR2
1Department of Computer Science, Pachaiyappa’s College for Women, Kanchipuram, Tamil Nadu, India
2Department of CSA, SCSVMV University, Kanchipuram, Tamil Nadu, India
- *Corresponding Author:
- Yasodha P
Department of Computer Science
Pachaiyappa’s College for Women
Tamil Nadu, India
Accepted on May 29, 2017
DOI: 10.4066/biomedicalresearch.29-17-155
Visit for more related articles at Biomedical ResearchAbstract
Now a day, a vast amount of medical data is available which can be efficiently utilized for diagnostic procedure by adopting data mining concepts. The main objective of this paper is to use effective data mining approach on huge amount of ovarian cancer dataset to identify the diseases in an efficient way. Thus, in this paper propose a novel approach for identifying ovarian cancer using combined Self- Organizing Maps Immune Clonal Selection (SOMICS) and Grammatical Evolution Neural Networks (GENN). SOMICS algorithm used for better feature selection which is used for extracting valuable, implicit, and interesting information from vast amount of medical data and GENN is used for classification process. The experimental results show the comparison of the proposed method and other classification methods using three various classifiers such as, Support Vector Machine (SVM), Multi- Layer Perceptron (MLP), Feed Forward Neural Network (FFNN). The combined SOMICS and GENN method yields promising results on classification and feature selection accuracy for ovarian cancer dataset with 98.23% classification accuracy, 0.0021% mean square error.
Keywords
Data mining, Grammatical evolution neural networks (GENN), Self-organizing maps immune clonal selection (SOMICS) algorithm, Ovarian cancer.
Introduction
Ovarian cancer is the fifth most cancer for the women after the breast cancer, uterus cancer, bowl cancer and lung cancer [1]. The ovarian cancer identified from the women during their menopause that is having the above 50 ages but this disease affects the women at any age. As same as the other cancer, the ovarian cancer also does not having the symptoms but the progress growth of the cancer has been noticed by pelvic pain, bloating, swelling [2]. The spreading areas of the cancer in the women body is bladder, lymph nodes, lungs, lining abdomen and the liver. The survey in the United States, 2012, which was conducted on 22,280 women for diagnoses the ovarian cancer 15,500 patients are died from it and the estimation provides that 95% percentage of the women has minimum ovarian cancer syndrome for one year [3]. Thus the risk level of the cancer increased women’s life time and it created several problems such as they never have the children, and creates problem in menopause stage during their life time [4]. So, the accurate identification, reliable cancer diagnoses and treatment process should be improved to overcoming these difficulties for the women. Then the evaluation of the ovarian cancer is enhanced by enormous amount of dataset because it has various potential information [5]. Thus the big data approach based huge volume of dataset is used for diagnose the ovarian cancer because it has different types of patient information not only size, complex, heterogonous, volumonial and longitudes data [6]. These data’s are used to make the smarter decision and also it consumes the lower cost. There are five reasons [7] how the big data approach helps during the cancer identification process. This paper handling the data mining and big data techniques to obtain the knowledge based system for identifying and detecting the ovarian cancer from the huge volume of unorganized dataset. Thus the main contribution of the paper is as per following: initially the feature set is minimized by applying the Self-Organizing Maps Immune Clonal Selection (SOMICS) approach which reduces the data set for selecting the optimized features. Then the classification is performed by Grammatical Evolution Neural Networks (GENN). The performance evaluation is made with the several existing methods such as Support Vector Machine (SVM) [8], Multi-Layer Perceptron (MLP) [9], and Feed Forward Neural Network (FFNN) [10].
Big Data Health Informatics
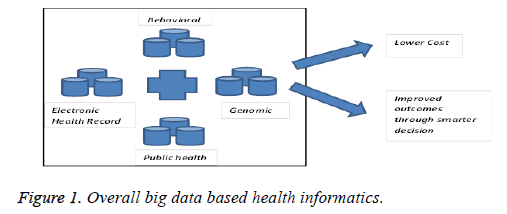
Big data in the sense it has huge collection of the information, it is normally represented by using the velocity, veracity, volume, variety and value terms [11]. In big data volume represents the size of the data, veracity represents that the genuineness of the information, velocity measures the pace at new data generation, variety represents that the different complexity of the information and value used to measure the quality of the data. Thus the big data approach is used to store the health informatics which is used during the disease diagnoses and the treatment. In this paper the big data based data set is used to analyse and detect the ovarian cancer because, the data set consist of multi scale information such as MRI details, recording, treatment, disease related symptoms, DNA micro data and so on. In the health informatics data set having different level [12] of health information which are mention as follows, bioinformatics, neuro informatics, clinical informatics, public health information, micro level data which means molecules, tissue level data, MRI details, patient level data such as monitored information, mission data and social data which are retrieved from the social medias such as Face book, twitter, Google and so. The social media data having the list of patient and user queries, doubts and general symptoms which are used during the diagnosis, treatment and prescription. Then the overall goal of big data analysis in the health informatics is providing the different variety of data’s with low cost and high quality. The sample big data based health informatics [13] is shown in the following Figure 1.
Thus the above Figure 1 shows that the different types of dataset and those data’s are used during the different disease diagnose process, treatments and etc. which provides the lower cost based improved outcomes through the smarter decision. This collected information is used for ovarian cancer detecting and analysing process.
Ovarian Cancer Identification Process Using Data Mining Concepts
The main goal of this paper is to detect and classify the ovarian cancer using the enormous volume of the data with the data mining concept. The dataset consists of various numbers of features and also having the noise, missing data’s and so on. So, that before processing the data set it has to be preprocessed by applying the following approach because the preprocessed data improves the performance of the system also increase the quality of the ovarian cancer detection process. But the data set has huge number of data’s which is difficult to pre-process, so the feature subset must be selected before processing the features. The feature subset must be selected by applying the self-organizing maps immune clonal Selection which is explained as follow.
Self-organizing maps immune clonal selection based feature subset selection
Feature subset selection [14] is the process of selecting the relevant features from the group of features which is used in the pattern recognition, classification and decision making process. The feature subset selection process is important in several applications because it simplify the models to interpret the functions, minimum training time and improved generalisation by minimized over fitting. In this paper Self- Organizing Maps (SOM) immune clonal selection feature subset selection method is used to reduce the features size which means selecting the important features from the huge data set without altering the semantics of the variables. The SOM algorithm is the one of the unsupervised algorithm which is works based on the two operations such as training and mapping. Then the feature subset is selected based on the similarity between the features which is estimated during the training and mapping. The SOM working process is explained as follows.
Self-organizing map: SOM is the one of the unsupervised learning method [15] which uses the competitive learning procedure that is it uses the wining node for further processing. It has the collection of nodes or neurons; each neuron has the particular weights in the vector space. The placing arrangement of the each node is placed in the hexagonal manner and it maps the high dimensional space into the low dimensional space. It has two important parameters namely training and mapping. During the training, the weights are updated continuously to obtain the wining node which has to by the similarity measures. The input vectors initialized for training process, then the distance between the each vectors are calculated by using the Mahalanobis distance [16] which is measured as follows
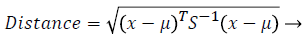 (1)
(1)
Where x is the input vectors and μ is the mean of the input vector and, S is the covarience of the input vector.
From the estimated distance, the best matching unit is identified by applying the immune clonal selection [17] approach which was explained as follows. From the best matching unit the input vectors are traced across the input vector space for selecting the best features. The feature matching has required for the input weight updating process which is calculated as follows.
Wv (s+1)=Wv (s)+Θ (u, v, s) α (s) (D (t)-Wv (s)) → (2) where, Wv represents that the node v current weight vector, s is the current iteration, Θ (u, v, s) is the distance of best matching unit usually called as the neighbourhood function, α (s) is the learning restraint D (t) target input vector. Then the weight updation process is repeated until to satisfy the threshold value. The selected features are used during the mapping. In the mapping, the similar features are grouped together and form the cluster which is used for further pre-processing and feature selection process.
Immune clonal selection algorithm: Immune clonal selection algorithm [18] works based on the immune response system which means it works how the antibodies of the immune system learn the features from the antigen. Initially the features (antibodies) are collected from the SOM training process then the affinity values of the antibody is estimated that is used when selecting the best matching unit. The affinity values have to be sorted for choosing the best features (antibodies). The selected features are mutated according to their affinity values. If the features are having the high value than it has to be cloned for less value else it has to be mutated highest value. Then the optimal features are selected using this mutation and cloning operation. This process is continued until to cluster the data’s in the largest data set. The feature subset selection Algorithm step is list out as follows.
Steps for feature subset selection
Step 1: Identify the dataset for selecting subset
Step 2: Assign the initial weights for each data in the data set
Step 3: Calculated the similarity between the data’s by Mahalanobis distance
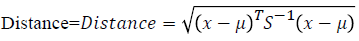
Step 4: Select the best matching unit by immune clonal selection approach
Step 5: Updating each data’s weights according to their important in the data base by
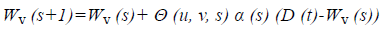
Step 6: Repeat the above steps 3 to 5.
Step 7: Mapping the each similar data’s into the single group (cluster)
Step 8: Select the minimum distance and minimum similarity clusters for further processing.
Step 9: Stop the condition at maximum condition.
By applying the above procedure, the feature subset has to be selected for further ovarian cancer identification and classification process.
Feature subset pre-processing
The feature subset selection process reduces the dimensionality of the features it makes easy to further classification process. The selected subsets have to be pre-processed for removing the noise from the data set that will be helpful during the classification for obtaining highest classification accuracy. The largest data set has to be pre-processed by using the two important steps. Weighting [19] and discretization [20] which is defined as follows,
Pre-processing by weighting approach: The subset selection process, selects the only relevant information and similar information, during that process each data has particular weights. The information weights are ranks are used in the preprocessing stage. Initially the information’s weights are analysed and eliminating the lowest weight because the lowest rank data does not provides any valuable information during the classification. The lowest rank data does not improve the accuracy at time of classification but it reduces the time and the efficiency of the classifier.
Pre-processing by discretization approach: Discretization is the process of partition the selected subset into the particular range by applying the binning and entropy method. These methods also ability to handle the missing and noisy data during the pre-processing approach. The discretization process minimizes the continuous attributes in the data set by dividing method and replaces the actual data by interval label. In this paper, discretization is done by entropy base method, initially the binning criteria are decided and then the information gain value is estimated. Then the information gain value is compared with the threshold for identifying the splitting criteria. If the information gain having the maximum value it has to be split into the mixed value else it has to be split according the similar data set. Then the maximum dissimilarity features or data’s are eliminated and the optimized features are selected by above discuss Self Organizing Map and the Immune Clonal Selection algorithm. The resultant features are used in the classification process.
Ovarian cancer classification using grammatical evolution neural networks
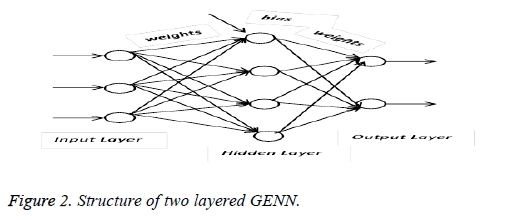
The next stage is the ovarian cancer classification which was done by using the Grammatical Evolution Neural Networks (GENN). In this paper the Neural Network is optimized by the grammatical evolution based evolutionary algorithm. It optimizes the neural network using the context free grammar and the genetic algorithm. The neural networks are constructed and trained by using the grammatical evolution based optimization [21] algorithm. In this paper two layered neural networks are used in which each node in the neural network represented as the Backus Naur form in context free grammar. The sample two layered grammatical evolution neural network is shown in following Figure 2.
In the neural network each node belongs to the particular context free grammar, and the neural network is represented as follows,
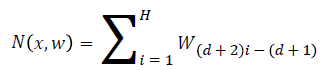
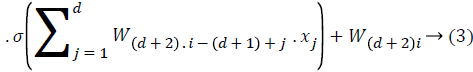
Where x represents that the input of the network w represents that the weights of the neural network. d is used to identify the number of hidden layers in the neural network.
In the neural network the sigmoid based activation function is used to find the classification output. The activation function is represented as follows.
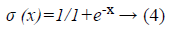
In the neural network the inputs are represented as the <xxlist> which is also called as the non-terminal node and the output of the neural network also represented as the non-terminal node. The every node in the neural network represented as the chromosome which is used during the network training and fitness calculation. Then the activation function is represented as the sig (.). The sample context free grammar based neural network is represented as follows in which each rules are having the rule number based on that the neural networks are trained.
| Rules | Rule number |
|---|---|
| S::=<sigexpr> | 0 |
| <sigexpr> ::= <Node> | 0 |
| |<Node> + <sigexpr> | 1 |
| <Node> ::= <number>*sig(<sum>+<number>) | 0 |
| <sum> ::= <number>*<xxlist> | 0 |
| |<sum>+<sum> | 1 |
| <xxlist> ::= x1 | 0 |
| |x2 | 1 |
| |... | (..) |
| |Xn | (n-1) |
| <number> ::= (<digitlist>.<digitlist>) | 0 |
| |(-<digitlist>.<digitlist>) | 1 |
| <digitlist> ::= <digit> | 0 |
| |<digit><digitlist> | 1 |
| <digit> ::= 0 | 0 |
| |1 | 1 |
| |2 | 2 |
| |.. | .. |
| |9 | 9 |
Then the neural network is constructed based on the above context free grammar and the training is performed by applying the following fitness function which is represented as follows,
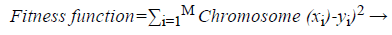 (5)
(5)
This GENN networks classify the incoming selected features into the ovarian cancer and the normal patient by using the above fitness function which is based on the chromosome node and the desired output based training algorithm. Then the structure of the neural networks does not limit by the genetic algorithm. The binary representation of the neural network classifies the incoming features with highest classification accuracy which was discussed in the following section.
Experimental Results and Discussion
The ovarian cancer is the most serious cancer for the woman which leads to death at the saviour stage. This cancer is classified by using the Self-Organizing Maps Immune Clonal Selection (SOMICS) and grammatical evolution neural networks. Then the proposed system demonstration is performed on the following data sets such as NCI PBSII data and curated ovarian data which is explained as follows,
Dataset
NCI PBSII data: NCI PBSII data set (http://datam.i2r.astar. edu.sg/datasets/krbd/OvarianCancer/OvarianCancer-NCIPBSII. html) is one of the ovarian cancer data set in which it consists of list of women’s details who having the highest risk factor. The dataset consists of 91 normal and 162 ovarian cancer information. In the dataset each samples having the related amplitude and intensity and molecular details, so totally there are 15154 identities. Then the identities are normalized according to the following formula.
NV=((V-Min))/((Max-Min)) → (6)
Where NV is the normalized value and V is the raw data set value.
So, finally the data set consists of only normalized value which is useful for further processing also it increases the performance during the ovarian cancer identification and classification process.
Curated ovarian data: The dataset consists of the Meta data which has the collection of gene expression data set that is used for analysing the ovarian cancer. It has the 2970 patients’ clinical Meta data set and it is implemented as the curated ovarian data (https://bitbucket.org/lwaldron/curatedovariandata) Bio conductor package for clinical analysis.
Discussions
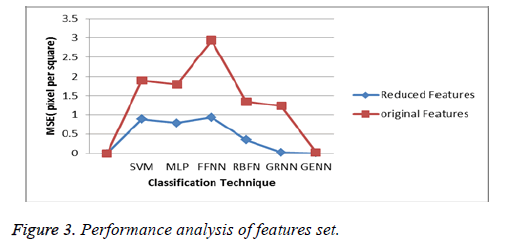
The performance of the proposed Self-Organizing Maps Immune Clonal Selection (SOMICS) and grammatical evolution neural networks is analysed with the help of the mean square error, mean absolute error, root mean square error, sensitivity, and specificity and classification accuracy. In this paper, the error rate is minimized by using the grammatical evolution neural networks, which reduces error during the weight calculation. The features selection process improves the above ovarian classification accuracy which is explained in the following results which was compared with the existing methods such as Support Vector Machine (SVM) [22], Multi- Layer Perceptron (MLP) [23], Feed Forward Neural Network (FFNN) [24], Radial Basis Function Network (RBFN) [25], General Regression Neural Network [26].
Then the above diagram clearly depicts that the reduced features set provides the better results during the classification technique (Figure 3). Then the mean square error value is shown in following Table 1.
| Classification technique | Mean square error value |
|---|---|
| SVM | 0.89 |
| MLP | 0.789 |
| FFNN | 0.934 |
| RBFN | 0.345 |
| GRNN | 0.0234 |
| GENN | 0.0021 |
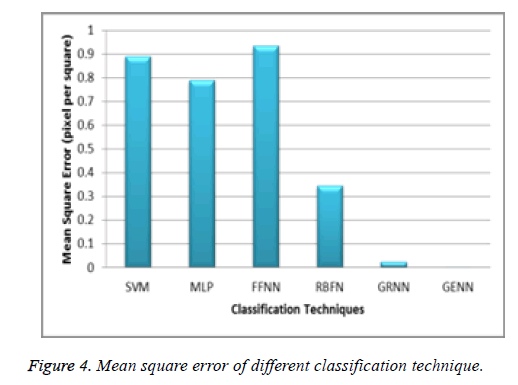
Table 1. Mean square error of different classification technique.
The above Table 1 clearly shows that the proposed GENN based classifier has the minimum mean square error which means it maximize the classification rate in the neural network. The Figure 4 shows that the proposed minimum means square error value.
Then the mean absolute error and root mean square value is calculated as follows.
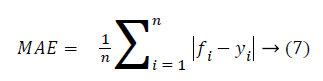
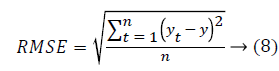
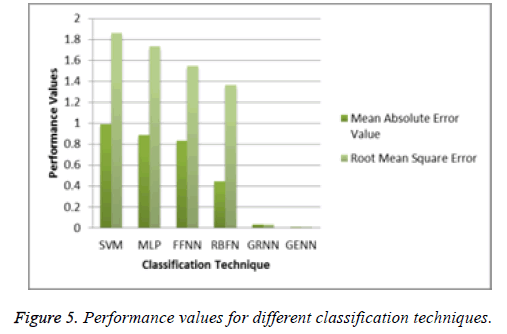
From the above Table 2, it clearly shows that the proposed system has a minimum MAE value and minimum RMSE value. The Figure 5 shows that the performance of the classification technique.
| Classification technique | Mean absolute error value | Root mean square error |
|---|---|---|
| SVM | 0.99 | 1.862 |
| MLP | 0.889 | 1.732 |
| FFNN | 0.834 | 1.543 |
| RBFN | 0.445 | 1.364 |
| GRNN | 0.0334 | 0.032 |
| GENN | 0.0022 | 0.0032 |
Table 2. Mean absolute error of different classification technique.
From the reduced mean square error, the proposed system classifies the huge data set with highest sensitivity and specificity rate. The sensitivity and specificity are calculated based on the following equation.
Sensitivity=TP/((TP+FN)) → (9)
Specificity=TN/((TN+FP)) → (10)
Where, TP=True Positive, TN=True Negative, FP=False Positive, FN=False Negative.
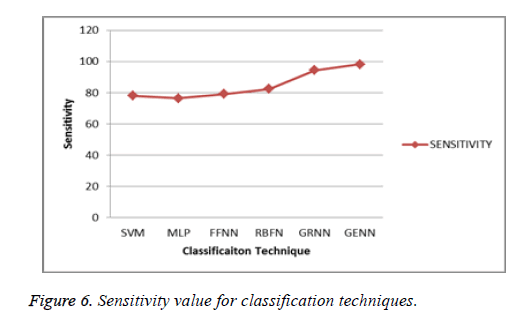
The following Figure 6 shows that the sensitivity and specificity value of the proposed system which is compared to the several classification methods such as SVM, MLP, FFNN, GRNN and GENN.
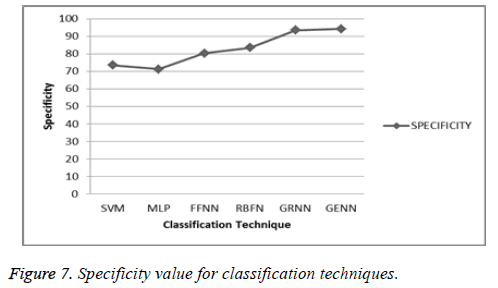
From the Figure 7, it is easy to justify that the proposed system produces the best classification result which is described by using the sensitivity and specificity. So, that classification accuracy of the proposed system in shown in following Table 3.
| S. No | Classification techniques | Classification accuracy (%) |
|---|---|---|
| 1 | SVM | 71 |
| 2 | MLP | 85 |
| 3 | FFNN | 75.6 |
| 4 | RBFN | 87.3 |
| 5 | GRNN | 93.21 |
| 6 | GENN | 98.23 |
Table 3. Classification accuracy for different classification techniques.
Thus the proposed system classifies the huge data set into ovarian cancer and normal by using the Self-Organizing Maps Immune Clonal Selection (SOMICS) and Grammatical Evolution Neural Networks (GENN).
Conclusion
Ovarian cancer is one of the critical cancers for the women which lead to death in the critical stage. Thus the paper proposed that new algorithm to identify and classify the cancer from the largest data set. From the large data set, the best sub set features are selected and grouped by using the selforganizing map and the immune clonal selection algorithm. Then the selected subsets are pre-processed by using the weighting and discretization method. The pre-processing stage removes the unwanted data also handle the missing data’s in the subset. The optimized features are selected by same selection algorithm. Finally the classification is done with the help of the grammatical evaluation neural network with highest classification accuracy. Thus the performance of the system is evaluated with the help of the experimental result.
References
- Ayako K, Yutaka U, Tetsuji N, Takayuki E. Therapeutic strategies in epithelial ovarian cancer. J Exp Clin Cancer Res 2012; 31: 14.
- Kataria, Shravan K. Role of radiotherapy in ovarian cancer. Ind J Med Paediatr Oncol 2007; 28.
- Kaveh D, Rick D, Laura H, Evan M. A branching process model of ovarian cancer. J Theor Biol 2012; 314: 10-15.
- Behtash N, Ghayouri Azar EF. Symptoms of ovarian cancer in young patients 2 years before diagnosis, a case-control study. Eur J Cancer Care 2008; 17: 483-487.
- Gemson Andrew EJ, Durga S. Big data analytics in healthcare: a survey. ARPN J Eng Appl Sci 2015; 10.
- Matthew H, Taghi MK, Randall W. A review of data mining using big data in health informatics, Herland. J Big Data 2014; 1.
- http://www.gene.com/stories/big-data-and-the-big-c
- El Sayed AW, Ibrahim Al E, Amr B. Feature selection for cancer classification: an SVM based approach. Int J Comp Appl 2012; 46.
- Renz R, Razvi L. Ovarian cancer classification with missing data. Neural 9th International Conference on Information Processing 2002; 2.
- Ankita T, Vijay M, Sunil KJ. Feed forward artificial neural network: tool for early detection of ovarian cancer. Sci Pharm 2011; 79: 493-505.
- Bellazzi R. Big data and biomedical informatics: a challenging opportunity. Yearb Med Inform 2014; 9: 8-13.
- Kamesh DBK, Neelima V, Ramya Priya R. A review of data mining using bigdata in health informatics. Int J Sci Res Publ 2015; 5.
- https://www.siam.org/meetings/sdm13/sun.pdf.
- Guangtao W, Heli S, Xueying Z. A feature subset selection algorithm automatic recommendation method. J Artificial Intel Res 2013; 47: 1-34.
- Vesanto J, Alhoniemi E. Clustering of the self-organizing map. IEEE Trans Neural Netw 2000; 11: 586-600.
- Esdras J, Pedro G, Rosa EL. The Mahalanobis distance for functional data with applications to classification. J Technometr 2015; 57: 1-37.
- Jason B. A review of the clonal selection theory of acquired immunity. Compl Intel Sys Lab 2007; 1-6.
- Muhammad Shoaib BS, Iqbal G, Laurence D. Communal neural network for ovarian cancer mutation classification. GSCIT Monash Univ 2005; 12: 1-10.
- Khaled AA, Abdul-Kader HM, Nabil AI. Artificial immune clonal selection classification algorithms for classifying malware and benign processes using API call sequences. IJCSNS Int J Comp Sci Netw Secur 2010; 10.
- SagarImambi S, Sudha T. Pre-processing of medical documents and reducing dimensionality. Adv Comp Int J 2011; 2.
- Ilia M, Krassimira I, Krassimir M, Vitalii V, Peter S, Koen V. Comparison Of discretization methods for preprocessing data for pyramidal growing network classification method. Int Conf Tec 2009; 31-39.
- Terrence SF, Nello C, Nigel D, David WB, Michel S, David H. Support vector machine classification and validation of cancer tissue samples using microarray expression data. Bioinformatics 2000; 16: 906-914.
- Renz C, Rajapakse JC, Khalil R. Ovarian cancer classification with missing data. Neural Information Processing 2002.
- Mabvuure NT, Klimach S, Eisner M, Rodrigues JN. An audit of best evidence topic reviews in the International Journal of Surgery. Int J Surg 2015; 17: 54-59.
- Chen CL, Weng KL, Chee PL. Dimensionality reduction of protein mass spectrometrydata using random projection. J Am Soc Mass Spectrom 2015; 26: 315-322.
- Feng C, Lipo W. Applying RBF neural networks to cancer classification based on gene expressions. Int Joint Conf Neur Netw 2006.