Short Communication - Journal of Cell Science and Mutations (2017) Volume 1, Issue 1
Comparison of a kanamycin versus hygromycin resistance gene in transgenic plant selection of Arabidopsis thaliana L.
Abdur Rashid*
Department of Biological Sciences, University of Alberta, Edmonton, Alberta, Canada
- *Corresponding Author:
- Abdur Rashid
Department of Biological Sciences
University of Alberta
Canada
Tel: 780-492-3111
E-mail: Abdur.Rashid@ARS.USDA.GOV
Accepted date: February 24, 2017
Citation: Abdur Rashid. Comparison of a kanamycin versus hygromycin resistance gene in transgenic plant selection of Arabidopsis thaliana L.. Adv cel sci tissue cul 2017;1(1):1-3.
DOI: 10.35841/cell-science.1.1.1-2
Visit for more related articles at Journal of Cell Science and MutationsShort Communication
The neomycin phosphotransferase II (nptII) and hygromycin phosphotransferase II (hptII) genes, originally isolated from E. coli [1,2] are widely used as selectable markers in transgenic research [3-5]. The recovery of transgenic plants in the presence of respective selection agent has been greatly facilitated by these genes. The nptII gene confers transgenic plants resistance to the antibiotic, kanamycin, kan, and the hptII gene provides transgenic plants resistance to the antibiotic, hygromycin, hyg [3,5]. These antibiotics normally inhibit protein synthesis in plants by binding to ribosomes [5]. However, in the transgenic plants, the encoded enzymes, NPTII and HPTII catalyze the phosphorylation of kan and hyg respectively, once they enter into plant cells [3,5]. The phosphorylated kan and hyg are inactive and therefore, unable to shutdown the protein synthesis machinery in plants. This enables transgenic plants to grow in the presence of these antibiotics. Because there is no endogenous NPTII or HPTII activity in plants, these antibiotics can normally inhibit the growth of non-transgenic plants [5].
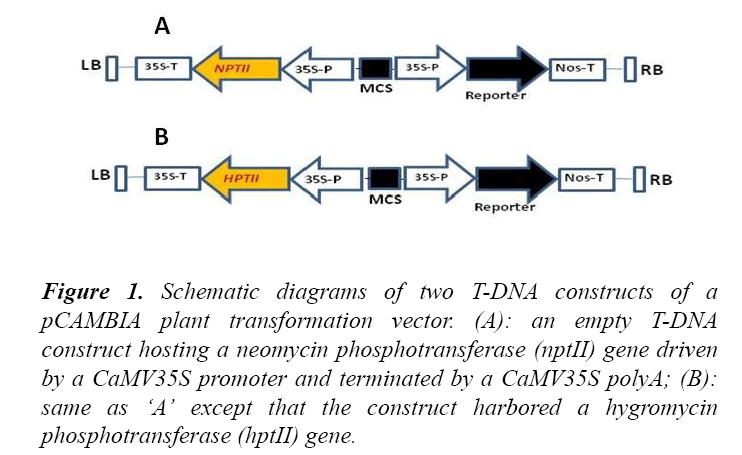
To compare the phenotypic differences of non-transgenic and transgenic plants in the presence of kan or hyg, transgenic plants were developed by Agrobacterium-mediated transformation of Arabidopsis thaliana plants as described previously [5] with a pCAMBIA vector construct hosting either a nptII or a hptII gene in the T-DNA region (Figure 1). While the mechanism by which kan and hyg exert inhibitory action on non-transgenic plants has been suggested to be similar [5], the phenotypic appearance of the plants growing in MS-agar plates containing either kan or hyg was not the same. For instance, kan caused a bleaching effect on non-transgenic A. thaliana L. plants, whereas hyg did not cause similar effect (Figure 2).
Figure 1: Schematic diagrams of two T-DNA constructs of a pCAMBIA plant transformation vector. (A): an empty T-DNA construct hosting a neomycin phosphotransferase (nptII) gene driven by a CaMV35S promoter and terminated by a CaMV35S polyA; (B): same as ‘A’ except that the construct harbored a hygromycin phosphotransferase (hptII) gene.
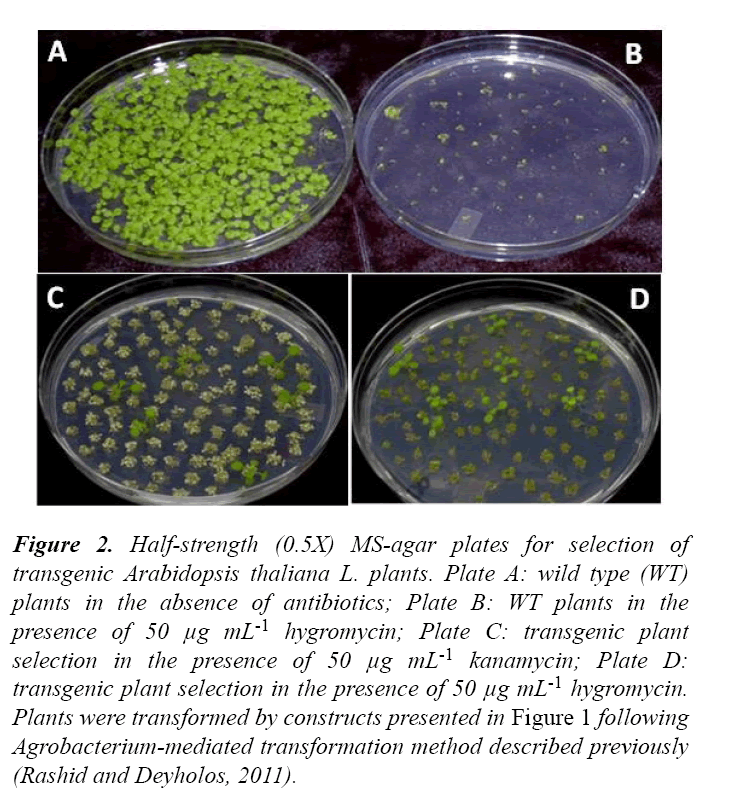
Figure 2: Half-strength (0.5X) MS-agar plates for selection of transgenic Arabidopsis thaliana L. plants. Plate A: wild type (WT) plants in the absence of antibiotics; Plate B: WT plants in the presence of 50 μg mL-1 hygromycin; Plate C: transgenic plant selection in the presence of 50 μg mL-1 kanamycin; Plate D: transgenic plant selection in the presence of 50 μg mL-1 hygromycin. Plants were transformed by constructs presented in Figure 1 following Agrobacterium-mediated transformation method described previously (Rashid and Deyholos, 2011).
When grown in 0.5X MS-agar plates with a concentration of either kan or hyg (50 μg mL-1) that discriminates between transgenic and non-transgenic plants, seeds of non-transgenic plants had germinated, grown up to two-leaf stage, and then bleached out and died in the presence of kan, but were either not germinated or germinated but failed to grow and died in the presence of hyg (Figure 2). Although the above observations are in agreement with previous reports indicating that hyg is potentially more toxic to non-transgenic plants than kan (for a review, 5), the current research showed that the hyg resistant transgenic plants produced more roots in hyg plate as compared to kan resistant transgenic plants in kan plate (Figure 2).
In conclusion, the transgenic selection plates presented in this communication (Figure 2C and 2D) are unique and would serve as a clear guidance for the researchers carrying out research in this field involving Arabidopsis thaliana L. as a model plant, nptII and hptII genes as selectable markers, and kan and hyg, respectively as selection agents.
References
- Berg DE, Davies J, Allet B, Rochaix JD. Transposition of R factor genes to bacteriophage lambda. Proc Natl Acad Sci. 1975;72(9):3628-3632.
- Gritz, L, Davies J. Plasmid-encoded hygromycin B resistance: the sequence of hygromycin B phosphotransferase gene and its expression in Escherichia coli and Saccharomyces cerevisiae. Gene. 1983;25(2-3):179-188.
- Ziemienowic A. Plant selectable markers and reporter genes. Acta Physiol. Plant. 2001;23(3):363–374
- Rashid A. Deyholos MK. PELPK1 contains a unique pentapeptide repeat and is a positive regulator of germination in Arabidopsis thaliana. Plant Cell Rep. 2011;30(9):1735-1745.
- Davey MR, Sonej JR, Rao MN, Kourmpetti S, Bhattacharya A. Generation of development of transgenic crop plants: an overview. Transgenic Crop Plants. 2010;1(1):1-30.
- Rashid A, Deyholos MK. PELPK1 contains a unique pentapeptide repeat and is a positive regulator of germination in Arabidopsis thaliana. Plant Cell Rep. 2011;30:1735-1745.