Research Article - Journal of Neuroinformatics and Neuroimaging (2017) Volume 2, Issue 1
COINS Tools for Automated NIMH Data Archive Submissions
Miller BN1*, Wang R1, Kelly R1, King MD1, Lake J1, Landis D1, Ragle J1, Reed C1, Stone M1, Dieringer C1, Calhoun VD1,21The Mind Research Network, 1101 Yale Boulevard NE, Albuquerque, NM, 87106, USA
2Departments of Electrical and Computer Engineering, Neurosciences, Computer Science, and Psychiatry, University of New Mexico, 1155 University Blvd SE, Albuquerque, NM, 87106, USA
- *Corresponding Author:
- Brittny Miller
1101 Yale Boulevard NE
Albuquerque, NM 87106 USA
Tel: 5052725028
Fax: 5052728002
E-mail: bmiller@mrn.org
Accepted date: February 06, 2017
Abstract
Grant funding agencies are continuing to move toward their own standardized database platforms in the pursuit of harmonizing data and reproducing results. The National Institute of Mental Health Data Archive (NDA) is one such mission. This initiative serves as a repository for data collected by investigators who have been funded by the NIH/NIMH and have committed to sharing their data with the NDA. Preparing data captured by assessment and imaging collection and management systems (e.g. XNAT, Qualtrics, and REDcap) for submission can be challenging. NDA data submissions are performed through standalone GUI-only applications, limiting automation possibilities. The Collaborative Informatics and Neuroimaging Suite (COINS), a web-based data collection and management platform that employs integrated assessment and imaging collection, has developed tools to assist in assessment and imaging submissions to NDA repositories. COINS is able to assist in the creation of the NDA participant identifier, Global Unique Identifier (GUID). The NDA application programmer interface (API) is then utilized to map COINS instrument questions and scan series to NDA data elements to provide a series of exports that can be directly submitted to the NDA applications. These exports do not require any manual manipulation or use of scripts to conform data to NDA standards. In July 2016, the first assessment submission using COINS tools was completed and the first imaging submission was performed in January 2017.
Keywords
Neuroinformatics, Data sharing, Coins, Brain imaging, Database.
Introduction
Producing reproducible results is a standard in scientific research, leading to more journals and funding agencies promoting or requiring data sharing for secondary analysis on datasets [1-3]. A challenge to this is that neuroimaging datasets are large in size and have no standard method of storage hierarchy or file naming conventions [4]. This is also true of the phenotypic measures generally collected alongside neuroimaging data. Although many of these measures are standardized, each research group tends to name variables in their own way. When this data is combined with similar data from other groups or is shared, it is often difficult to make sense of it [5]. Many initiatives, such as National Institute of Mental Health Data Archive (NDA) (https://data-archive.nimh.nih.gov/), Federal Interagency Traumatic Brain Injury Research (FITBIR) (https://fitbir.nih.gov/), and Brain Imaging Data Structure (BIDS) [6], have been started to help harmonize data, making sharing and second analysis easier. The goal of increasing sharing and reproducibility is at the forefront for the NDA. The NDA encompasses several initiatives, including National Database for Autism Research (NDAR) [7], Research Domain Criteria (RDoC), and National Database for Clinical Trials Related to Mental Illness (NDCT). Before submitting data, the NDA GUID Tool is used to generate Global Unique Identifiers (GUIDs) for participants whose data is to be submitted to the database. Then researchers are to use the NDA Validation and Upload Tool to begin the data submission process. The NDA Validation and Upload Tool will ensure the data queued for submission is properly formatted, conforms to the corresponding data dictionaries, and has valid GUIDs. The NDA does not provide or require any particular data collection tools, allowing researchers to use collection platforms they are familiar with or prefer. A complication of this is that researchers are then required to map their various forms of collected data to the NDA Data Dictionary form elements. This has led many researchers to manipulate their data by-hand or with complicated scripts, both of which can be error prone and are time consuming to perform and write. The Collaborative Informatics and Neuroimaging Suite (COINS) [8], a centralized, web webbased data collection and management platform designed for integrated assessment and imaging collection, have developed automated tools to help researchers overcome the challenges of mapping their data to NDA specifications. COINS also provides many study management tools, automated participant assessment queuing, granular querying abilities, data sharing capabilities, and a permission system designed to protect personal identifying information (PII). Currently COINS is serving over 600 studies consisting of over 580,000 completed participant questionnaires, 46,000 MRI and MEG scan sessions from more than 42,000 participants and is continuing to grow. As a data collection platform dedicated to advancing research, COINS is continuously seeking integrations with other neuroinformatics services and databases. This has led to the development of automated COINS tools that allow users to collect and store GUID information as well as map and export assessment and MRI scans to NDA Data Dictionary specifications.
Materials and Methods
A Review of COINS
COINS was developed at the Mind Research Network (MRN) to provide a centralized repository to capture all the data collected during multisite brain imaging studies [8]. The goal was to have a centralized place for all sites to access imaging and assessment data in real-time, as well as reporting and other functionality. This homegrown effort has led to the development of several different tools that work together to provide a comprehensive participant and data management system, while streamlining the collection, storage, and sharing of data. All tools have a front end interface, eliminating the need for users to have any programing knowledge. There are many checks and balances in place to ensure study staff can only access designated participant information, whether it be PII, assessment, or imaging data. Site administrators also have tools to ensure only studies that are compliant with their IRB requirements are able to enroll participants and collect data, preventing violations from occurring.
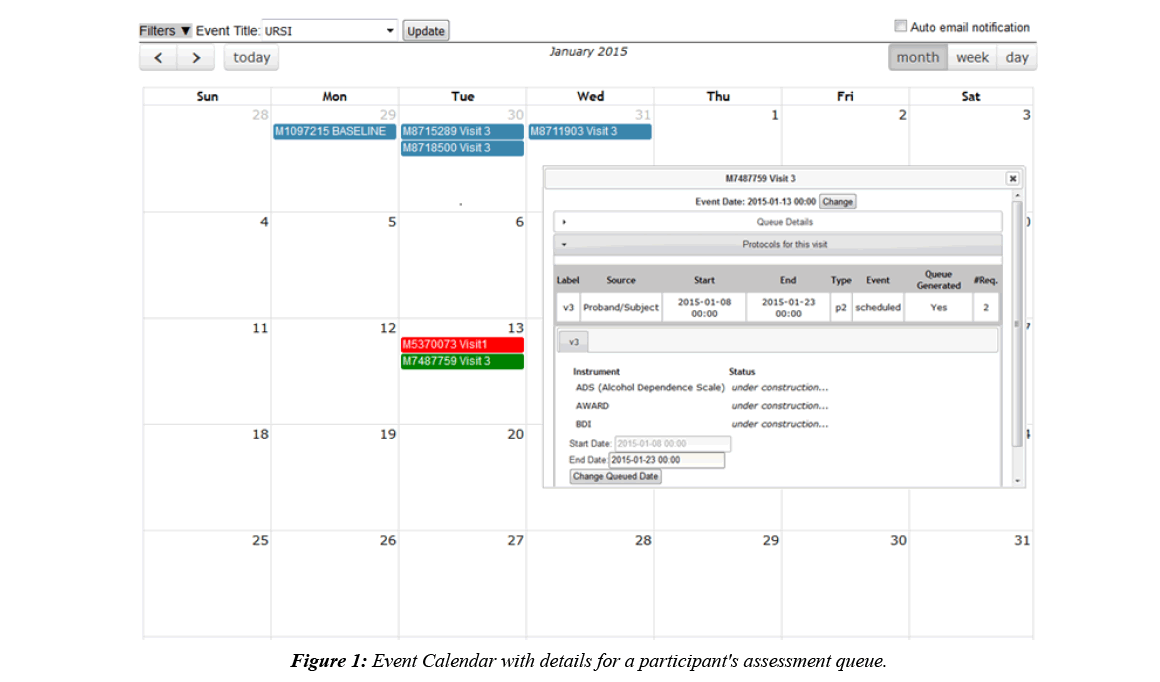
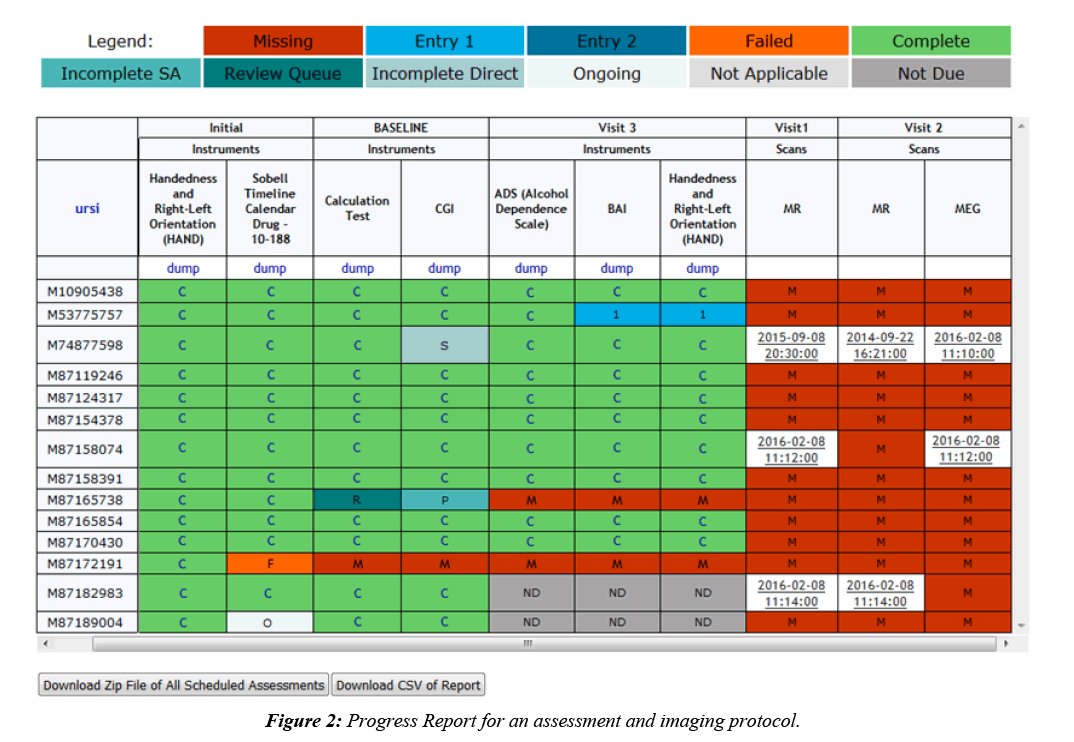
Imaging collection is done by a DICOM receiver that automatically validates scan data by checking participant enrollment and study IRB status, then transfers the DICOMs to an archive location of choice and stores scan metadata in the database. DICOMs can also be imported through the web interface, allowing for the same archiving and metadata storage. Once the data is stored, fMRI behavioral data can be associated with it, scan comments can be recorded, and radiologists can perform reviews. Assessment data is typically collected alongside imaging data, which is why COINS supports very robust assessment administration tools, allowing researchers to take control over the way their data is formatted [9]. Assessment data is entered into instruments that can be copied from an existing library or handcrafted, using many question types and features like, automatic calculations, skip logic, critical response flagging and text formatting. Study staff can enter data directly into the instruments or use the dual entry system for recording paper assessments. Participants can also directly enter data by logging into an instrument queue created by staff or a queue that has been automatically generated via the auto queuing system. The auto queuing system uses a combination of study defined visits, subject types, and assessment protocols to generate instrument queues that are accessible by participants for a set time frame. The auto queuing feature works with an event calendar that allows study staff to view and schedule visits and provides the ability to send automatic email reminders to participants (Figure 1). Assessment and imaging data that has been collected can be managed through a series of tools that allow study staff to validate the collected data and track the collection process. A progress report is used to provide a visual representation of the study’s data collection schedule and compliance for each participant (Figure 2). As soon as data collection starts, de-identified assessment and imaging data can be exported separately or together for analysis through a very granular query tool. Data sharing can also be performed through Data Exchange, which allows users from all over the world to query and download anonymized datasets [10].
GUID
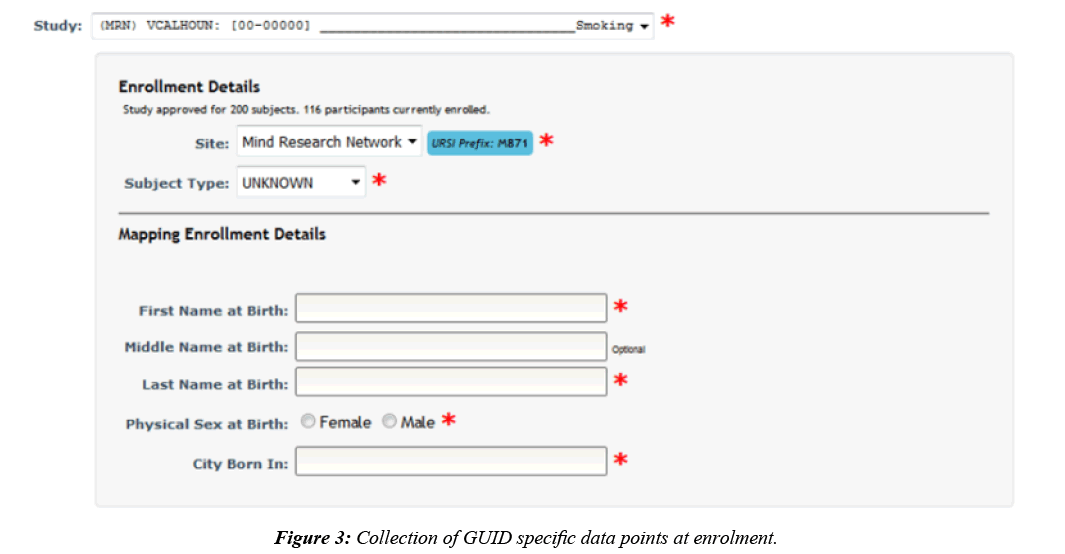
To submit data to the NDA, it is required that each participant has an assigned GUID [11], which is the NDA’s participant identifier. GUIDs are created by entering or importing a participant’s personal information to a downloadable, locally installed application called the GUID Tool. A user can generate GUIDs through the application by entering participants oneby- one, which requires double entry of the required fields, or in a batch via CSV. Basic participant information is already collected in COINS when a participant is enrolled into a study, generating a Unique Research Subject Identifier (URSI). URSIs are the COINS participant identifier and are required to be created prior to data collection. Although participant details are collected at the time of enrollment, GUID creation requires first and last name at birth, physical sex at birth, and city born in, which are not details collected in COINS at the time of participant enrollment. To accommodate the collection of this extra required information, upon enrollment study staff are asked to also include the GUID required fields for any participant being enrolled into a study configured as an NDA study (Figure 3). The GUID specific data is stored in the COINS database alongside the default COINS enrollment information.
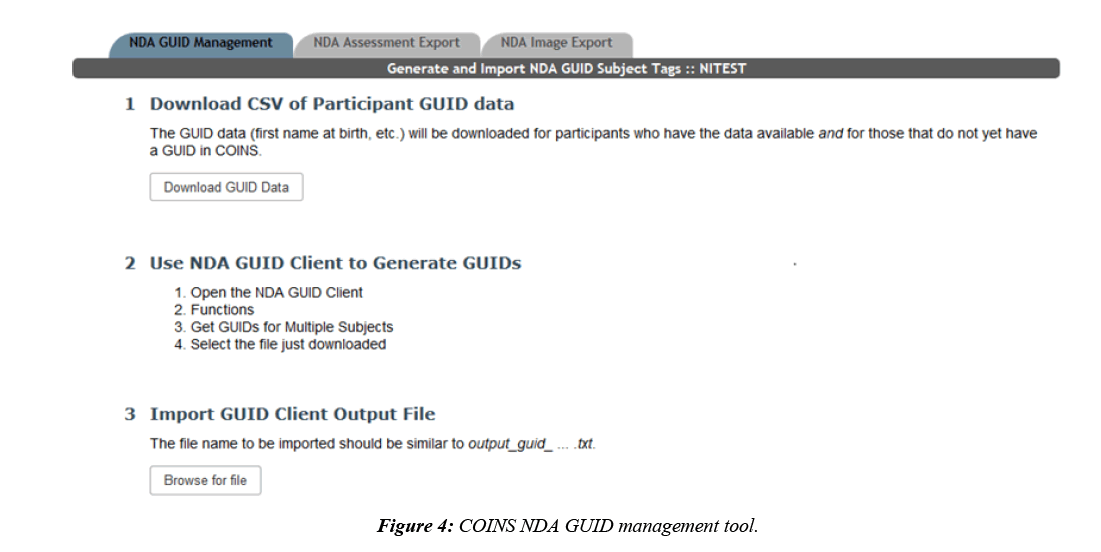
At this time, there is no automated way for COINS to directly interact with the NDA GUID Tool, so COINS relies on the batch import CSV for GUID creations. COINS provide the completed batch CSV that is ready to be directly uploaded to the NDA GUID Tool. Once the CSV is submitted, the NDA GUID Tool outputs a text file with the resulting GUIDs for each submitted participant. This text file can then be imported back into COINS and each NDA GUID will be linked to the COINS URSI using a subject tag (Figure 4). Subject tags in COINS are used as an alternative way to label participants. If a study tracks participants by an identifier other than the URSI, this feature can be used to document the link between the URSI and the other identifier. Participants can be looked up by subject tag in addition to URSI in many COINS tools. Once the text file has been uploaded to COINS, the subject tag link will exist for each submitted URSI. Only participants with completed NDA specific enrollment fields who do not already have a GUID subject tag are exported for each NDA study. This prevents participants who have already been assigned a GUID and any participants with missing GUID required fields from being submitted or re-submitted.
Instrument Mapping and Export
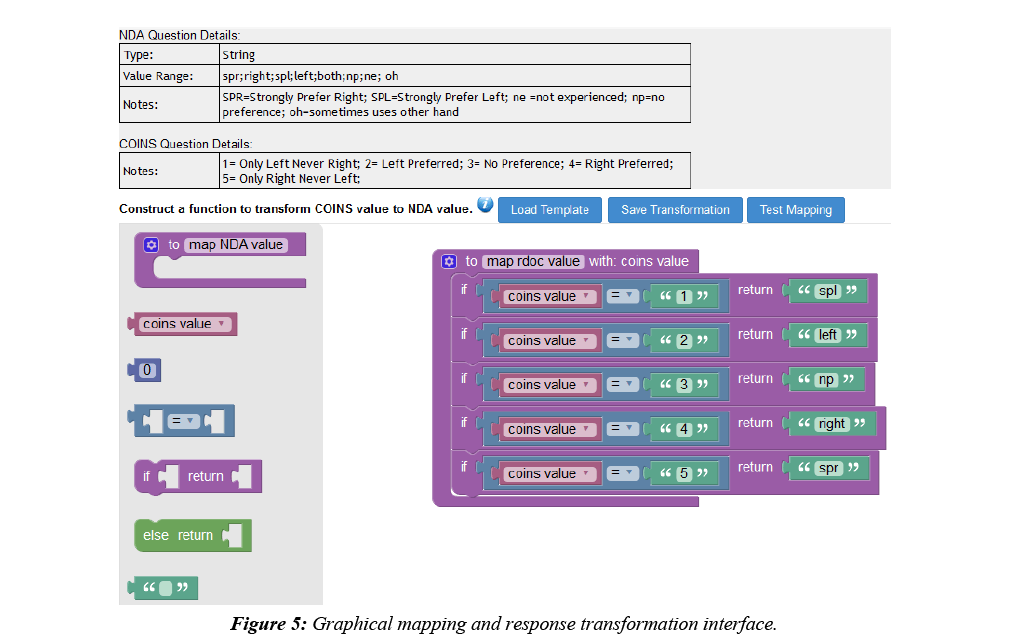
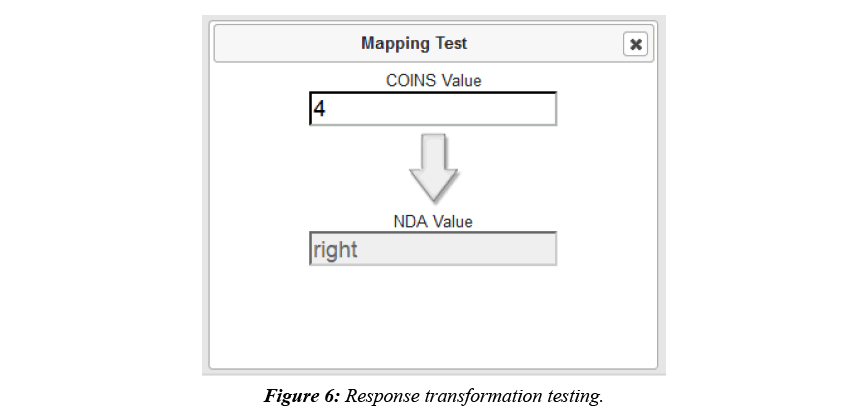
COINS allow users to easily create, import, copy, or share data collection instruments. These instruments can be used for data entry or for participant self-assessment. After instruments have been set up for NDA studies in COINS, users can start mapping the COINS instrument questions to NDA Data Dictionary questions and performing response value transformations. In COINS, the mapping can be done individually, or in batch. In doing individual question mapping, the list of COINS questions for the instrument and list of NDA data structures and item level elements are presented for the user to select. The list of NDA data structures and elements are fetched through the NDA API in situ. Basic information (i.e. data type, value range, etc.) for the NDA data element are also shown to the user to assist with the mapping process. After the user selects the targeted pair of questions for mapping, a graphical drag-and-drop mapping interface, implemented with Google’s open source Blockly library (https://developers.google.com/blockly/), will become available. Users can then build the response transformation mapping from scratch or edit the pre-built, sample template provided. This response mapping allows users to transform the stored COINS value to match the NDA Data Dictionary value (Figure 5). Before saving a response transformation mapping, users can test the mapping function to be sure the transformation is valid and producing the intended values (Figure 6).
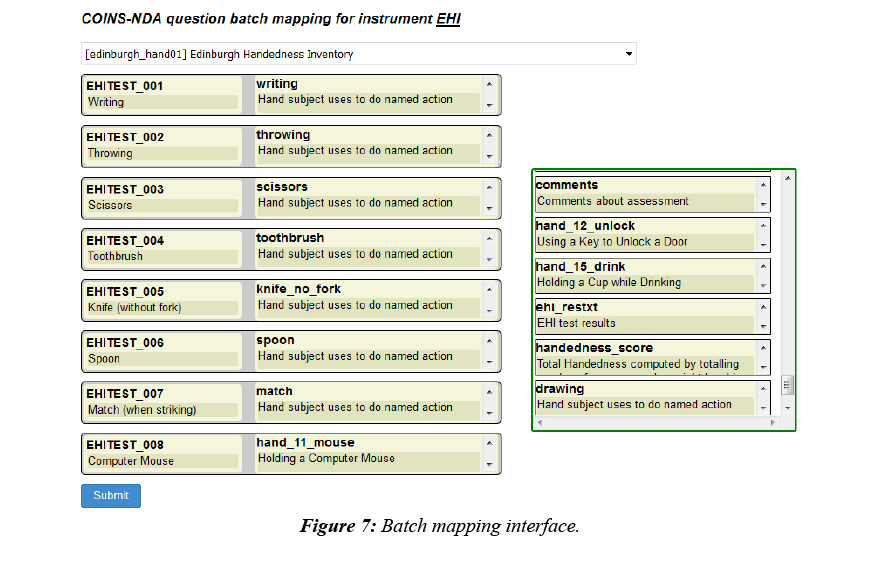
To facilitate a quicker mapping process, COINS also provides batch mapping functionality. This feature allows users to map multiple pairs of questions in one submission (Figure 7). The questions on the left are questions from a COINS instrument, listed in the order they are stored in the database. After users select the NDA data structure the NDA elements will be automatically populated in the order they are listed in the data structures and aligned with the COINS instrument questions. Users can then drag and drop the NDA questions to adjust the matching pairs. Once pairs are defined, the user can perform question response transformations.
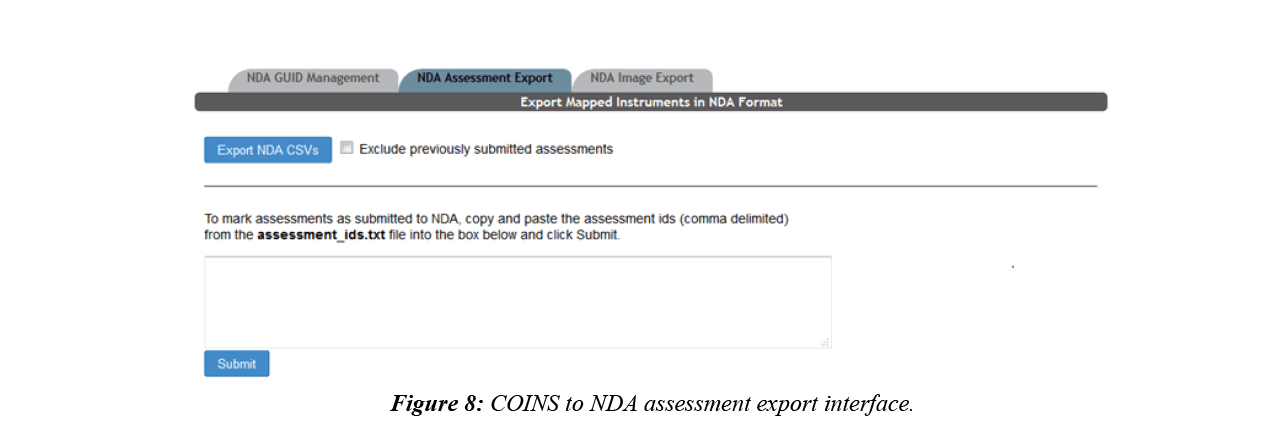
After the mappings between an NDA data structure and a COINS instrument is set up, the user will then be able to export the assessments from COINS, which will conform to the NDA submission format without further modification. COINS also enables the user to record which assessments have been submitted (Figure 8). The previously submitted assessments can be omitted from future export, allowing users to control which data is submitted.
Imaging Mapping and Export
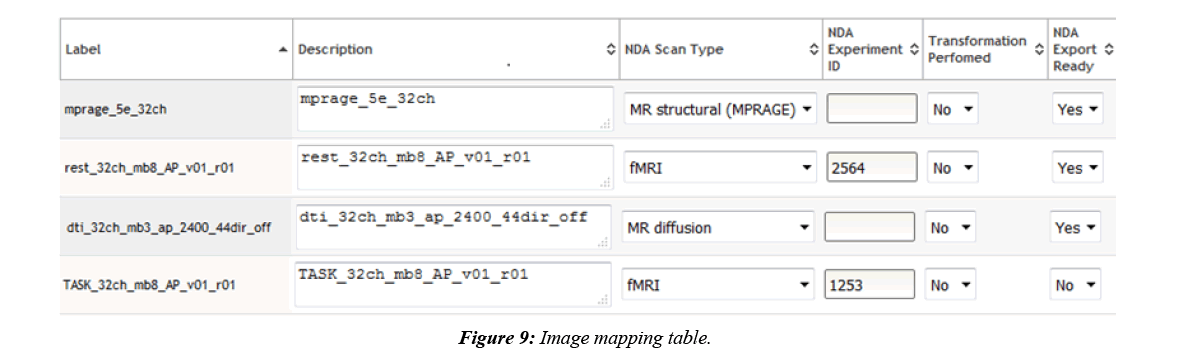
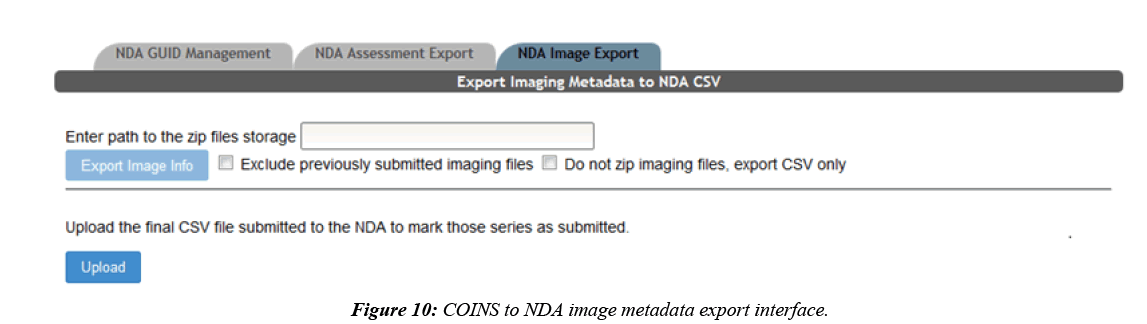
COINS are also able to export MRI imaging metadata that conforms to the NDA format. Most of the imaging metadata required for the NDA CSV is already collected in the COINS database when the imaging data was transferred. Some extra metadata is required by the NDA that is not collected in COINS, such as fMRI experiment ID and if transformations were performed to the imaging data. COINS provide a series/ label management tool for users to enter this extra information, and to control which imaging series should be exported for NDA submission (Figure 9). When a user submits the imaging metadata export request, they will get a CSV containing all requested information in minutes, while the requested raw imaging data is zipped on the backend. Similar to the assessment export, users are able to mark series already submitted to the NDA by importing the CSV submitted to the NDA back into COINS (Figure 10). This will ensure that these submitted series will not be exported again in the future.
Results
This development has resulted in COINS providing GUID, assessment, and imaging CSVs that can be directly submitted to NDA tools with little or no user manipulation of the data. There are currently four NDA studies in COINS. Over 660 GUIDs have been created using the COINS GUID export and have been added to participants as subject tags. In July 2016, the first submission using COINS tools were performed to the RDoC database consisting of 53 subjects and from 18 different data structures (697 elements or assessments total). A second submission was made in January 2017 consisting of 59 participants. Also in January, the first image submission was performed, which included almost 4,400 MRI imaging series.
Interfacing data collection platforms with sharing initiative databases is achievable. The easy-to-use COINS support tools help to lessen the burden placed on study staff to prepare a submission. An automated submission process that does not require exports can be implemented when a method for direct interaction with NDA repositories is supported. Until this time, these front-end COINS web tools give researchers a hasslefree way to continue to collect their data in formats of their choosing, and the ability to conform to NDA data dictionaries. This implementation eliminates the need for the use of scripts, which need to be created by each user, are extensive, and can be error prone, or sending your data to a paid submission service to be organized and reconciled.
Discussion
The development of these features was driven by a study being conducted (MRN). This study uses self-assessment along with direct and dual entry to collect assessment and imaging data. During their first submission using COINS, it was discovered that a COINS instrument that needed to be submitted lacked the appropriate NDA data structure mappings. The mappings were put in place in minutes and the data was properly exported and validated for submission. However, this first submission brought up a challenge that was unexpected, which was that users need more control over which assessments are exported. Tools had been created to exclude assessments from export, but no tools existed to re-include assessments in the export. This was an obstacle because this particular study needed to submit current assessments for participants that had completed the study. If a participant was still in the follow up phase, their data was to not be submitted until the next submission. Study staff were able to exclude the assessments for participants in the follow-up phase from exporting in this submission, however, on the front end there was no way for them to re-include the assessments so they would export for the next submission. This led to the creation of better tools that give users more control over which assessments are exported for submission. These tools also increase transparency, allowing the users see to which assessments are flagged for export and which are not. These new developments will give users the power to manage their own data without needing additional back end help. Much like the first assessment submission, the first MRI submission helped to shed light on needed tools that increase user control. It was apparent that users need an easier way to include and exclude MRI series in the export, similar to the including and excluding of assessments. Also, there are new required fields for certain scan types that did not exist when the COINS tools were originally built, requiring the manual addition of these fields for this one scan type. These items will be addressed in future development.
The development of the NDA tools has led to requests for interacting with other data sharing initiatives. A TBI study being conducted at MRN has requested similar mapping tools for the FITBIR data sharing initiative. Since both initiatives are from the NIH, the thought was that both would be structured similarly and NDA tools could easily be reused for FITBIR mapping. However, it was quickly learned this is not the case and that there are many differences between the two initiatives. GUID requirements and resulting text file outputs were the first differences encountered. Compared to the NDA initiative, FITBIR currently does not provide the APIs for fetching any instrument or question data contained in their data dictionaries, presenting additional difficulty. The current solution is to use scripts to scrape the FITBIR webpages and gather all the useful data, which is then saved into COINS database for mapping purposes. It was also noticed that, in contrast to the NDA instruments where one question only appears once in one data dictionary, in a FITBIR data dictionary questions are organized by group and the same question can appear multiple times in different groups. These differences will require tweaks to the implementation for providing a similar GUID collection and assessment mapping interface.
Limitations
Upon starting the development of these tools, it was quickly realized that there were going to be several obstacles to overcome to make them work intuitively with minimal user effort. As development of these tools has continued to grow, it is becoming apparent that limitations exist in the current architecture. One limitation is that a COINS instrument can only be mapped to one data sharing initiative. Although this has not been an issue as of yet, it is potentially problematic in the future, however it can be addressed with additional development. The submission of DICOM imaging data is another difficult task to tackle. Submission of this data requires each scan task or series to be zipped and the path for the zipped DICOM series needs to be added to the CSV. This path will then be used to locate the referenced imaging data for upload to the NDA. Performing this for MRN users is a simple task because the MRI data is onsite. This becomes more difficult when dealing with users from other sites, because they need to be able to reference the zipped data path with their local computers, which they cannot do if the data is zipped and stored at MRN. Most COINS sites typically store their data locally and at MRN as a backup. Since the data local to the site is transferred with the COINS DICOM receiver, it is archived in the same storage hierarchy as the data stored at MRN. This simplifies the approach as the same process that zips the DICOM data at COINS can also be used at the various sites so the files can be zipped locally and referenced properly in the CSV. This, however, does not solve the problem for sites that only store their imaging data with COINS and do not have local copies. A solution would need to be put into place in the event a study was not storing imaging data locally. Solutions could include rsyncing zip files to the site, a method of download from COINS, or even a cloud-based file access tool until the zip files can be directly sent to the NDA from COINS.
Through the many challenges, the biggest hurdle has been the lack of a method for automated data validation and submission, forcing data to be exported into CSVs that can then be submitted to the NDA. Fortunately, this is something the NDA has committed to improving. They are currently working on updating their API to allow automated validation and submissions (https://data-archive.nimh.nih.gov/API). This will eliminate the need to use CSVs and the NDA Validation and Upload Tool, allowing for easier and quicker submissions. Once these changes are in place, they will be integrated into COINS, allowing users to do direct submissions from COINS to NDA with ease.
Acknowledgements
The COINS development team would like to thank Dan Hall and Svetlana Novikova from the NDA for their guidance as we have built these interfacing tools.
References
- Savage CJ, Vickers AJ. Empirical study of data sharing by authors publishing in PLoS journals. PLoS One, 2009; 4(9): p.7078.
- Poline JB , Poldrack RA. Frontiers in brain imaging methods grand challenge. Front Neurosci. 2012; 6:p.96.
- Eickhoff S, Nichols TE, Van Horn JD, et al. Sharing the wealth: Neuroimaging data repositories. Neuroimage. 2016; 124(Pt B): p.1065-8.
- Van Horn JD, Gazzaniga MS. Why share data? Lessons learned from the fMRIDC. Neuroimage. 2013; 82: 677-682.
- Poline JB, Breeze JL, Ghosh S, et al., Data sharing in neuroimaging research. Front Neuroinform. 2012. 6: 9.
- Gorgolewski KJ, Auer T, Calhoun VD et al. The brain imaging data structure, a format for organizing and describing outputs of neuroimaging experiments. Sci Data. 2016; 3: 160044.
- Hall D, Huerta MF, McAuliffe MJ et al., Sharing heterogeneous data: the national database for autism research. Neuroinformatics, 2012; 10(4): 331-339.
- Scott A, Courtney W, Wood D. COINS: An Innovative Informatics and Neuroimaging Tool Suite Built for Large Heterogeneous Datasets. Front Neuroinform. 2011; 5: p. 33.
- King MD, Wood D, Miller B, et al. Automated collection of imaging and phenotypic data to centralized and distributed data repositories. Front Neuroinform, 2014; 8: 60.
- Landis D, Courtney W, Dieringer C, et al., COINS Data Exchange: An open platform for compiling, curating, and disseminating neuroimaging data. Neuroimage, 2016; 124(Pt B): 1084-1088.
- Whitney G, Johnson SB, McAuliffe M. Using global unique identifiers to link autism collections. J Am Med Inform Assoc. 2010; 17(6): p. 689-695.