Research Article - Journal of Clinical Research and Pharmacy (2018) Volume 1, Issue 1
Phenotypic identification and antibiogram profile of citrobacter species.
Oludare Temitope Osuntokun1*, Taiye Anangwureyi Jemilaiye1, Adisa O Thonda21Department of Microbiology, Adekunle Ajasin University, Akungba Akoko
2Department of Biological Science, Kings University, Ode Omu, Nigeria
- *Corresponding Author:
- Oludare Temitope Osuntokun
Department of Microbiology
Adekunle Ajasin University
Akungba Akoko
Nigeria
Tel: 08063813635; 08056096672
E-mail: osuntokun4m@gmail.com
Accepted date: December 30, 2017
Citation: Osuntokun OT, Jemilaiye TA, Thonda AO. Phenotypic identification and antibiogram profile of Citrobacter species. J Clin Res Pharm. 2018;1(1):1-6
Abstract
The administration of antimicrobial agents on patients had a major impact on the rate of survival from infections. However, the evolving and the changing patterns of antimicrobial resistance posed a threat and caused a demand for review of the existing antibiotic policy or a demand for new antibacterial agents. This study was designed to determine the phenotypic identification in Citrobacter species and Antibiogram profile of citrobacter species isolated from University College Hospital (UCH), Ibadan. Twenty clinical isolates of clinical specimens were recovered from patients using the laboratory services of UCH between December 2016 and March 2017. Preliminary confirmatory test to identify the isolates as Citrobacter was then performed using Oxidase strips and Microbact™ 12A and 24E (Oxoid, UK) Identification Kits. Antibiotic susceptibility tests (Antibiogram profile) was carried out on the identified Citrobacter isolates using some common antibiotics and Minimal Inhibitory Concentrations (MICs) of Piperacillin/ Tazobactam tested against the Citrobacter spp were determined. Ofloxacin was active against all the four Citrobacter isolates (100%), with diameter zone of inhibition ranging from 18 mm to 25 mm. Closely followed is Piperacillin/Tazobactam which was active against three out of the four isolates (75%) with diameter zone of inhibition ranging from 26 mm to 40 mm. Two of the isolates (50%) showed susceptibility to Co-trimoxazole (24 mm and 30 mm), and Imipenem (18 mm and 35 mm). All the isolates were, however, resistant to Cephalosporins (Ceftazidime, Cefotaxime and Cefoxitin), Cloxacillin, Oxacillin, Piperacillin, Meropenem, Nystatin and combination of Ampicillin+Clavulanic acid. MIC values were quite high ranging from 100 μg/mL to 110 μg/mL. This study highlighted the prevalence of Citrobacter species among clinical samples in southern Nigeria and multi-drug resistant nature of the isolated Citrobacter species.
Keywords
Phenotypic identification, Antibiogram profile, Citrobacter species
Introduction
The administration of antimicrobial agents on patients had a major impact on the rate of survival from infections. However, the evolving and the changing patterns of antimicrobial resistance posed a threat and caused a demand for review of the existing antibiotic policy or a demand for new antibacterial agents. Bacteria antimicrobial resistance is a worldwide problem that is exacerbated by the diminishing number of new antimicrobial drugs in the pharmaceutical pipeline [1]. This is an emerging public health problem, especially in hospitals of the developing and the newly industrialized countries of Africa, Asia and the Pacific [2,3].
In developing countries such as Nigeria, the spread of antimicrobial resistance is a matter of concern as it compromises the management of infectious diseases, the leading causes of morbidity and death in these countries [4,5]. Much of the current discourse on infectious disease and drug resistance as it affects Sub-Saharan Africa is limited to the pressing problems associated with emerging and re-emerging resistant organisms. Hence, the available therapeutic options for antibiotics-resistant organisms are severely limited, as these organisms frequently display a multidrug-resistant phenotype.
The excessive use of antimicrobials produces selective pressure for resistance. Also, empirical treatment with ineffective antibiotics prescribed by physicians and poor patient adherence to antibiotic regimens could potentially lead to mutation and drug resistance [6]. Furthermore, the routine use of antibiotics in animal foods to promote their growth is another aspect that may enhance selective pressure for resistance [7,8]. The rate of emergence of antibiotic resistance in the isolates of Enterobacteriaceae and in other Gram-negative bacilli in many parts of the world is very alarming and this is a major threat to the successful treatment of infections in hospitals [9]. Citrobacter species are aerobic, motile, Gram negative bacilli of Enterobacteriaceae. Citrobacter species have been reported to cause a range of infections which include urinary tract infections, wound infections (surgical site infections), respiratory infections, middle ear infections, infections of meninges, osteomyelitis, peritonitis, endocarditis, and bacteremia [10].
The leading three species causing human infections are C. freundii, C. koseri, and C. braakii. Those predispose to Citrobacter infections are the neonates, who may develop sepsis and meningitis (usually less than 2 months of age), and the debilitated aged (≥ 65 years) or immunocompromised patients. Many Citrobacter infections are nosocomially acquired, however, they can also be community acquired. In the hospital settings, Citrbacter spp. might account for 3-6% of all Enterobactericeae causing nosocomial infection. The significance of nosocomial infection lies not only in its ability to substantially alter morbidity and mortality statistics, but also in its economic implications. Nosocomial infections pose a problem of enormous magnitude globally as hospital localities have proven favourable in transmitting Citrobacter infection due to existing suitable pathogen-host-environment relationship [5].
The Citrobacter species are emerging as important nosocomial pathogens with antibiotic-resistant strains, though less frequent [11]. Patient with invasive devices are more prone for invasive Citrobacter infections and are associated with a high mortality rate with majority of patients perishing or wilting to Citrobacter bacteremia. The high mortality rate associated with Citrobacter infections may be due in part to ineffective empirical antibiotic therapy.
Antibiotic resistance has increased significantly among species of Citrobacter, making infections caused by these pathogens are difficult to treat [12]. Citrobacter species are often resistant to the routinely used antibiotics, like the Penicillins, Cephalosporins, Aminoglycosides and Tetracyclines. Cephalosporins form the major component of empirical antibiotic treatment especially in teaching hospitals. Indiscriminate use of cephalosporins, the 3rd generation cephalosporins, has contributed to the evolution of extended spectrum beta lactamases (ESBLs) production and Multiple Drug Resistance (MDR) [13]. MDR strains are becoming frequent in Citrobacter spp, where clinical isolates resistant to a wide range of antibiotics are increasingly being reported [14,15].
Materials and Methods
Equipment and materials used
The equipment used for this study include: Autoclave (LDZX-30FBS, England), Incubator (Genlab, UK), Oven (Gulfex Medical and Scientific, England), Refrigerator (Haier Thermocool, China), Weighing balance, Hot plate, Bunsen burner, and Laminar flow hood.
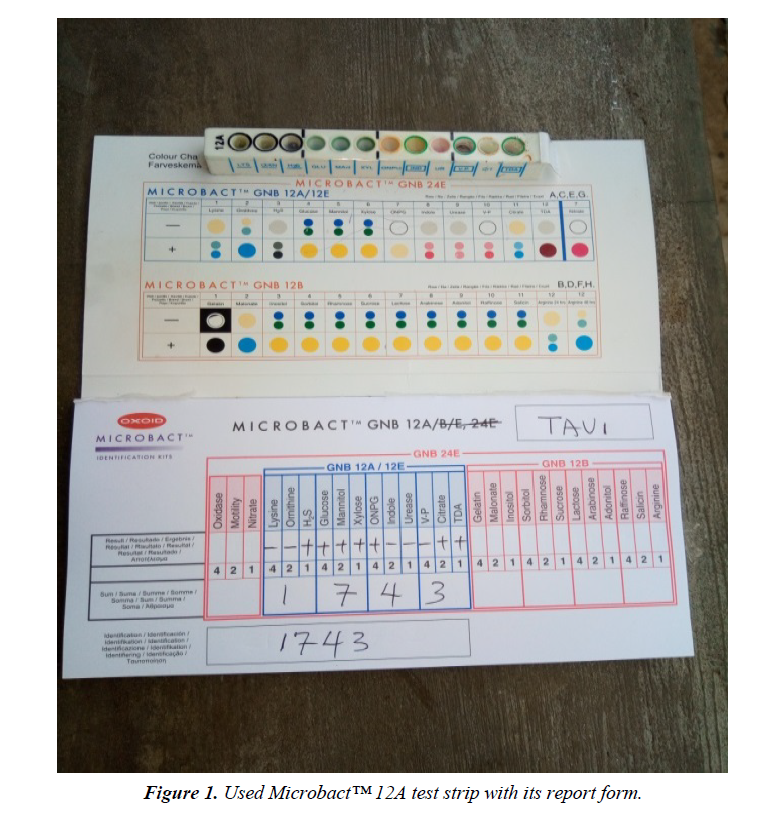
The materials used for this study include: Test tubes, Beakers, Conical flasks, Inoculating loop, Sterile needles and syringes, McCartney bottles, Hand gloves, non-absorbent cotton wool, Aluminum foil, Alcohol (95%), Potassium hypochlorite solution, Paper tape, Petri dishes, Universal sterile bottles, Sterile distilled water, Mueller Hinton agar (Oxoid, UK), Microbact™ 12A and 24E identification Kits (Oxoid, UK), and some selected Antibiotics discs (Oxoid, UK) (Figure 1).
Sample collection and identification of bacteria isolates
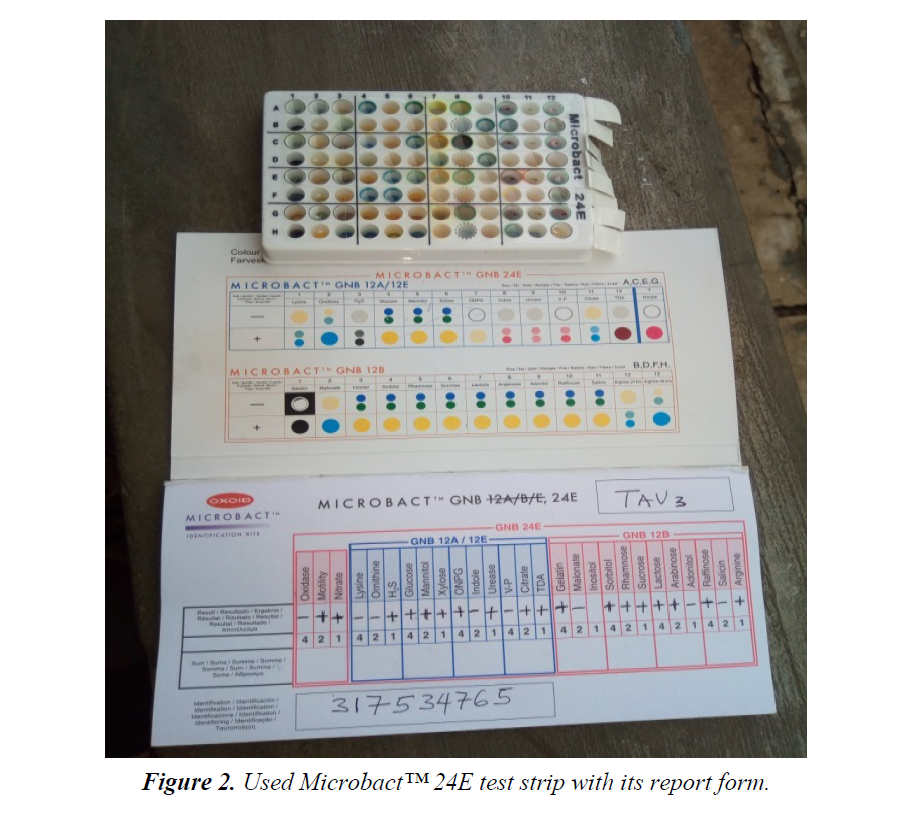
Twenty (20) non-duplicate non-repetitive presumptive clinical isolates of clinical specimens were recovered from patients using the laboratory services of University College Hospital (UCH), Ibadan, between December, 2016 and March, 2017. The isolates were then transported using prepared Nutrient agar slant to the Microbiology laboratory of Adekunle Ajasin University for further microbiological analysis. The clinical isolates were recovered from the following specimens: sputum, blood, urine and stool. Preliminary confirmatory test to identify the isolates as Citrobacter was then performed using Oxidase strips and Microbact™ 12A and 24E (Oxoid,UK) identification kits according to the manufacturer’s instruction. The recovered isolates were then prepared and sub cultured into nutrient agar slants and refrigerated at -4°C in readiness for further use (Figure 2).
Antibiogram profile
Antibiotic susceptibility tests were performed to determine the phenotypic resistance patterns of the identified Citrobacter spp isolates to some common antibiotics available in Nigeria. This test was carried out by Kirby-Bauer’s disc diffusion method on Mueller-Hinton agar (CM337-Oxoid, UK) using 10⁴ normal saline dilution of a 24 hours broth culture (Figure 3). The results were interpreted according to the Clinical Laboratory Standard Institute (CLSI, 2016) guidelines and the diameter of zones of inhibition measured and recorded in millimeter. The choice of antibiotics was based on their mechanism of action on bacteria and also on the principle of the commonly prescribed antibiotics in Nigeria’s Tertiary Health Institutions. The standard antibiotic discs (Oxoid, UK) used with their concentrations include Ceftazidime (30 μg), Cefoxitin (30 μg), Cefotaxime (30 μg), Amoxycillin+Clavullanic acid (30 μg), Cotrimoxazole (25 μg), Ofloxacin (5 μg), Imipenem (10 μg), Piperacillin/ Tazobactam (110 μg), Cloxacillin (5 μg), Oxacillin (1 μg), Piperacillin (100 μg), Meropinem (10 μg), Vancomycin (30 μg), Teicoplanin (30 μg) and Nystatin (100 IU) (Figure 3).
Screening for ESBL producers
The Citrobacter spp isolates being resistant to the cephalosporins were further processed for ESBL detection by the combined disc potentiation method according to the Clinical Laboratory Standard Institute (CLSI, 2016) guidelines. Discs of Cefotaxime (30 μg), Ceftazidime (30 μg) and Cefoxitn (30 μg) were each placed 20mm away from a combination disc of Amoxycillin+Clavullanic acid (30 μg) on a Mueller-Hinton agar plate inoculated with a 24 hours broth culture of the isolates diluted to 10⁴ inoculum density with normal saline. The results were then read after 16-24 hours incubation.
Minimal inhibitory concentration (MIC)
Minimal inhibitory concentration (MIC) was carried out using the agar dilution method according to the Clinical Laboratory Standard Institute (CLSI, 2016). Various concentrations of Piperacillin/Tazobactam were tested against the clinical Citrobacter isolates that were susceptible to the antibiotic with a zone of diameter ≥ 18 mm. A 24 hour broth culture was inoculated into Mueller-Hinton agar plates containing varying concentration of Piperacillin/Tazobactam incubated aerobically for 24 hour at 37°C under appropriate conditions.
Results
The distribution of the isolates among the collected clinical samples and profile of patients from whom samples were obtained is shown in Table 1. The distribution show that out of the 20 Gram negative isolates recovered, 4(20%) belonged to the Citrobacter species (C. freudii-3, C. sakazakii-1) with 50% of them recovered from urine sample. The antibiogram profile of the recovered isolates of Citrobacter spp to some selected antibiotics is represented in Table 2. Antibiogram of the study clearly demonstrated maximum sensitivity to Ofloxacin (100%), followed by Piperacillin/Tazobactam (75%) and Cotrimoxazole (50%).
Table 1. Distribution of isolates among clinical samples and profile of patients from whom samples were obtained.
| S/No | Hospital Assigned Code | Clinical Sample | Sex of Patient | Age of Patient | Organism Isolated | Laboratory Assigned Code |
|---|---|---|---|---|---|---|
| 1 | UCH P1 | Blood | ----- | ----- | Proteus mirabilis | Nil |
| 2 | UCH P2 | Sputum | ----- | ----- | Citrobacter freundii | TAV 1 |
| 3 | UCH P3 | Ear swab | ----- | ----- | Proteus mirabilis | Nil |
| 4 | UCH P4 | Blood | ----- | ----- | Acinetobacter iwoffii | Nil |
| 5 | UCH P5 | Ear swab | ----- | ----- | Citrobacter freundii | TAV 2 |
| 6 | UCH P6 | Stool | ----- | ----- | Salmonella marcescens | Nil |
| 7 | UCH P7 | Stool | ------ | ----- | Klebsiella oxytoca | Nil |
| 8 | UCH P8 | Urine | ----- | ----- | Citrobacter freundii | TAV 3 |
| 9 | UCH P9 | Sputum | ----- | ----- | Klbsiella pneumonia | Nil |
| 10 | UCH P10 | Blood | ----- | ----- | Providentia stuartii | Nil |
| 11 | UCH P11 | Stool | Male | 7 yrs | Enterobacter agglomerans | Nil |
| 12 | UCH P12 | Urine | Female | Adult | Acinetobacter iwoffi | Nil |
| 13 | UCH P13 | Urine | Female | 37 yrs | Hafnia alvei | Nil |
| 14 | UCH P14 | Urine | Male | 27 yrs | Citrobacter sakazakii | TAV 4 |
| 15 | UCH P15 | Urine | Female | 27 yrs | Enterobacter agglomerans | Nil |
| 16 | UCH P16 | Urine | Female | 27 yrs | Acinetobacter iwoffi | Nil |
| 17 | UCH P17 | Stool | Female | 22 yrs | Tetmella ptyseos | Nil |
| 18 | UCH P18 | Urine | Female | 21 yrs | Yersinia pseudotuberculosis | Nil |
| 19 | UCH P19 | Urine | Female | 22 yrs | Salmonella liquefaciens | Nil |
| 20 | UCH P20 | Urine | Male | 27 yrs | Salmonella marcescens | Nil |
Table 2. Antibiogram profiles of citrobacter spp against common antibiotics (measured in mm) ± sem.
| Antiobiotic (Concentration In µg) | TAV 1 | TAV 2 | TAV 3 | TAV 4 |
|---|---|---|---|---|
| Ceftazidime (30 µg) | NI | 40 ± 0.5 | NI | NI |
| Cefoxitin (30 µg) | NI | NI | NI | NI |
| Cefotaxime (30 µg) | NI | NI | NI | NI |
| Cotrimoxazole (25 µg) | 24 ± 0.3 | NI | 30 ± 0.2 | NI |
| Ofloxacin (5 µg) | 18 ± 0.1 | 20 ± 0.2 | 22 ± 0.2 | 25 ± 0.5 |
| Imipenem (10 µg) | 18 ± 0.2 | Ni | 35 ± 0.5 | NI |
| Piperacillin/Tazobactam (110 µg) | 26 ± 0.5 | 40 ± 0.2 | 36 ± 0.3 | NI |
| Cloxacillin (5 µg) | NI | NI | NI | NI |
| Oxacillin (1 µg) | NI | NI | NI | NI |
| Piperacillin (100 µg) | NI | NI | NI | NI |
| Amoxycillin+Clavulanic acid (30 µg) | NI | NI | NI | NI |
| Meropenem (10 µg) | NI | NI | NI | NI |
| Nystatin (100 iu) | NI | NI | NI | NI |
Keys: NI=No inhibition, SEM=Standard error of magnitude.
Result of the screening for ESBL producers is shown in Table 3. Phenotypic resistance pattern of the isolates to glycopeptides is shown in Table 4. The result showed maximum resistance (100%) to Teicoplanin and Vancomycin. The Minimal Inhibitory Concentration (MIC) of Piperacillin/Tazobactam against the susceptible strains of the recovered Citrobacter spp is presented in Table 5.
Table 3. Phenotypic screening for ESBL producer among the citrobacter isolates (measured in mm) ± sem.
| Isolates | Ceftazidime (30 µg) | Cefotaxime (30 µg) | Cefoxitin (30 µg) | Amoxycillin+Clavulanic acid (30 µg) |
|---|---|---|---|---|
| TAV 1 | NI | NI | NI | NI |
| TAV 2 | 38 ± 0.5 | NI | NI | NI |
| TAV 3 | NI | NI | NI | NI |
| TAV 4 | NI | NI | NI | NI |
Key: NI=NO Inhibition, SEM=Standard Error of Magnitude.
Table 4. Phenotypic resistance profile of citrobacter isolates to glycopeptides.
| Isolates | Teicoplanin (30 µg) | Vancomycin (30 µg) |
|---|---|---|
| TAV 1 | 13 ± 0.5 | NI |
| TAV 2 | NI | NI |
| TAV 3 | NI | NI |
| TAV 4 | 13 ± 0.5 | NI |
Key: NI=No Inhibition
Table 5. MIC pattern of susceptible citrobacter isolates to piperacillin/tazobactam.
| Isolates | MIC values |
|---|---|
| TAV 1 | 110 µg/mL |
| TAV 2 | I00 µg/mL |
| TAV 3 | 110 µg/Ml |
Discussion
This study was designed to determine the phenotypic resistance in Citrobacter species and Antibiogram profile of Citrobacter species isolated from University College Hospital (UCH), Ibadan. The present study indicates that, of the 20 clinical isolates from the University College Hospital (UCH), Ibadan, 4 (20%) were Citrobacter species (Citrobacter freundii-3, Citrobacter sakazakii-1) with 1 (25%). This is much higher than that reported by Khanna A et al. [14] (4.70% of the Citrobacter spp out of a total of 1830 gram negative bacilli), but similar to the report of Avinash et al. (25.1% out of 1000 clinical samples). This high prevalence might be due to lack of preventive measures during the handling of the patients in these hospitals. A majority of the isolates were recovered from urinary samples (50%), while the remaining 50% were from sputum and ear swab. This low prevalence of Citrobacter spp from this research is similar to the report of Onoh RC et al. [16] who isolated five Citrobacter spp from a total of 252 bacterial isolates, cultured from urinary samples. Metri BC and Jyothi P [17] reported that invasive procedures like, catheterization helps the bacteria to colonize the urinary bladder. The presence of intact immune system in the host helps in defending against the pathogen to certain extent, but when the patients are immunocompromised, the condition would be fatal.
The study by Whitby has found Citrobacter in 12% of acute infections. Some studies reported that Citrobacter species appears to be one of the contributing organisms in causing urinary tract infection [17,18]. The antibiogram profile of the Citrobacter spp (Table 2) showed a significantly high level of resistance among all the isolates. In this study, Ofloxacin was active against all the four Citrobacter isolates (100%), with diameter zone of inhibition ranging from 18 mm to 25 mm. Closely followed is Piperacillin/Tazobactam which was active against three out of the four isolates (75%) with diameter zone of inhibition ranging from 26 mm to 40 mm. Two of the isolates (50%) showed susceptibility to Co-trimoxazole (24 mm and 30 mm), and Imipenem (18 mm and 35 mm). The Citrobacter spp also demonstrated resistance to the β-lactam antibiotics. The rate of resistance among the isolates to Cephalosporins was high. Cefoxitin and Cefotaxime resistance were seen in all the four isolates (100%) while Ceftazidime resistance was observed in three of the isolates (75%). All the isolates were, however, resistant to Cloxacillin (100%), Oxacillin (100%), Piperacillin (100%) and combination of Ampicillin+Clavulanic acid (100%), Meropenem (100%) and Nystatin (100%). Citrobacter sakazakii demonstrated extreme multidrug resistance to all the drugs used except Ofloxacin. Over-prescription of antibiotics, lack of compliance, extensive use of antibiotics in Agriculture, poor infection control in health care settings, poor hygiene and sanitation and absence of new antibiotics being discovered have been linked to the development of resistant bacterial strains. Another likely cause of antibiotic resistance is poor drug quality which has been a major threat to healthcare system in Nigeria. In this study, Citrobacter freundii was the commonest species which was isolated, as also was the case in other studies [14,18]. ESBL production was found to be phenotypically absent in all the isolates (Table 3), using the combined disc potentiation method which differs from the 25% (1 out of 4) occurrence reported by Yusha’u (2010). It has been reported that the occurrence and distribution of ESBL producers among Enterobacteriaceae differs from country to country and hospital to hospital. However, the resistance shown by these Citrobacter spp isolates might be due to other mechanisms of resistance other than β-lactamase production. As shown in Table 5, the MIC values of Piperacillin/Tazobactam tested against the Citrobacter isolates are very high (100 μg/mL and 110 μg/ mL). This may be a pointer that these isolates could develop resistance to these combined antibiotics if proper antibiotic administration measures are not put in place.
The strength of this study is the broad population of interest, involving both the in and out-patients samples reflects the circulating resistance, and the wide range of antibiotics used, involving almost all classes of antibiotics. There are however some limitations. This study had a relatively small sample size. This is due to the low prevalence of Citrobacter species among the isolates from the two tertiary health institutions, which has also been reported across the globe. Susceptibility testing was also carried out using antibacterial from few groups of antibiotics, excluding the aminoglycosides, tetracyclines, and macrolides.
Further area to be exploited for future research is the genetic diversity and the characterization of these antibiotic resistant Citrobacter strains. This will enable us confirm the mechanisms of resistance exhibited by these strains and the various genes responsible for their resistance.
Conclusion
Citrobacter species are emerging as important resistant pathogens. In this study, Citrobacter freundii was the commonest species isolated and the University Teaching Hospital (UCH), Ibadan in Southern Nigeria. Multidrug resistance is quite high among the Citrobacter species. These multidrug resistant strains were mostly sensitive to a combination of Piperacillin/ Tazobactam. Greater caution is required in the selection of the antibiotic/antibiotics to be used in treatment of patient with Citrobacter infections to avoid treatment failure. This bacterium can become resistant to these drugs at any time, and hence, the indiscriminate, inadequate and the prophylactic use of these antibiotics should be avoided. Infection control measures like disinfection of the wards and equipment, barrier precautions against colonized patients and hand hygiene measures should be strictly adhered to, to prevent the spread of the pathogen. The antibiogram is a cost effective, easy and rapid typing tool in a low resource country like Nigeria, where molecular typing tools are not easily accessible. Only in case of emergency, empirical therapy which is based on the antibiotic sensitivity pattern of that institution should be given to the patients.
References
- Birdsall PD, Hayes JH, Cleary R, et al. Health outcome after total knee replacement in the very elderly. J Bone Joint Surg. 1999;81(4):660-662.
- Baker PN, van der Meulen JH, Lewsey J, et al. The role of pain and function in determining patient satisfaction after total knee replacement: data from the National Joint Registry for England and Wales. J Bone Joint Surg.2007;89(7):893-900.
- Kurtz S, Ong K, Lau E, et al. Projections of primary and revision hip and knee arthroplasty in the United States from 2005 to 2030. J Bone Joint Surg. 2007;89(4):780-5.
- Jones CA, Voaklander DC, Johnston DW, et al. Health related quality of life outcomes after total hip and knee arthroplasties in a community based population. J Rheumatol. 2000;27(7):1745-52.
- Bachmeier CJ, March LM, Cross MJ, et al. Arthritis Cost and Outcome Project Group. A comparison of outcomes in osteoarthritis patients undergoing total hip and knee replacement surgery. Osteoarthritis Cartilage. 2001;9(2):137-46.
- Robertsson O, Dunbar M, Pehrsson T, et al. Patient satisfaction after knee arthroplasty: a report on 27,372 knees operated on between 1981 and 1995 in Sweden. Acta Orthop Scand.2000;71(3):262-7.
- Gandhi R, Davey JR, Mahomed NN. Predicting patient dissatisfaction following joint replacement surgery. J Rheumatol.2008;35(12):2415-8.
- Gunarate R, Pratt DN, Banda J, Fick DP, et al. Patient dissatisfaction following total knee arthroplasty: A systematic review of the literature. J Arthroplasty. 2017;32(12):3854-60.
- Ross CK, Steward CA, Sinacore JM. A comparative study of seven measures of patient satisfaction. Med Care. 1995;33(4):392-406.
- Kim TK, Chang CB, Kang YG, et al. Causes and predictors of patient’s dissatisfaction after uncomplicated total knee arthroplasty. J Arthroplasty.2009;24(2):263-71.
- Eisler T, Svensson O, Tengstrom A, et al. Patients expectation and satisfaction in revision total hip arthroplasty. J Arthroplasty. 2002;17(4):457-62.
- Mahomed NN, Liang MH, Cook EF, et al. The importance of patient expectations in predicting functional outcomes after total joint arthroplasty. J Rheumatol.2002;29(6):1273-9.
- Scott CE, Howie CR, MacDonald D, et al. Predicting dissatisfaction following total knee replacement: a prospective study of 1217 patients. J Bone Joint Surg. 2010;92(9):1253-8.
- Hamilton DF, Lane JV, Gaston P, et al. What determines patient satisfaction with surgery: a prospective cohort study of 4709 patients following total joint replacement. BMJ Open. 2013;3(4):e002525.
- Bourne RB, Chesworth BM, Davis AM, et al. Patient satisfaction after total knee arthroplasty: who is satisfied and who is not? Clin Orthop Relat Res. 2010;468(1):57-63.
- Zacharia B, Paul M, Thanveeruddin Sherule M. Patient-based outcome analysis is important to determine the success of total knee arthroplasty: result of a focus group discussion. Med Devices. 2016;9:125-130.
- Culliton SE, Bryant DM, Overend TJ, et al. The relationship between expectations and satisfaction in patients undergoing primary total knee arthroplasty. J Arthroplasty. 2012;27(3):490-2.
- Lopez-Olivo MA, Landon GC, Siff SJ, et al. Psychosocial determinants of outcomes in knee replacement. Ann Rheum Dis. 2011;70(10):1775-81.
- Mantyselka P, Kumpusalo E, Ahonen R, et al. Patients’ versus general practitioners’ assessments of pain intensity in primary care patients with non-cancer pain. Br J Gen Pract. 2001;51(473):995-7.
- Scuderi GR, Bourne RB, Noble PC, et al. The new knee society knee scoring system. Clin Orthop Relat Res. 2012;470(1):3-19.
- Chesworth BM, Mahomed NN, Bourne RB, et al. Willingness to go through surgery again validated the WOMAC clinically important difference from THR/TKR surgery. J Clin Epidemiol. 2008;61(9):907-18.
- Thambiah MD, Nathan S, Seow BZ, et al. Patient satisfaction after total knee arthroplasty: an Asian perspective. Singapore Med J. 2015;56(5):259-63.