Research Article - Biomedical Research (2016) Computational Life Sciences and Smarter Technological Advancement
Extraction of perfect protein sequences with minimal processing cost using enhanced B+ tree algorithm
Mary Posonia A1*, Jyothi VL21Department of Computer Science and Engineering, Sathyabama University, Chennai, India
2Department of Computer Science and Engineering, Jeppiaar Engineering College, Chennai, India
- *Corresponding Author:
- Mary Posonia A
Department of Computer Science and Engineering
Sathyabama University
India
Accepted on August 18, 2016
Abstract
The sequence of protein comparison and classification is increasing hugely in current era of ‘omics’ revolution. The functionality of proteomics and genomics are creating a large prediction over protein functionality. Query based protein sequence information retrieval from protein Extensible Markup Language (XML) dataset is a challenging scenario in a real world application. The major problem that rises in medical information retrieval is that it cannot retrieve the exact protein content or its ingredients information. Also in retrieving such information the computation cost finds to be much higher and also retrieval accuracy about the particular protein sequence is very minimal. Apart from this another major issue focused is of the communication loss which happens when users communicating with protein Extensible Markup Language (XML) dataset for information retrieval. In order to overcome the above mentioned issues here proposed a new method which resolves the complexity of communication loss of protein Extensible Markup Language (XML) dataset and to retrieve exact protein content or its ingredients information with minimum computation cost. The proposed new method which is of an enhanced B+ tree indexing algorithm helps in retrieving the protein information from protein Extensible Markup Language (XML) dataset with minimal computation cost and accurate result pattern. The proposed approach is designed in such a way to extract the perfect protein similarity matrix from the protein sequence that is extracted from the protein Extensible Markup Language (XML) dataset. All the similarity matrices of protein sequence are generated and collected by the collection of sequence of protein based on users query request. Thus Experimental result show how efficient the proposed enhanced B+ tree indexing algorithm helps in retrieving the protein information from protein Extensible Markup Language (XML) dataset with minimal computation cost and accurate result pattern.
Keywords
B+ tree indexing, Protein sequence, Information retrieval, Data mining, Protein family, Omics, Protein functionality, Proteomics, Genomics.
Introduction
Biomedical Engineering is one of the latest fields in modern science and it is evolving constantly in the promising field. The analysis of proteins is a huge challenge in the field of biomedical Engineering because of its large volume of biological Protein data [1]. Due to the increased size of biological protein data the sequence retrieval of protein pattern became a big challenge. The requirement of efficient searching of proteomics and the retrieval tolls became most extreme process because of its higher execution time. The efficient retrieval of proteomics within this process requires a huge size for storing the data and it utilizes a large volume of memory [2]. Also there are several techniques existing for the retrieval of proteins sequence [3]. There is one technique that could help to provide the efficient speed for accessing and searching the content from biological protein database is of biological protein database indexing. The main reason for utilizing this biological protein database indexing is, since it is faster and efficient when compared to other local biological protein database storages. Several numbers of researchers have made a lot of effort in several previous years for developing biological tools for fast biological query retrieval.
The searches over protein sequences are commonly done for analysing the uncharacterized sequences of proteins within a high degree of implied similarity in the pair of protein sequence homology [4]. These searches are based on the similarity and are really helpful in identifying protein sequence patterns. The similarity sequences of protein are being represented based on the global or local searches and a pair wise alignment of sequences with a mathematical optimization [5]. These alignments of protein sequence are being detected based on the significant relationship over a huge dataset within some basic tools of a bioinformatics task [6]. The process of annotating the sequence task for protein sequence is being used as an evidence for the knowledge from known or unknown protein dataset sequence. The presentation of protein family through several alignments required comparison over pair wise and are created as a sequence profile based on the pair wise alignments.
Initially protein sequence files are pre-processed using preprocessing algorithm like lexical preparation [7]. After preprocessing the protein sequence document the group nodes will be constructed. Then the grouped protein sequence document will take as an input for protein sequence indexing process. Thus this protein sequence indexing process indexes the nodes and stores in biological protein database.
Whenever the user submits a protein name as a query in search box, the protein name request is forwarded to biological protein Extensible Markup Language (XML) database server. Then the biological protein Extensible Markup Language (XML) database server searches for protein sequence value using the sequence of keywords and the relevant protein sequence data will be retrieved. The mechanism even with online and offline scenario, i.e., even if there is any overload or communication breakage in network. The existing drawback in biological protein Extensible Markup Language (XML) database server is when it receives more number of requests from many users then the communication of some users may get break and the searching process will start again and the user has to resubmit their biological query. Since due to this the searching time and cost gets increased [8].
Here in this paper proposed an enhanced B+ tree indexing algorithm and log file creation approach for biological protein Extensible Markup Language (XML) database server. Thus this approach creates a log file whenever a user login and communicate and also the log maintain each individual user id, Internet Protocol (IP) address, user protein query and output which is of protein sequence. Also the detailed explanation of the proposed work is explained in Section 3.
Related Works
The author has proposed indexing of bitmap structural technique that using the database encode and condense sequence for indexing smaller part of protein sequence [9]. Mainly, the profit of this approach is to reduce the response time and to add additional space. The algorithmic component is the procedure of Builder Information System (BIS) construction, phase of filtering and analysis of result.
Weimin et al. proposed a technique of indexing with space matrix, which is investigating two major and crucial techniques for supporting the evaluation of fast query with the alignment [10].
Hsiao et al. proposed IDC (Incrementally Decreasing Coverbased) extraction technique for genomic based homology dataset [11]. The novel concept was being introduced within this approach, that approach is having filtration process within “homology candidate” and “annote query sequence”. There is one important characteristic about this algorithm is that it has a lossless filtration property.
Elberrichi et al. has proposed an N-Gram technique for identifying the protein sequence through several fields including the process of natural language statistical science and the process for analysis of genetic sequence [12]. The search item could be a protein sequence, letters or words based on application.
B+ tree is extensively used for indexing Extensible Markup Language (XML) blobs in relational databases and it is commonly a selected indexing mechanism for indexing. This B + tree efficiently estimates the queries on indexed Extensible Markup Language (XML) blobs. Query execution should reassemble the results of Extensible Markup Language (XML) from B+ tree indexing to conserve document structure and document order. Many operators in X Path 2.0 particularly are self-axis or descendant and recursively navigate down on Extensible Markup Language (XML) tree [13]. Therefore B+ tree lookups also be recursive.
In the study [14], a ranking model named XRank is proposed to rank Lowest Common Ancestor (LCA) results from Extensible Markup Language (XML) element level to page level. A new ranking method RACE is introduced for ranking trees by textual and structural similarity [15]. In XSearch, the content and structure are considered for ranking Extensible Markup Language (XML) documents [16]. A novel approach called XReal is proposed for ranking relevant and creates a formula for searching the recognized nodes and a query ranking mechanism Extensible Markup Language (XML) documents using an adaptive keyword search on nodes [17]. A study examined the descendant nodes placed at top position then their antecedent nodes in the stack. From this examination a new stack-based join algorithm is proposed [18].
Williams et al. explains partitioned related method [19]. In this method coarse searching technique with inverted index process is used for similarity ranking. The subset of database with query is utilized in a consequent fine search. Index of this method contains three components that are search structure, mapping table and inverted lists. Here compression idea is used for make the more manageable index size. Particularly, CAFE approach contains of fine search and coarse technique is slightly fewer accurate than FASTA and BLAST1. In search point of vision, CAFE method is 8 times quicker and proficient than method of BLAST.
Proposed Methodology
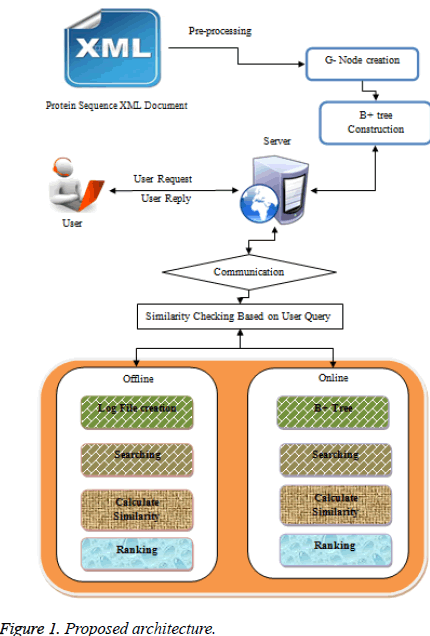
The proposed architecture mainly concentrates to reduce the cost of query based result computation. In Figure 1 initially the protein sequence documents are pre-processed by creating Gnode and B+ tree construction and the processed documents are stored in biological protein Extensible Markup Language (XML) database server. Whenever user request a protein query in online-biological protein Extensible Markup Language (XML) database server it performs indexing and searching based on keywords present in the user protein query request.
In internet scenario due to overload or link breakage user cannot retrieve the data, in this situation server will generate the log file for reducing the searching time. The detailed explanation of log file creation and indexing procedure is explained in the following section.
A. Sample Extensible Markup Language (XML) protein sequence document
<? Extensible Markup Language (XML) version="1.0"?>
<! DOCTYPE repository system "/projects/null/Extensible Markup Language (XML) tk/Extensible Markup Language (XML) data/data/repository.dtd">
<repository>
<Path>Extensible Markup Language (XML) tk/Extensible Markup Language (XML) data/data/pir</path>
<name>pir</name>
<human-name>protein sequence database</human-name>
<entry-descr>integrated collection of functionally annotated protein sequences.</entry-descr>
<author>Margolliash, E</author>
<contents>annotation</contents>
<contents>composition of chymotryptic peptides</contents>
<classification>
<superfamily>cytochrome c</superfamily>
<superfamily>cytochrome c homology</superfamily>
</classification>
<keywords>
<keyword>acetylated amino end</keyword>
<keyword>chromo protein</keyword>
, keyword>electron transfer</keyword>
…………
<keywords>
</classification>
GNode= (PNode, NNode, IDNode)
B. Algorithm for group-node creation GNode = (PNode , NNode , IDNode)
Input: Protein sequence Extensible Markup Language (XML) file
Output: G-Node
Step 1: consider every node as input
Step 2: if Node1>1 contains child node
Step 3: Construct G-node for Node1
Step 4: GNode= (PNode, NNode, IDNode)
Step 5: Repeat Step 2, 3 and 4 for every node
C. Procedure for B+ tree insert operation
If (Data page! =full && index page! =full)
Return leaf page sort
Else if (Data page=full && index page! =full)
leaf page/Right side leaf
leaf page/left side leaf
if (middle key generation=Ascending)
Left leaf page<middle key values
if (middle key generation=Descending)
Right leaf page ≥ middle key values.
Else if (Data page=full && Index Page=full)
leaf page/Right side leaf
leaf page/left side leaf
if (Records key value<middle key values)
go left leaf page
if (Records key value>middle key value )
go right leaf page
index page/right index
index page/left index
If (key value<middle key value)
Go left index page
If (key value>middle key value)
Go right index page
middle key → Hindex
Return;
Call left index;
Call right index;
End if
D. Similarity-based-search
The B+ tree consists of root node and leaf node. The records stored in root node once root node is full then split into internal or leaf nodes. The leaf node doesn’t contain any node in B+ tree. Each leaf node contains balanced records. Normally each node contains the keys. The n number of nodes having keys such as K1, K2…Kn.
K1<K2<………………Kn-1
In B+ tree number of nodes formed a tree structure in balanced manner. The minimum number of nodes in B+ (n/2)-1 tree to maximum n-1.
(n/2)-1 ≤ n ≤ b-1
The B+ tree improves the query processing performance. The query consider as a token tag. The tokens are transformed in the form of internal and check the similarity. Each node having the key based on the key it compares the key value and indexing Extensible Markup Language (XML) blobs. Based on the similarity of key value every related tag will be rank.
E. Protein sequence retrieval annotation in the entries of protein sequence with similarity
B+ tree is completely able to retrieve the complete set of annotated protein sequence in the protein bank protein sequence. Extensible Markup Language (XML) database entries. The sequence of proteins chromosome is containing the genes annotation corresponding to proteins. The protein contents are retrieving all the information of sequence from Extensible Markup Language (XML) file. When G-Node identifier of a protein is entered then the ingredients of the proteins will be present in the respective sequence entry.
A protein sequence entry id representing a huge sequence such as complete chromosome might be having several properties within different sequence entries rather than a huge protein string sequence.
The hierarchical Extensible Markup Language (XML) dataset of proteins structure is being observed for calculating the Extensible Markup Language (XML) node similarity that preferred for searching based on keyword. The created G-Node is computing similarities among leaf node of protein Extensible Markup Language (XML) data and the queries, then it recursively computing the similarities amid of internal nodes and value of each chromosome with queries.
G. Protein sequence characteristics
The process of non-redundant sequence of the protein sequence is being supplemented within the characteristics of proteins and the references of secondary element from the database of protein families. The information of associated dataset is being automatically updated when novel sequences are imported into the protein Extensible Markup Language (XML) dataset.
H. Communication loss log file creation
This operation is an important task in our research. Normally in network scenario, more number of users can send the request and retrieve the data. But main disadvantage in wireless network is communication overhead. For example, if more number of user’s access the network channel at the same time, communication of some user gets break. At that time the server creates log file.
Log file: Log file is a file to resume the searching process again. This log file contains user Internet Protocol (IP), query, tag name, path. Whenever the communication loss occurs while searching the G-nodes in B+ tree the server creates the log file.
Log file=LF (Internet Protocol (IP), time, node name, value);
Algorithm:
Input: every Extensible Markup Language (XML) file, user query
Output: log File
Step 1: get keyword from query
Step 2: Search values in Extensible Markup Language (XML) file using B+ tree
Step 3: if more numbers of user accessing the data (or) bandwidth gets overload
Step 4: create log file (Internet Protocol (IP), time, node id, node name, filename)
Step 5: server receives same request
Step 6: if user Internet Protocol (IP) exists in Log file and query exists in `log file and time<threshold value
Step 7: start to search using node id, node name and filename.
Step 8: End
For example, normally the user enters the query and sends to server. The server receives the request and searches in stored G-nodes. But the main drawback in wireless network is communication loss whenever more number of users sends the request to server. So the communication gets break while searching the data and the server creates log file. This log file contains user entered query, tag name, path and user Internet Protocol (IP). The server stores the log file after creating the log file. Whenever the same user enters the same query in particular time period (5 min) the server checks in log file in data database. If the Internet Protocol (IP) and entered query is same, then the server will take the path and tag name as input and starts to search the query.
(i) Execution cost:
Execution cost of the individual query can be calculated based on processing time, number of files and memory utilization.
Ecpq=Tp*nf+Tp*Mu → (1)
Where, Ecpq-execution cost per query
Tp-processing time
nf-number of files
Mu-memory utilization
Ecent=Qn (Tp*nf+Tp*Mu) → (2)
Where, Ecent-execution cost for entire network
Qn-number of queries
Tp-processing time
nf-number of files
Mu-memory utilization
Results and Analysis
For Experiments were carried out based on the following configuration Windows 7, Intel Pentium (R), CPU G2020 and processor speed 2.90 GHz and jdk 1.7. Experiments have been done with the protein sequence Extensible Markup Language (XML) document from Extensible Markup Language (XML) data repository protein sequence Extensible Markup Language (XML). The number of elements 21305818 and the attributes are 1290647 with the max-depth is 7. Initially the server starts to pre-process the data and stores in database. This pre-process process contains generating keywords, indexing, constructing B+ tree and constructing G-nodes. Then the user starts to search the required details by entering the query. Then the server receives and starts to search the requested details and finds the most relevant details based on similarity and response to requested user. Whenever the channel gets overloaded, the server creates the log file for that particular user and stores in a temporary folder. If within that particular time period the server receives the request from same user, then the server uses the stored data and starts to search. After that particular time period the server deletes the file automatically. Table 1 explains about the number of users accessing the data in parallel and time for searching and retrieving the data from data storage. Table 2 shows the validation time based on number of users and log files. Table 3 shows the system performance can be described based on number of request and searching time. Table 4 proves the execution cost of individual query generated by user. The execution cost is depends on time, number of files and memory utilized, however the entire network computation cost is depends on Ecpq.
| S.No. | No of users (parallel access) | Time for transmitting request (Ms) | Time for protein sequence retrieval/630Mb (Ms) |
|---|---|---|---|
| 1 | 30 | 300 | 233 |
| 2 | 50 | 320 | 250 |
| 3 | 70 | 350 | 266 |
Table 1. Protein sequence data accessing details.
| S. No. | No. of users (Parallel access) | No of log Files | Validation time (in minutes) |
|---|---|---|---|
| 1 | 400 | 30 | 3 |
| 2 | 600 | 40 | 3 |
| 3 | 700 | 43 | 3 |
Table 2. Protein sequence data access details with user log file.
| S. No | Number of requests (Parallel access) | Searching time (in ms) |
|---|---|---|
| 1 | 400 | 200 |
| 2 | 600 | 300 |
| 3 | 700 | 356 |
Table 3. Biological protein Extensible Markup Language (XML) database server performance.
| Query | Tp (ms) | nf | Mu (KB) | Ecpq(ms per query) |
|---|---|---|---|---|
| Q1 | 300 | 50 | 20 | 21000 |
| Q2 | 320 | 110 | 40 | 48000 |
| Q3 | 350 | 160 | 45 | 71750 |
Table 4. Biological protein Extensible Markup Language (XML) database server execution cost.
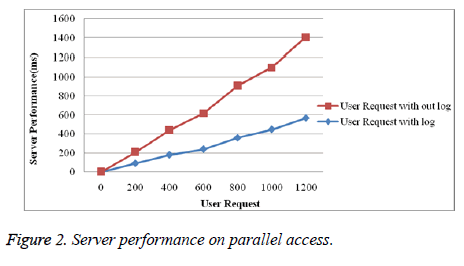
Figure 2 shows performance graph of server with log file and without log file. Here with log file system works much better than without log file if more number of users access is increase in parallel.
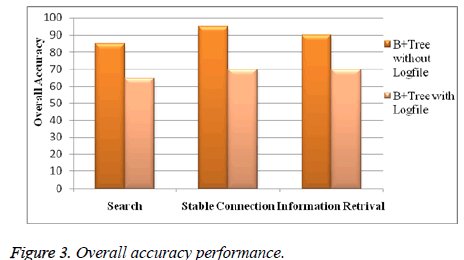
B. Overall accuracy performance
Figure 3 shows accuracy performance of overall system. By this system retrieving protein information from protein Extensible Markup Language (XML) database is achieved faster and accurately.
This system gives better result compare with other system in area of exact information retrieval, fast result and stable connection with protein Extensible Markup Language (XML) data base.
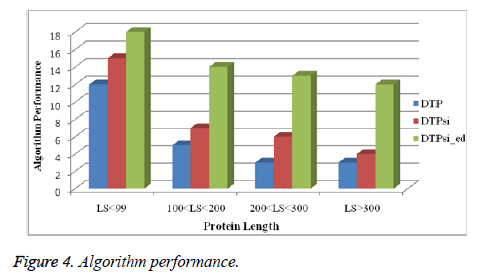
The above mentioned Figure 4 is describing the performance of algorithm with several protein lengths. The protein length has been counted amid of 99 to 300.
The algorithm is considering three term that is named as DTP, DTPsi and DTPsi_ed.
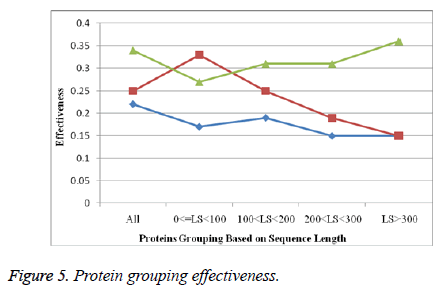
D. Protein grouping effectiveness
The below mentioned Figure 5 is describing about the protein grouping effectiveness, where the sequence length of protein is showing the relation amid of several length and their similarity.
The effectiveness of similarity is being described here.
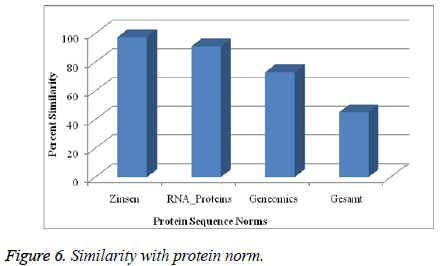
E. Similarity with protein norm
The above mentioned Figure 6 is describing about the similarity percentage over the several norm of proteins. The Zinsen, RNA, Genomics and Gesamt have been considered for the similarity measurement. The protein norm has similarity function approximately with the entire proposed norm.
Conclusion
The proposed enhanced B+ tree indexing algorithm retrieves a perfect protein sequence pattern from protein Extensible Markup Language (XML) dataset with a very minimal computation cost. Apart from this it also reprocesses a perfect protein sequence pattern from user query which is a protein name thus all the proteins are being annotated in single entry of DNA sequence. Also the proposed algorithm produces a perfect protein sequence pattern as uninterrupted and effective communication from protein Extensible Markup Language (XML) dataset even if the network traffic or communication breaks stage. Thus this approach completely avoids reprocessing of user query due to this its processing time will be efficiently utilized. Also a G-Node is constructed for grouping a perfect protein sequence pattern from protein Extensible Markup Language (XML) dataset for user query and a log file is created for retrieving the protein Extensible Markup Language (XML) query result without reprocessing the user query even if in traffic condition or communication loss. Thus the proposed approach obtains a faster processing of protein query result output from the protein Extensible Markup Language (XML) dataset even if in idle or busy network condition. Thus the proposed technique produces an enhanced visual inspection of proteins sequence pattern entries in local system from protein Extensible Markup Language (XML) dataset, where as it store millions of protein sequence records locally and retrieve efficiently based on the user request for any particular protein name.
References
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J MolBiol 1990; 215: 403-410.
- Pearson WR. Flexible sequence similarity searching with the FASTA3 program package. Methods MolBiol 2000; 132: 185-219.
- Gojobori T, Li WH, Graur D. Patterns of nucleotide substitution in pseudo genes and functional genes. J MolEvol 1982; 18: 360-369.
- Smith T, Waterman M. Identification of common molecular sub-sequences. J MolBiol 1981;147: 195-197.
- Krause A, Haas SA, Coward E, Vingron M. SYSTERS, GeneNest, SpliceNest: Exploring sequence space from genome to protein. Nucleic Acids Res 2002; 30: 299-300.
- OBrien KP, Remm M, Sonnhammer EL. Inparanoid: A comprehensive database of eukaryotic orthologs. Nucleic Acids Res 2005; 33: D476-480.
- Li L, Stoeckert CJ Jr, Roos DS. OrthoMCL: Identification of ortholog groups for eukaryotic genomes. Genome Res 2003; 13: 2178-2189.
- Pellegrini M, Marcotte EM, Thompson MJ, Eisenberg D, Yeates TO. Assigning protein functions by comparative genome analysis: Protein phylogenetic profiles. ProcNatlAcadSci USA 1999;96: 4285-4288.
- Beng Chin O, HweeHwa P, Hao W, Limsoon W, Cui Y. Fast filter-and-refine algorithms for subsequence selection. ProcInt 2002; 243-254.
- Weimin C, Aberer K. Efficient querying on genomic databases by using metric space indexing techniques. IEEE Comp Soc 1997.
- Hsiao Ping L, Yin Te T, ChingHua S, Tzu Fang S, Chuan Yi T. An IDC-based algorithm for efficient homology filtration with guaranteed seriate coverage.Proc IEEE BioinformBioeng 2004; 395-402.
- Elberrichi Z, Aljohar B.N-grams in texts categorization. Sci J King Faisal Univ Basic ApplSci 2007; 8: 25-38.
- Li G, Feng J, Wang J, Yu B, He Y. Race-finding and ranking compact connected trees for keyword proximity search over XML documents. WWW 2008; 1045-1046.
- Hucka M, Finney A, Sauro HM, Bolouri H, Doyle JC. The systems biology markup language (SBML): A medium for representation and exchange of biochemical network models. Bioinformatics 2003; 19: 524-531.
- Bao Z, Ling T, Chen B, Lu J. Effective XML keyword search with relevance oriented ranking. Data Eng2009; 517-528.
- Al-Khalifa S, Jagadish HV, Koudas N, Patel JM, Srivastava D, Wu Y. Structural joins: A primitive for efficient XML query pattern matching.ProcInt Data Eng2002; 141-152.
- Bruno N, Srivastava D, Koudas N. Holistic twig joins: Optimal XML pattern matching. ProcAcmSigmodIntManag Data 310-321.
- Waugh A, Gendron P, Altman R, Brown JW, Case D. RNAML: A standard syntax for exchanging RNA information. RNA 2002; 8: 707-717.
- Williams HE, Zobel J. Indexing and retrieval for genomic databases, knowledge and data engineering. IEEE Trans Knowl 2002; 14: 63-78.