Research Article - Biomedical Research (2017) Volume 28, Issue 6
Two novel Ni (II) complexes with two different Schiff bases: inhibiting glioma cells growth
Guohu Weng1,2, Suyue Pan1*, Bo Zhou3, Guoqiang Wen2 and Shixiong Huang21Department of Neurology, Nanfang Hospital, Southern Medical University, Guangzhou, Guangdong, PR China
2Department of Neurology, People’s Hospital of Hainan Province, Haikou, Hainan, PR China
3Department of Intensive Care Unit, the First Affiliated Hospital of Human University of Chinese Medicine, Changsha, Hunan, PR China
- *Corresponding Author:
- Suyue Pan
Department of Neurology
Nanfang Hospital
Southern Medical University, PR China
Accepted on November 03, 2016
Abstract
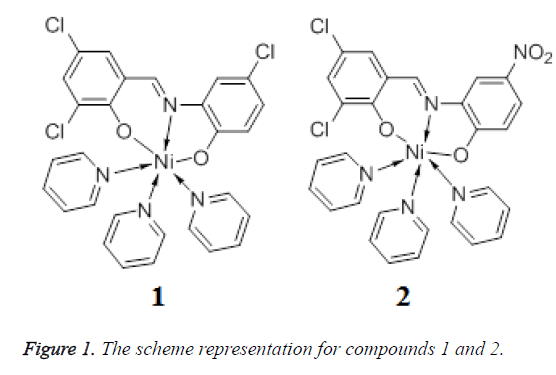
By using two flexible Schiff bases H2L1 and H2L2, two new Ni(II)-coordination compound, namely, Py3NiL1 (1) and Py3NiL2 (2) (Py=pyridine, L1=3,5-Cl2C6H2(O)C=NC6H3(O)-4-Cl, L2=3,5- Cl2C6H2(O)C=NC6H3(O)-4-NO2) have been synthesized under solvothermal conditions. Single crystal X-ray structural analysis reveals that compounds 1 and 2 are both six-coordinate in a distorted octahedral geometry. For 1, the 1D chain-like structure was observed by the C-H…π interaction, and the 1D ribbon-like structure was formed by the C-H…π and π…π interaction; for 2, the 1D ribbon-like structure was formed by the C-H…O hydrogen bonding interaction, and the π…π interaction on the side of 1D ribbon-like structure also result in the formation of a 2D structure. The in vitro antitumor activities of 1, 2 and their corresponding organic ligands Py, L1 and L2 were studied and evaluated, in which three human glioma cells (U373, U251 and SHG44) were used in the screening tests.
Keywords
Schiff base, Octahedral, Antitumor.
Introduction
A glioma is a type of malignant brain tumour. A malignant tumour is a mass of abnormal cells that is cancerous [1]. Tumours can develop in any part of the brain or its nerves and covering tissues. The two major types of brain tumour are primary and secondary. Primary brain tumours start in the brain. Secondary brain tumours start in another part of the body, then spread to the brain [2,3]. A glioma is a primary brain tumour, accounting for 45% of cancers that begin in brain cells. The three main types of glioma include: astrocytoma, ependymoma, andoligodendroglioma. Each of these types can be assigned a grade, either low grade or high grade, with high grade being more malignant and aggressive [4].
Schiff bases have played an important role in the development of coordination chemistry since the late 19th century [5]. The metal complexes with Schiff bases have been widely investigated for their versatile structures and potential applications in the fields of medicine, catalyst, and magnetism [6,7]. Furthermore, nickel is one of the most important transition metal ions in biological systems and plays an important role in transport and activation of molecular oxygen [8]. In this work, two new Ni(II) complexes Py3NiL1 (1) and Py3NiL2 (2) (Figure 1) (Py=pyridine, L1=3,5- Cl2C6H2(O)C=NC6H3(O)-4-Cl, L2=3,5- Cl2C6H2(O)C=NC6H3(O)-4-NO2) were solvothermally prepared by employment of two different Base ligands and their antitumor activities was then evaluated.
Experimental
Apparatus and materials
All the starting materials and reagents used in this work were obtained commercially and used without further purification. Element analyses (C, H and N) were determined with an elemental Vairo EL III analyser [9]. Powder X–ray diffraction data were collected using PANalytical X'Pert Pro powder diffractometer with Cu-Kα radiation and 5° ≤ 2θ ≤ 50°. Thermogravimetric experiments were performed using a TGA/ NETZSCH STA449C instrument heated from 30 to 800°C (heating rate of 10 °C/min, nitrogen stream). Single crystal X-ray diffraction was carried out by an Oxford Xcalibur E diffractometer.
Three human glioma cells (U373, U251 and SHG44) were obtained by paying to Institute of Basic Medical Sciences (IBMS) of Chinese Academy of Medical Sciences (CAMS) (Beijing, China).
Synthesis and characterization of Py3NiL1 (1) and Py3NiL2 (2)
A mixture of 3,5-dichlorosalicylaldehyde-2-amino-4- chlorophenol Schiff base (H2L1, 1.0 mmol, 0.316 g) and 25 ml methanol were mixed in the reaction flask. After heating and dissolving in air, 20 ml methanol solution of NiCl2 (50 mmol/L) was added to the reaction. Most of the solvent was removed by evaporators in the vacuo behind 2 hrs. Then pyridine was dropped to dissolve, continue to reflux 2 hrs. The solution was cooled down to room temperature and filtered; the brown crystals were obtained after a few days. Analytical found for compound 1 (C28H21Cl3N4NiO2): C, 55.10; H, 3.45; N, 9.19%. Calculate: C, 55.08; H, 3.47; N, 9.18%.
The synthesis method for 2 is similar with that of 1. Analytical found for compound 2 (C28H21Cl2NiN5O4): C, 51.99 H, 3.06; N, 10.34%. Calculate: C, 51.93; H, 3.08; N, 10.34%.
Crystal structure determination
Suitable single crystals of compounds 1 and 2 were carefully selected under optical microscope and glued on thin glass fibers. The intensity data of 1 and 2 was collected on Oxford Xcalibur E diffractometer. The empirical absorption corrections were applied to the data using the SADABS system. This structure was solved by direct method and refined by full-matrix least-squares method on F2 using the SHELXS-97 program [10]. All non-hydrogen atoms of 1 and 2 were refined anistropically, and all the hydrogen atoms attached to carbon atoms were fixed at their ideal positions. Pertinent crystal data and structural refinement results for compounds 1 and 2 were summarized in Table 1.
| 1 | 2 | |
|---|---|---|
| Formula | C28H21Cl3N4NiO2 | C28H21Cl2N5NiO4 |
| Mr | 610.55 | 621.11 |
| Temperature/K | 293 (2) | 293 (2) |
| Crystal system | Triclinic | Triclinic |
| Space group | Pī | Pī |
| a/Å | 8.972 (4) | 8.9007 (6) |
| b/Å | 9.010 (4) | 9.0563 (7) |
| c/Å | 17.574 (8) | 17.7195 (13) |
| α/° | 104.410 (5) | 104.701 (1) |
| β/° | 92.753 (5) | 92.252 (1) |
| γ/° | 100.603 (6) | 100.093 (1) |
| V/Å3 | 1345.9 (10) | 1354.91 (17) |
| Z | 2 | 2 |
| Dcalc/g·cm-3 | 1.507 | 1.522 |
| μ(Mo Kα)/mm-1 | 1.052 | 0.958 |
| θ range/° | 2.32 to 25.99 | 2.33 to 26.00 |
| Reflections collected | 14933 | 15037 |
| No. unique data [R(int)] | 5251 [0.0189] | 5284 [0.0174] |
| No. data with I ≥ 2σ(I) | 4442 | 4558 |
| R1 | 0.0288 | 0.0281 |
| ωR2(all data) | 0.0792 | 0.0725 |
Table 1. Crystal data and structure refinement for 1 and 2.
In vitro antitumor activity
Three human glioma cells (U373, U251 and SHG44) were used for cytotoxicity test in vitro using the SRB (sulforhodamine B) assay. They were maintained as stocks in RPMI 1640 (Gibco) supplemented with 10% fetal bovine serum (Gibco). Cell cultures were passaged once or twice weekly by using trypsin-EDTA to detach the cells from their culture flasks. The rapidly growing cells were harvested, counted, and incubated at the appropriate concentration (1-2 × 104 cells/well) in 96-well plates. After incubation for 24 h, the compounds dissolved in culture medium were applied to the culture wells in triplicate and incubated for 48h at 37°C under 5% CO2/95% air atmosphere in a humidified incubator. The culture cells were fixed with 10% cold TCA and stained with 0.4% SRB dissolved in 1% acetic acid. After solubilizing the bound stain with 10 mM of unbuffered Trisma base solution (pH 10.5) using gyratory shaker, the absorbance at 520 nm was measured spectrophotometrically in a microplate reader. Cytotoxic activity was evaluated by measuring the concentration of a compound, which was required to inhibit the protein synthesis by 50% (IC50) and compared with that of aclacinomycin.
Results and Discussion
Molecular structure
The crystal structure determined by single-crystal X-ray diffraction showed that 1 and 2 both crystallizes in the triclinic system, space group Pī. The asymmetric unit comprises one Ni(II) atom, three pyridine molecular, one 3,5- dichlorosalicylaldehyde-2-amino-4-chlorophenol (H2L1) or 3,5-dichlorosalicyde-2-amino-4-nitrophenol (H2L2) Schiff base respectively.
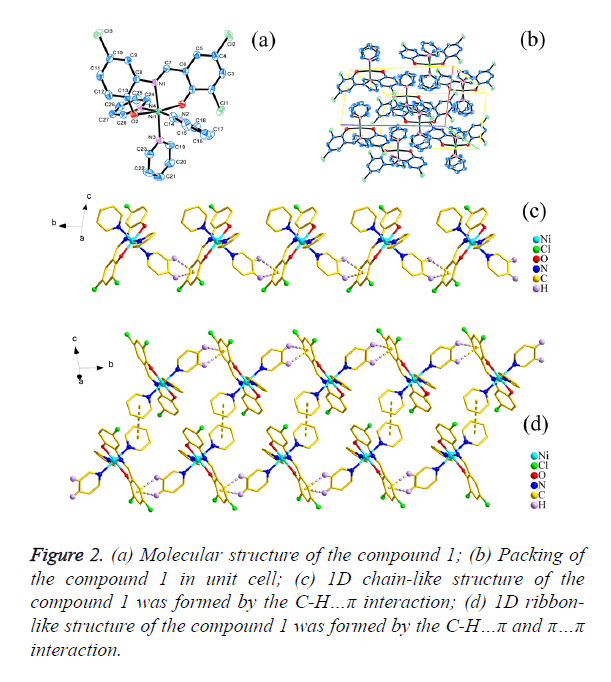
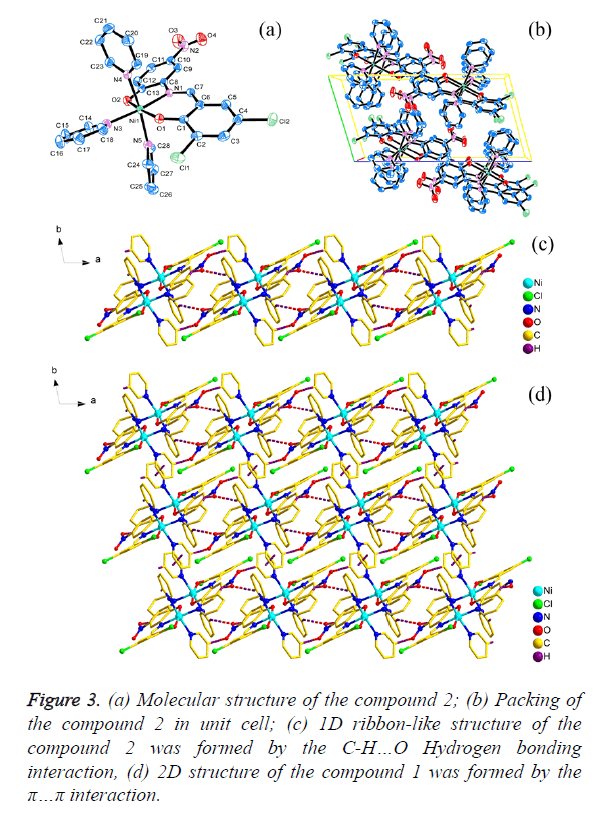
As shown in Figures 2a and 3a, the central Ni1 atom is six-coordinate in a distorted octahedral geometry and it is surrounded by two oxygen atoms (O1 and O2) and one nitrogen atom (N1) from the ligand, three nitrogen atoms (N2, N3 and N4 for 1; N3, N4 and N5 for 2) from three different pyridine. The axis position were occupied by two nitrogen atoms (N1 and N3) from Schiff base ligand (H2L1 and H2L2) and one pyridine molecular, respectively. The angle of N1-Ni1- N3 is 174.950(6)° and 174.598(6)° respectively, which is obviously deviates from linear angle 180°. The four atoms (O1, O2, N2 and N4 for 1; O1, O2, N4 and N5 for 2) were placed in equatorial sites, the sum of the angles [for 1, O1-Ni1- N2=87.30(6)°, <O2-Ni1-N2=89.74(7)°, <O1-Ni1- N4=92.29(6)°, <O2-Ni1-N4=91.71(7)°; for 2, <O1-Ni1- N5=87.27(6)°, <O2-Ni1-N5=89.35(6)°, <O1-Ni1- N4=92.60(6)°, <O2-Ni1-N4= 91.91(6)°] is 361.04° and 361.13 respectively, which shows that the four atoms are basically planar, but the bond lengths [for 1, Ni1-O1=2.0244(14) Å, Ni1- O2=2.0374(15) Å, Ni1-N2=2.1490(17) Å, Ni1-N4=2.1674(18) Å; for 2, Ni1-O1=2.0177(12) Å, Ni1-O2=2.0603(13) Å, Ni1- N4=2.1641(15) Å, Ni1-N5=2.1475(15) Å] are unequal, it indicated that the central Ni atom is six-coordinate in a distorted pentagonal pyramid geometry.
The packing of the compounds 1 and 2 in unit cell was shown in Figures 2b and 3b respectively. Moreover, for 1, the C-H… π(C1i-C6i, i: x, -1+y, z) interactions [d(H15…π)=3.0174(10) Å; d(H16…π)=3.5839(10) Å] were observed between adjacent molecular, which leads to the forming of a interesting 1D chain-like structure (Figure 2c) and the π…π interaction [d(π… π)=4.2701(19) Å] in the adjacent 1D chain-like structure also result in the formation of a 1D ribbon-like structure (Figure 2d); for 2, the C-H…O Hydrogen bonding interactions [H22i… O4=2.6115(19)Å, <C22-H22i…O4=173.421(136)°, i: -1+x, y, z; H15ii…O3=2.6269(19) Å, <C15-H15ii…O3=132.295(175)°, ii: 2-x, 2-y, 1-z; H28iii…O3=2.5657(21) Å, <C28-H28iii… O3=128.854(126)°, iii: 1-x, 2-y, 1-z] were observed between adjacent molecular, which leads to the forming of a interesting 1D ribbon-like structure (Figure 3c), and the π…π interaction [d(π…π)=4.3985(3) Å] on the side of 1D ribbon-like structure also result in the formation of a 2D structure (Figure 3d).
Biological activity
The in vitro cytotoxicities of Schiff ligands L1 and L2, Py and compounds 1 and 2 were evaluated in three human glioma cells (U373, U251 and SHG44) by the SRB (sulforhodamine B) method and the results are summarized in Table 2. Because of the IC50 values of L1, L2 and Py are all higher than those of aclacinomycin, we can conclude that the three compounds showed no antitumor activity. When organic ligands Py, Schiff bases L1 and L2 are in coordination with Ni ion respectively, the resulting compounds 1 and 2 showed significantly increased antitumor activity with the IC50 values of 1.12-4.10 μm, which are lower than those of aclacinomycin.
| Compounds | IC50 | ||
|---|---|---|---|
| U373 | U251 | SHG44 | |
| L1 | 7.88 | 7.21 | 8.1 |
| L2 | 8.12 | 9.33 | 6.46 |
| Py | 9.33 | 7.41 | 7.09 |
| 1 | 1.54 | 2.88 | 3.65 |
| 2 | 1.12 | 3.78 | 4.1 |
| Aclacinomycin | 1.23 | 3.39 | 4.67 |
| aIC50: concentration which produces 50% inhibition of proliferation after 72 h of incubation. | |||
Table 2. In vitro antitumor activity tested compounds and reference drug aclacinomycin.
Conclusion
In conclusion, we successfully obtained two new Ni(II)- coordination compounds (1 and 2) by employing two different flexible Schiff base ligands. From the biological activity investigation, we can see that the antitumor activity of compounds 1 and 2 has been advanced greatly when organic ligands pyridine and Schiff bases are in coordination with Ni ion. However, additional studies are needed to define the mechanism underlying its antitumor activity and evaluate the correlations of its drug efficacy in vivo.
References
- Khosh N, Brown CE, Aboody KS. Contact and Encirclement of Glioma Cells In Vitro Is an Intrinsic Behavior of a Clonal Human Neural Stem Cell Line. PLoS One 2012; 7: e51859.
- Song JH, Song DK, Pyrzynska B. TRAIL Triggers Apoptosis in Human Malignant Glioma Cells Through Extrinsic and Intrinsic Pathways. Brain Pathol 2003; 13: 539-553.
- Zhang Z, Zeng Q, Liu Y. Assessment of the intrinsic radiosensitivity of glioma cells and monitoring of metabolite ratio changes after irradiation by 14.7-T high-resolution 1H MRS. Nmr Biomed 2014; 27: 547-552.
- Annabi B, Thibeault S, Moumdjian R. Hyaluronan cell surface binding is induced by type I collagen and regulated by caveolae in glioma cells. J Biol Chem 2004; 279: 21888-21896.
- Ma P, Liu Y, Zhang LP. Synthesis, BSA Interaction and Antimicrobial Activities of two Amantadine Schiff Bases. Lat Am J Pharm 2015; 34: 999-1005.
- Wang JJ, Zhang WH. A Novel Schiff Base Copper(II) Complex for Inhibiting Growth of Human Hemangioma Cells. Lat Am J Pharm 2016; 35: 1026-1028.
- Zeng Y, Xu T. Anti-Oxidative and Anti-Inflammatory Activity of Baicalin Complexes with Trivalent Lanthanum, Yttrium and Neodymium. Lat Am J Pharm 2016; 35: 1228-1233.
- Zhao CL, Zhang HQ, Jin Y. Crystal structure of bis(2-hydroxy-2-phenylacetato-κ2O,O′)bis(pyridine-κN)nickel(II), C26H24N2NiO6. Z KRIST-NEW CRYST ST 2015; 231: 87-88.
- Yan Y, Qi Y. Crystal structure of poly[(μ2-1,4-bis((1H-imidiazol-1-yl)methyl)benzene-κ2N:N′)-(μ2-4,4′-(1,2-phenylenebis(oxy)dibenzoato-κ4O,O′:O′′,O′′′)nickel(II)], C34H26O6N4Ni. Z KRIST-NEW CRYST ST 2015; 231 (1): 109-111.
- Sheldrick GM. SHELXL-97, Program for Solution Crystal Structure and Refinement, University of Götingen, Germany, 1997.