Short Communication - Biomedical Research (2016) Health Science and Bio Convergence Technology: Edition-I
Preliminary report on the incidence of pathogenic Entamoeba histolytica in patients attending out-patient clinic of university of Nigeria teaching hospital, Ituku Ozalla, Enugu-Nigeria
Ameh LO1, Ikekpeazu JE1, Eze AA1, Udeogaranya PO2, Okoli UA1, Ogbunude POJ1*1Department of Medical Biochemistry, Faculty of Basic Medical Sciences, University of Nigeria, Enugu-Campus, Nigeria
2Department of Pharmacy, University of Nigeria, Nsukka, Nigeria
- *Corresponding Author:
- Ogbunude POJ
Department of Medical Biochemistry
Faculty of Basic Medical Sciences
University of Nigeria
Nigeria
Accepted date: April 04, 2016
Abstract
Entamoeba comprises six species that inhabit human intestinal lumen, of which only Entamoeba histolytica (E. histolytica) is positively shown to be pathogenic. It is the causative agent of human intestinal and extra intestinal ameobiasis and responsible for significant morbidity and mortality in many developing countries as well as several communities in developed countries. Entamoeba histolytica is in the WHO list of neglected tropical diseases that requires funding for drug discovery, epidemiology, surveillance and vector control. Indeed, WHO recommends treatment of pathogenic Entamoeba histolytica infection while treatment of non-pathogenic species is unnecessary; hence, the need to use technique that positively identifies the pathogenic specie. Stool samples from patients attending University of Nigeria Teaching Hospital Ituku-Ozalla, for gastrointestinal complaints or diarrhoea were analysed by combined use of polymerase chain reaction and restriction endonuclease digest to positively show the presence of pathogenic Entamoeba histolytica in stool. Thirty samples extracted with commercial genomic DNA extraction kit (DNA extraction buffer, Life River, UK) according to manufacturer’s protocol were subjected to amplification by polymerase chain reaction using primers (Forward: 5′TAAAGCACCAGCATATTGTC3′ and Reverse: 5′GATGACATATCCTCTTCTTG3′). Two of the genomic DNAs were amplified but only one of them contained EcoR1 digestion site suggesting that it is pathogenic Entamoeba histolytica. This represents approximately 3.3% of the total samples studied and is considered significant considering the population at risk. Furthermore the study represents improvement in diagnosis of Entamoeba histolytica using molecular based technique in a local laboratory.
Keywords
Pathogenic Entamoeba histolytica, Non-pathogenic Entamoeba histolytica, Polymerase chain reaction, Endonuclease enzyme.
Introduction
Entamoeba histolytica (E. histolytica) is a protozoan parasite that is primarily dominant in developing countries and causes amoebic diarrhoea, colitis and amoebic liver abscesses. Although about eighty per cent of individuals infected with the parasite remain asymptomatic, the remaining twenty per cent develop clinically overt disease resulting in 40,000 to 1, 00,000 deaths annually [1]. Clinical diagnosis classically depends on the visualization of parasites by light microscopy of wet smear or stained specimens. This method, however, does not distinguish between the trophozoites of disease causing Entamoeba histolytica and the non-pathogenic E. dispar [2-5]. Reliable distinction would have a medical impact as infections due to pathogenic specie would be treated and patients with non-pathogenic types spared the ordeal of undergoing unnecessary treatment plus the savings on the costs of drugs to the health system.
In the present study, we utilized the primers described by Tachibana et al. [6]. The primers (Forward: 5′TAAAGCACCAGCATATTGTC3′ and Reverse: 5′GATGACATATCCTCTTCTTG3′) were designed to target the genomic DNA coding 30,000 molecular weight antigen. The antigen is coded by a 714 base pair gene with an open reading frame encoding 218 amino acids common to both pathogenic and non-pathogenic Entamoeba histolytica. Three restriction endonucleases including Hinc II, EcoRI and TaqI, however, digested DNAs from pathogenic isolates showing different zymodeme patterns but did not digest DNA from nonpathogenic species.
Materials and Methods
Fresh stool samples were collected from patients who presented with gastrointestinal complaints or diarrhoea at Microbiology Laboratory Department of University of Nigeria Teaching Hospital (UNTH). The samples were extracted with commercial genomic DNA extraction kit (DNA extraction buffer, Life River, UK) according to the manufacture’s instruction. Briefly 0.1 g of stool sample was taken into 2 ml test tube to which 1.0 ml of sterile normal saline was added. The mixture was vigorously vortexed and then centrifuged at 600 g for 2 minutes. To 0.9 ml of supernatant was added 0.1 ml of the DNA extraction buffer, mixed and allow to stand for 5 minutes before centrifugation at 600 g for 30 minutes; the supernatant resulting from the centrifugation was incubated at 100ºC for 15 seconds to inactivate any DNA denaturing enzyme. Absorbance of the DNA extracts was read at 260 nm with Eppendorf Bio photometer and the concentration of DNA calculated using the formula C=260 χ 20 where C=concentration of DNA in mg/ml.
PCR amplification
The PCR amplification of the genomic DNA was carried out with the primers described above as described by Saiki et al., with modification [7]. Briefly, the amplification was performed in 100 μl reaction containing 10 mM Tris-HCl, pH 8- pH 3; 1.5 mM MgCl2; 50 μM KCl; 200 μM each, of dNTPs; 40 ng of each of the primers and 1 unit of Taq DNA polymerase. The tube containing the reactants were placed in the thermo cycler programmed for 45 cycles: a single cycle at 94ºC for 90 seconds was followed by 45 cycles at 60ºC for 15 seconds and 72ºC for 30 seconds with a final cycle at 72ºC for 60 seconds.
Restriction endonuclease digest
Restriction endonuclease digest of the amplified DNA was performed as described by the manufacturer. The reactants were added in the following order; 17 μl nuclease-free water, 2 μl of 10X buffer, 10 μl amplified DNA product and 10 μl of EcoRI enzyme. This was gently mixed and incubated at 37ºC for 20 minutes. Aliquots (10 μl) were electrophoresed in a 1% agarose gel containing 0.1 μg of ethidium bromide per ml and bands visualized by UV trans illumination.
Results and Discussion
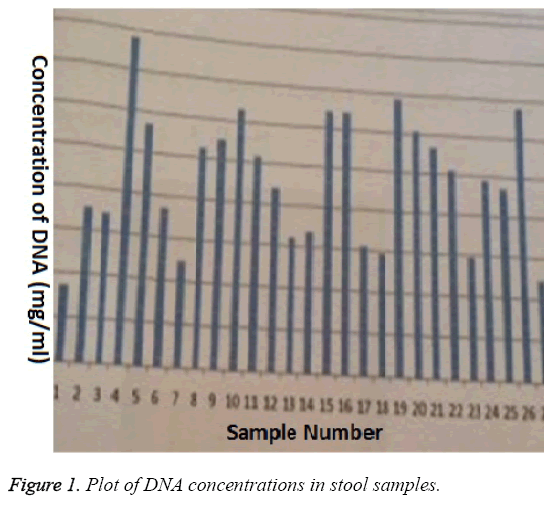
The commercial extraction kit (Life River UK) efficiently extracted DNA from stool samples (Figure 1) however; the extracts did not necessarily represent DNA from amoeba infection but could be from any of many sources present in the stool. Using primers specific for pathogenic and nonpathogenic amoeba, two of the thirty DNA samples were amplified, thus, approximately 6.6 % of patients who presented with gastrointestinal infection were due to amoeba. Since the study did not investigate the effect of the commercial extraction kit on the efficiency of the PCR amplification, we cannot rule out the possibility that we may have higher infection rate due to amoeba than seen in the study. We do know that extraneous conditions affect PCR amplification, for example, sensitivity of PCR decreased within two days in faeces stored in sodium-acetic acid-formalin fixative [8].
To determine the nature of the two amplified DNA samples, the amplicons were subjected to restriction endonuclease digest using EcoR1. The amplicon from the pathogenic amoeba contains the enzyme recognition site while that from the nonpathogenic amoeba is devoid of the restriction site. Our result shows that one of the two amplicons was digested by the restriction enzyme, showing that one individual infection was definitely due to pathogenic Entamoeba histolytica.
It is important to emphasize that the study does not claim that the 30,000 molecular weight antigen is responsible for the pathogenicity of the pathogenic strain but rather that its digestion with restriction endonuclease could be used to differentiate strains. Equally, amplification of this gene with other primers and subsequent digestion with AccI, TaqI and XmnI was also useful in distinguishing Entamoeba histolytica isolates [9]. Other studies had shown that other genes, like 125- kDa antigen of Entamoeba histolytica could also be useful in distinguishing pathogenic and non-pathogenic isolates [10]. The study showed that, although, only one of the 30 stool samples contained pathogenic Entamoeba histolytica, this is considered significant when population at risk is taken into account. Nigeria has a population of over 160 million people of which over 80% are living in rural areas where running water is almost nonexistence and standard of hygiene is very poor so the inhabitants are likely to be exposed. The study is a first step in comprehensive epidemiological study that will look into the prevalence of pathogenic and non-pathogenic Entamoeba histolytica in our geographical region. No doubt, there are limitations to this kind of study using highly sensitive technique in poor resource laboratory. The most significant limitation is general problem of inefficient power system in the country. In a limited study as this, the problem can be handled but poses serious handicap if extensive epidemiology is carried out with power outage for several days.
References
- Petri WA Jr, Singh U. Diagnosis and management of amoebiasis. Clinical Infectious Disease 2009; 29:1117-1125.
- Hamzah Z, Petmitr S, Mungthin M, Leelayoova S, Chavalitshewinkoon-Petmitr P. Differential detection of Entamoeba histolytica, Entamoeba dispar and Entamoeba moshkovskii by a single round PCR assay. J Clin Microbiol 2006; 44: 3196-3200.
- Fotedar R, Stark D, Beebe N, Marriot D, Ellis J, Harkness J. PCR detection of Entamoeba histolytica, Entamoeba dispar, and Entamoeba moshkovskii in stool samples from\ Sydney, Australia. J Clin Microbiol 2007; 45: 1035-1037.
- Lebbad M, Svard SG. PCR differentiation of Entamoeba histolytica and Entamoeba dispar from patients with amoeba infection initially diagnosed by microscopy. Scand J Infect Dis 2005; 37: 680 685.
- Nunez YO, Fernandez MA, Torres-Nunez D, Huva JA, Montano I, Maestre JL, Fonte L. Multiplex polymerase chain reaction amplification and differentiation of Entamoeba histolytica and Entamoeba dispar. Am J Trop Med Hyg 2001; 64: 293-297.
- Tachibana H, Ihara S, Kobayashi S, Kaneda Y, Takeuchi T, Watanabe Y. Differences in genomic DNA sequences between pathogenic and non-pathogenic isolates of Entamoeba histolytica identified by polymerase chain reaction. J Clin Microbiol 1991; 29: 2234-2239.
- Saiki R, Gelfand DH, Stoffel S, Scharf SJ, Higuchi R, Horn GT, Mullis KB, Erlich HA. Primer\ directed enzyme amplification of DNA with a thermostable DNA polymerase. Science 1988; 239: 487-491.
- Troll H, Marti H, Weiss N. Simple differential detection of Entamoeba histolytica and Entamoeba dispar in fresh stool specimens by sodium acetate acid-formalin concentration and PCR. J Clin Microbiol 1997; 25: 1701-1705.
- Tannich E, Burchard GD. Differentiation of pathogenic from non-pathogenic Entamoeba histolytica by restriction fragment analysis of a single gene amplified in vitro. J Clin Microbiol 1991; 29: 250 255.
- Edman U, Meraz UMA, Rausser S, Agabian N, Meza I. Characterization of an immune-dominant variable surface antigen from pathogenic and non-pathogenic Entamoeba histolytica. J Exp Med 1990; 172: 879-888.