Review Article - Biomedical Research (2015) Applications of Rapid Prototyping Techniques in Bio-Materials ARTBM2015
Studies on Molecular Dynamics Simulation and Solvent Stability analysis of Collagen Mimetic Peptide GFO with Cyclodextrin ? An Insilico analysis for Tissue Engineering.
Lavanya Gunamalai1*, Chellam Jaynthy21PhD Scholar, Department of Bioinformatics, Faculty of bioengineering, Sathyabama University, Chennai, Tamilnadu, India
2Associate Professor, Department of Bioinformatics, Faculty of bioengineering, Sathyabama University, Chennai, Tamilnadu, India
- *Corresponding Author:
- Lavanya Gunamalai
Department of Bioinformatics
Faculty of Bioengineering
Sathyabama University
Chennai, Tamilnadu
India
Accepted date: July 07 2015
Abstract
Collagen mimetic peptides (CMPs) and collagen-like proteins (CLPs) that mimic either structural or functional characteristics of natural collagens have many applications to engineering collagen-like materials for potential biomedical use. Molecular modeling studies on collagen and collagen-like peptides suggest that triple-helical stability can vary along the amino acid chain. GFO is collagen-like peptide constructed using building script (BuScr) triple helix builder, whose stability is calculated in same scripting software and the Tm is found to be 30.74 OC.The stabilized GFO peptide is docked with α- cyclodextrin using auto dock 4.01v freeware and found atom-atom interaction with amino acids like Phe 11, Gly7 respectively. The estimated free energy of binding is calculated using auto dock was found to be as -8.26 kcal/Mol with Inhibition Constant, Ki = 880.75 nM at 298.15K. The interaction and negative binding shows there is stability between the docked complex, this study is conducted experimentally to confirm the same process in a wet lab.To the docked complex, solvent is added in two ways either for the whole complex else to specific docked region using specific options protocol available in Accelrys’ Discovery Studio 2.1v.Cascade dynamics studies are carried out for two different solvated complex with ensembles NVT and NPT, the changes in energy are observed only in the production stage for both complexes. Finally, the trajectory path is generated using principle component analysis(PCA) for the complex shows a small unit of changes in total energy at 302.526K.This computational study is correlated with experimental study shows, the stability of the Peptide complex in solvent environment.
Keywords
Collagen mimetic peptides (CMPs), CLPs, Biomedical application, GFO peptide, cyclodextrin, The BuScr , Cascade dynamics, principle component analysis.
Introduction
Collagen is more important protein that found in the extracellular matrix form in the body, it is the major insoluble fibrous protein in connective tissue [1].It helps to maintain the structural integrity, suppleness and firmness in tissues, also maintains the constant renewal of skin cells [2]. The elasticity of the skin is mainly due to the presence of collagen, due to aging of cell production of collagen in the cell becomes slow and other cell structures will lose its uniqueness [3].The decrease in production of collagen leads to skin thinner, sags, wrinkles, sometimes the skin will damage easily that in turn may lead to hair on the skin gets lieless [4], collagen plays a very important role in maintaining the skin and body of mammals [3,4]. Chemically the collagen is made up of amino acids, that too specifically Glycine, Proline, Hydroxyproline, and Arginine. Structurally, it is, triple helical domain in twisted peptide, under microscope collagen is observed like elongated fibrils. It is mostly found in tissues such as tendon, skin, bone, blood vessels, and the digestive tract and abundant in the cornea [5]. Collagen like peptide is also same a collagen structurally and functionally and it finds its more applications in the biomedical field. Computer simulation study of collagen-like peptide GFO is necessary before carrying out the experimental analysis of this GFO collagen-like peptide. Simulation can provide the detail information about the motion of particles as a function of time; many studies on simulation of biomolecules using different kind’s software are carried out in many Bioinformatics research organizations [6]. CHARMM based simulated of biomolecules is more common than other simulating methods, the computational methods of study the molecular dynamics methods provide useful information on biomolecules and its physical motions, a small change in conformation of protein with respective to time plays an important role in the function of their own [7]. In our research, we studied the various energy parameters for GFO peptide, cyclodextrin with water and without water; perhaps simulation is carried with complex for peptide, cyclodextrin with water. The objective of this research of collagen-like peptide begin been computed designing peptides using the appropriate software its energy values must be investigated at each step. After docking peptide with α- cyclodextrin it was introduced into the water environment to find the stability of the complex. The simulations were carried in three different steps such as dynamics heating/cooling, equilibrate and production using different ensemble for a complex of GFO collagen- like peptide, α- cyclodextrin and water.
Experimental detailsExperimental details
Collagen builder
The triple helix collagen builder script or THe BuScr is used to build the GFO peptide that mimics as like collagen. The three chains A, B and C in. fasta format is uploaded individually to perform this task. GFO collagen like peptide was built as per instructions given in the user defined manual of BuScr and its time and stability profiles is calculated in BuScr collagen builder. Tcl/Tk scripting language is an inbuilt program in THe BuScr and also provide a user friendly platform to perform the scripting task [8,9].
Peptide-cyclodextrin interaction
The structure, based drug designing concept is applied to perform docking here the two molecules, among peptide will act as receptor and cyclodextrin act as a ligand. It follows the old concept of lock and key model, only the appropriate key can fit into the lock. Similarly, only the molecules that as binding site or shape complementary can fit into it. This task was performed using Auto- Dock 4, it is freely available under the GNU General Public License. In this current study, Docking studies were carried out using the program Auto dock 4, GFO collagen-like peptide with α -Cyclodextrins.
Analysis of docking Interaction calculation
The energy parameters of the docked complex, GFO collagen like peptide with α –Cyclodextrins Interaction energy is carried out between the complexes to confirm interaction is strong between the docked complex mole6 cules. On the other hand, for the sample complex overall initial energy for stability is calculated.
Solvation of complex
All the biomolecules are functioning in an aqueous en10 vironment, some molecules may lose its stability when interaction with water, but in other cases increases its stability in aqueous environment. In this current study, water molecules are added to the docked complex using the periodic boundary conditions with optioning of or15 thorhombic cell shape. In addition, it prepares special constraints to allow simulation of these waters in the explicit periodic boundary [10].
Cascade molecular dynamics
The output of the solvation was employed as input for the cascade molecular dynamics simulation, in this process initially energy minimization is performed on the complex structure prior to dynamics study to relax the conformation and remove steric overlap that pro25 duces bad contacts. Most commonly Steepest Descent, Conjugate Gradient (CONJ), the methods available in CHARMm was used [11]. The objective of the molecu28 lar dynamics is to generate a realistic model of a struc29 ture's motion, perform conformational searching, pro30 duce a time series analysis of structural and energetic properties, explore the energy decay, and analyze sol32 vent effects. The method used in CHARMm to constrain these motions is the SHAKE algorithm [12]. SHAKE is available in the Equilibration, Heating, Minimization, and Production protocols, as well as the Standard Dy36 namics Cascade are most commonly used to perform this task in Accelrys discovery studio 2.01v.
Results and DiscussionResults and Discussion
Triple helix and its stability
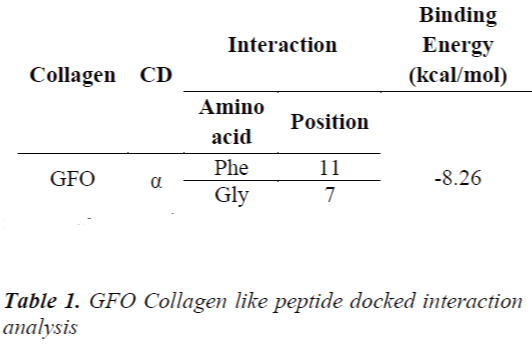
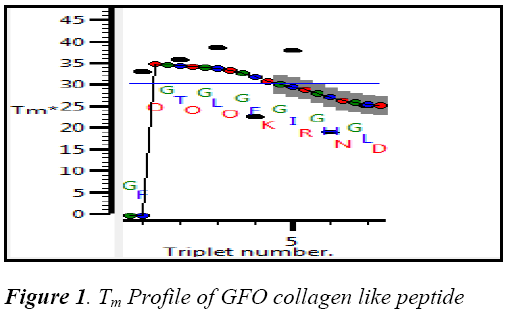
The collagen type 1 sequence starts from 260‐site‐α2 (G254‐‐ D277):GFOGTOGLOGFKGIRGHN GLD was built in THeBuScr triple helix builder and its stability Tm was found by 30.740C .The stability profile of GFO collagen like peptide is shown in the Fig 1.
GFO Peptide -cyclodextrin interaction
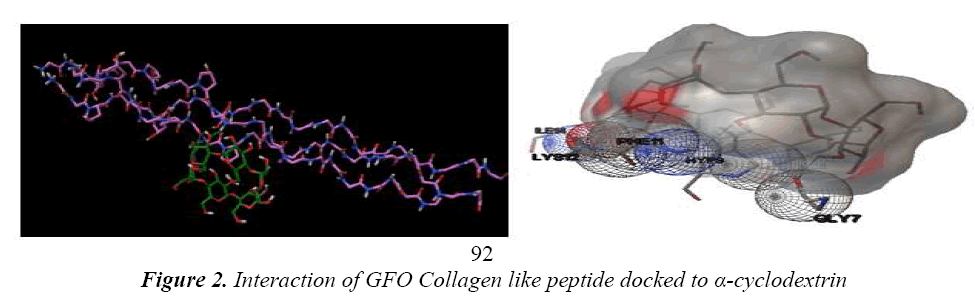
Leikina et al., And other collagen researchers stated that it is less stable, so complex should be formed to make it structurally and functionally stable [13]. The concept of same is applicable for GFO Collagen like peptide, GFO triple helix built collagen like peptide GFO is docked with α- cyclodextrin to confirm the atom-atom contact in between them. For this peptide-chemical complex, we docked the two molecules using AUTODOCK 4.01 with Auto grid and auto dock calculations. In auto grid, Grids were laid over a three-dimensional cubic box where the GFO Collagen like peptide occupied the center of the box. On the other hand, α- cyclodextrin root maker, torsions, and aromatic criteria were initialized and prepared further fed for the docking. Docking is processed with Genetic algorithm; this algorithm is commonly used method for receptor –ligand interaction. During docking, it was about nearly 2500000 number of energy evaluations with 150 individual population sizes and 27000 generations is carried out to get the result of one run, the same process is carried for next nine runs. Among them, least estimated binding energy run is chosen to analyze the interaction and it was considered as the best run in docking and interaction. Estimated binding Energy is the summation of the intermolecular Energy, the torsional energy and the Internal energy. The interaction between them confirms the peptide is stability. Hence, it confirms the experimental results of collagen stabilized by adding the cyclodextrin. Fig 2 shows GFO Collagen like peptide-docked interaction with α – Cyclodextrin and its interaction with aminoacid are stated in Table 1.
Initial energy estimation of complex
The docked GFO collagen like peptide with α- cyclodextrin form a single complex structure, interaction energy is calculated in various energy parameters. Such as, potential energy is 713.87221 kcal/mol, Van der Waals energy is -37.54061 kcal/mol, Electrostatic energy is -67 6.33160 kcal/mol and their respective Interaction Energy is -713.873 kcal/mol with VDW Interaction Energy of -37.5406 kcal/mol and Electrostatic Interaction energy of -676.332 kcal/mol. Now the same complex is proceeded to estimate its initial energy and it was found, its potential energy, Van der Waals energy, Electrostatic energy and RMSD gradient as 2132.47175kcal/mol, 1096.24050 kcal/mol, - 1685.66742 kcal/mol and 175.07678 (kcal/(mol x Ang Interaction energy of -676.332 kcal/mol. Now the same complex is proceeded to estimate its initial energy and it was found, its potential energy, Van der Waals energy, Electrostatic energy and RMSD gradient as 2132.47175kcal/mol, 1096.24050 kcal/mol, - 1685.66742 kcal/mol and 175.07678 (kcal/(mol x Angstrom)) respectively.
Solvation of complex-GFO collagen like peptide and α-cyclodextrin
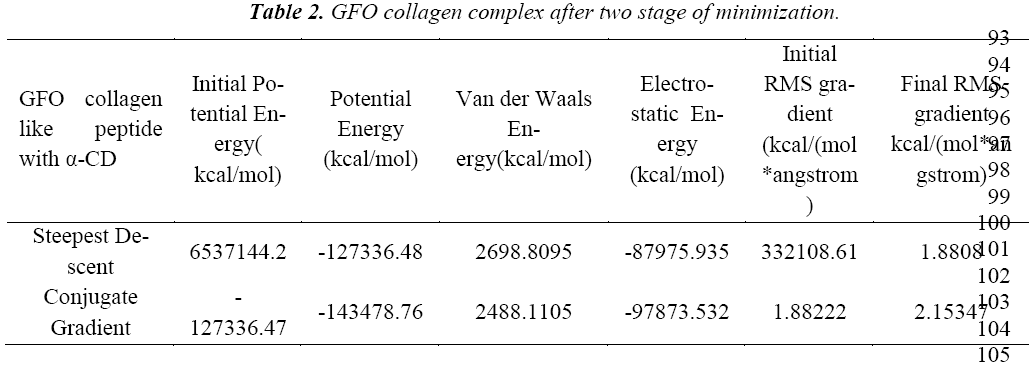
Water molecules are added to the complex structure of GFO collagen like peptide and α-cyclodextrin in the cell shape of orthorhombic arrangement with Explicit Peri62 odic Boundary solvation model. The radius of 20Å around the complex with 7Å minimum distance bound64 ary and random speed of 314159 was kept, as default to process this study. The aim of this solvation study is to find the stability of the complex after adding the water molecule to the system. We further calculated the all the energy parameters and found the positive energy value, which implies the less stable after adding the water molecules. Henceforth, before studying the dynamics of the system, we tried to minimize the complex structure using the steepest descent and conjugate gradient. An initial minimization stage, typically using the robust steepest descent (SD) algorithm to resolve any initial poor contacts within the system without creating large distortions in the overall structure. Steepest descent (SD) algorithm with Minimization Max Steps of 500 and the Minimization RMS Gradient of 0.1, now second minimization is applied to complex with a Conjugate Gradient algorithm concept with Minimization Max Steps of 500 and the Minimization RMS Gradient from 0.0001 respectively(Table 2).
Cascade Simulation of complex with water molecules
Then dynamics simulation studies have performed in a docked complex with water by adjusting the tempera87 ture of a molecule from 277K-313 K using the program called dynamics by heating under simulation package in Discovery Studio. Then molecular dynamics simulation heating stage was employed to add thermal energy to the system to reach a target temperature.
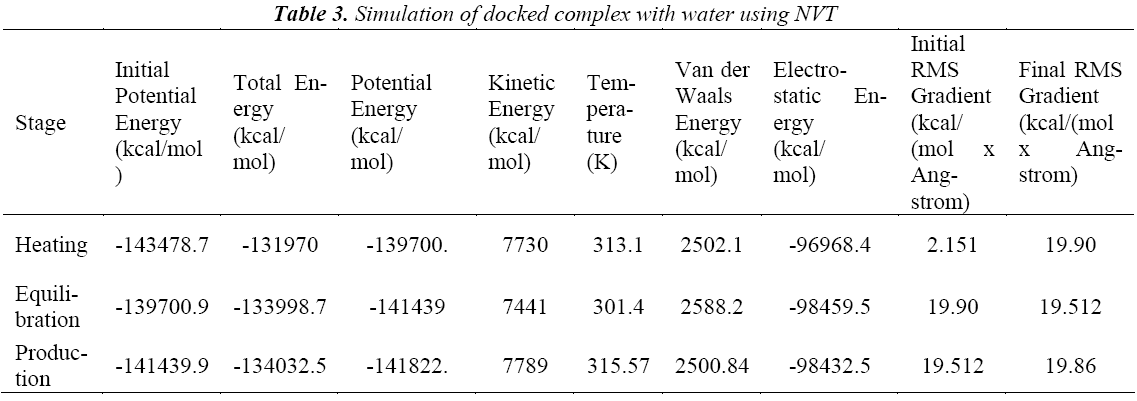
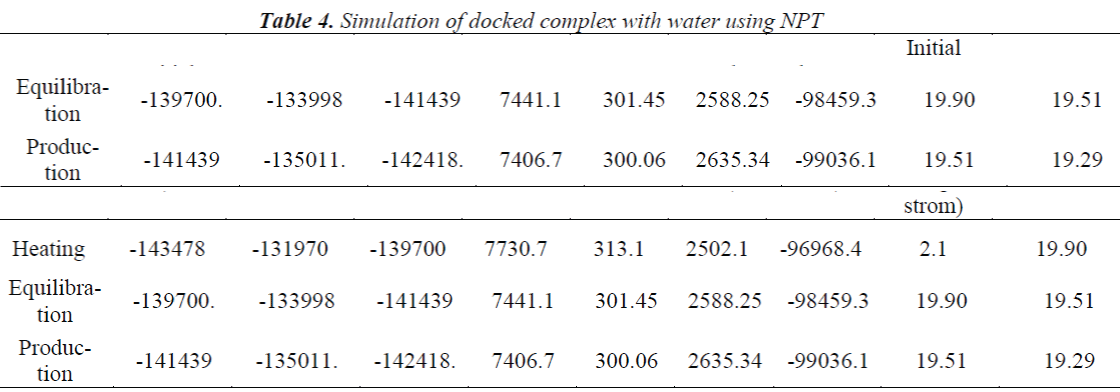
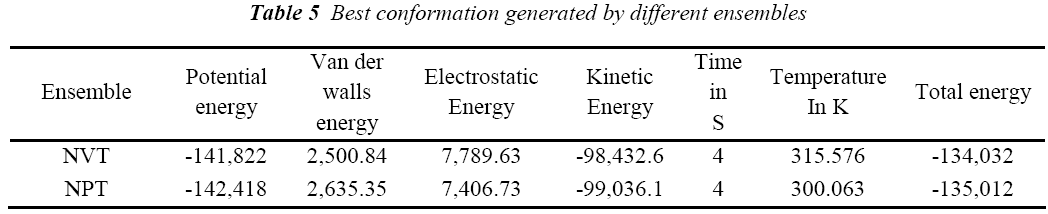
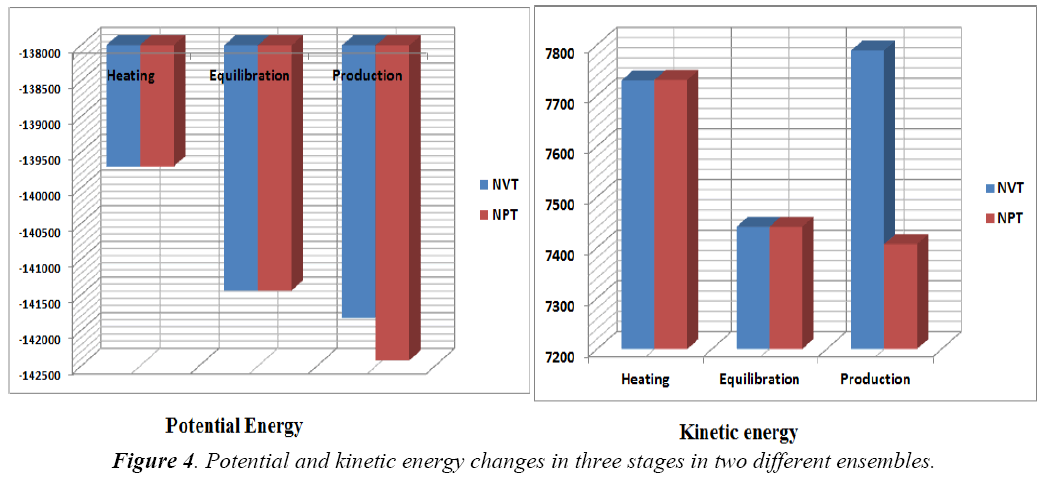
A standard molecular dynamics simulation stage is then employed to equilibrate the system at a target temperature. The purpose of the equilibration stage is to ensure that the energy in the system was distributed appropriately among all degrees of freedom. This allows the system to achieve thermal equilibration at the target temperature. Finally, production of dynamics is carried out to calculate the final energy parameters of the complex GFO collagen like peptide with cyclodextrin (CD) and water. The two different types of ensembles NVT (Table 3) and NPT (Table 4) was studied, for docked complex with water and observed the variations in dynamics heating, equilibration and production. Comparison Kinetic and potential energy of the two ensembles is shown in the Fig 4. Finally, the production in dynamics results different conformation of the docked complex with various energy parameters with respect to time and temperature. Table 5 shows best conformations generated using NVT and NPT with energy parameters. The best conformation is fed as input for generating the motion of the GFO collagen like peptide with cyclodextrin (CD) and water complex, the process was carried out individually for both ensembles and found a very small change in energy parameters and pressure. The fundamental motions of a trajectory path of complex are created by means of Principal Components using charm, Principal Components Analysis (PCA) is similar to the quasi-harmonic analysis derived from vibrational models [14].The same process is carried out for the solvation of GFO collagen like peptide with cyclodextrin (CD) complex with harmonic restraint. In this study the water molecules are added only in the specific docked region instead of the whole complex as shown in figure 3. Fig 5 shows the water molecules added to specific docked region, even in this complex the cascade dynamics and trajectory motion study is carried and the results show very small unit change in energy, temperature and pressure.
Conclusion
Hence, from this study, we conclude that docked complex of GFO collagen like peptide with cyclodextrin (CD) with solvation in the specific docked region or in whole complex shows small changes in energy parameters. Extended cascade molecular dynamics studies using two different ensembles such as NVT and NPT show variation in final production stages in both complex of GFO collagen like peptide with cyclodextrin (CD) with solvation in the specific docked region and GFO collagen like peptide with cyclodextrin (CD) with solvation in the whole complex molecule. Finally the trajectory path is generated using principle component analysis for the complex shows a small unit of changes in total energy at 302.526K.This computational study is correlated with experimental study shows, the stability of the peptide complex in solvent environment.
Acknowledgment
We would like to thank Sathyabama University for providing Bioinformatics lab facility to carry out the research work.
References
- Di Lullo GA, Sweeney SM, Korkko J, Ala-Kokko L, San Antonio JD. “Mapping the ligand-binding sites and disease-associated mutations on the most abundant protein in the human, type I collagen.” J.Biol.Chem.2002; 277(6):4223–4231.
- Smith JG, Davidson EA, Clark WM, “Alterations in human dermal connective tissue with age and chronic sun damage” J Invest Dermatol 1962; 39: 347–356.
- Fisher GJ, Wang Z-Q, Datta SC, Varani J, Kang S, Voorhees JJ, “Pathophysiology of premature skin aging induced by ultraviolet light” New Eng J Med1997; 337: 1419–1428.
- Pieraggi MT, Julian M, Bouissou H, “Fibroblast changes in cutaneous aging” Virchows Arch A Pathol Anat Histopathol1984; 402: 275–287.
- Viidik A,Ekholm R, “Light and electron microscopic studies of collagen fibers under strain” A. Anat. Entwicklung 1968; 127(2): 154-164
- Bodian DL, Radmer RJ, Holbert S, Klein TE, “Molecular dynamics simulations of the full triple helical region of collagen type I provide an atomic scale view of the protein's regional heterogeneity. Pac” Symp Biocomput.2011; 193- 204.
- Mirijanian DT,Mannige RV, Zuckermann RN, Whitelam S., “Development and use of an atomistic CHARMM-based forcefield for peptoid simulation.” J Comput Chem 2014; 35(5): 360-370
- Ousterhout JK. Tcl and the Tk Toolkit. Addison- Wesley, Reading 1994
- Persikov AV, Ramshaw JA., Kirkpatrick A., Brodsky B. “Amino acid propensities for the collagen triple-helix” Biochemistry 2000; 39: 14960- 14967
- Allen MP. and Tildesley DJ, “Periodic Boundary Conditions and Potential Truncation. In: Computer Simulations of Liquids”1987; Oxford University Press, Oxford.
- Fletcher R and Reeves CM, “Function minimization by conjugate gradients.Computer” Journal, 1964, 7: 148–154,.
- Straatsma TP, Berendsen HJC, and Stam AJ, Estimation of statistical errors in molecular simulation calculations. Molecular Physics,1986, 57: 89, .
- Leikina E,Mertts MV,Kuznetsova N, and Leikin S,Type I collagen is thermally unstable at body temperature,Proc Natl Acad Sci USA, 2002, 99(3): 1314–1318,.
- Balsera MA, Wriggers W, Oono Y, Schulten K, Principal component analysis and long time protein dynamics. The Journal of Physical Chemistry, 1996, 100(7), 2567-2572, .