Research Article - Biomedical Research (2018) Volume 29, Issue 2
Expression of eNOS and CAV gene in trabecular meshwork cells and effects on glaucoma pathogenesis
Yan Chen, Shudan Ge, Zhihui Lin and Zhihong Liu*
Changchun University of Chinese Medicine, Changchun, Jilin, 130021, PR China
Accepted date: October 27, 2017
DOI: 10.4066/biomedicalresearch.29-17-3139
Visit for more related articles at Biomedical ResearchAbstract
Objective: Glaucoma is the second popular blinding disease worldwide, and is mainly caused by abnormal function or structure of trabecular meshwork and consequent aqueous humor effluent dysfunction, ocular hypertension, leading to vision deficit. eNOS and CAV1 might be correlated with glaucoma onset. This study thus investigated their expression in trabecular meshwork cells and effects on glaucoma pathogenesis.
Patients and Methods: Trabecular meshwork tissues were collected during surgery from POAG patients, and eNOS/CAV1 gene expressions were analyzed by qRT-PCR and Western-Blot. eNOS and CAV1 genes were over-expressed in cultured HTMC cells, whose survival rate was analyzed by MTT assay, whilst cell free radicals clearance rate was analyzed by chemiluminescence method.
Results: HE staining revealed significant pigment deposition in trabecular meshwork of POAG patients compared to normal group, accompanied with stenosis of mesh. Expression levels of eNOS and CAV1 were significantly higher in POAG group (p<0.05 compared to healthy control group). In vitro overexpression of eNOS and CAV1 significantly suppressed cell survival rate and free radical clearance rate (p<0.05).
Conclusion: Trabecular meshwork cells in glaucoma patients showed elevated eNOS and CAV1 expression. Up-regulation of eNOS expression may suppress cell survival rate at cellular level, impaired ability to clear free radicals, eventually leading to cell or tissue damage.
Keywords
Glaucoma, Trabecular meshwork, eNOS, CAV1
Introduction
Glaucoma is presented as intermittently or persistently elevated intraocular pressure, which can impair ocular tissue and visual functions [1]. Glaucoma is the second blinding disorder worldwide, having an incidence of about 1% [2]. Glaucoma severely threatens patient visual acuity. However, pathogenesis mechanism of glaucoma is still unclear yet. Currently major stream of glaucoma treatment in clinical practice targets the management of ocular hypertension, including medicine and surgery. However, commonly used drugs may develop adverse effects to various extents with sometimes major complications, and surgical approaches such as filtration operation still cannot obtain satisfactory effects [3]. Therefore, the identification of molecular mechanism related with glaucoma pathogenesis is of critical importance of preventing and treatment of glaucoma and hence force the vision dysfunction correction.
Major reason for ocular hypertension in glaucoma is due to efflux congestion of aqueous humor [4]. Trabecular meshwork is one filtrating structure of human eyeball. It is located at anterior chamber angle, and is one cavernous-like structure composed of cross-liked multi-laminar and spindle meshwork tissues. As the major efflux pathway of aqueous humor, abnormal structure or function of trabecular meshwork blocks outflow of aqueous humor, leading to angle-closure glaucoma [5]. Under normal physiological conditions, trabecular meshwork structure can be maintained by homeostasis of cell proliferation and apoptosis. Various factors [6] including inflammatory response, cytokines or oxidative stress can lead to abnormal proliferation of trabecular meshwork cells [7].
Study showed that nitric oxide (NO) and superoxide anion (O2-) quickly reacted to generate potent oxidant peroxynitrite (ONOO-), which is the major step for pathological injury of NO [8]. NO synthesis under the catalyse of nitric oxide synthase (NOS). Previous study showed expression of NOS and NO in trabecular meshwork cells in normal people, whilst glaucoma patients showed significantly higher NO level [9]. Caveolin is the major membrane protein on cell surface caveolae [10]. As one tumor suppressor gene, CAV mutation has been identified in various cancers [11,12]. Moreover, one sequencing study found existence of CAV mutation in some patients with primary open angle glaucoma (POAG), and such mutations were potentially related with abnormal trabecular meshwork cells [13]. Therefore, we aimed to investigate the expression of eNOS and CAV gene in trabecular meshwork cells and their effects on glaucoma pathogenesis.
Materials and Methods
Research subject
A total of 32 patients (52 eye balls from 20 males and 12 females, aging between 38 and 64 years, average age=49.1 ± 6.5 years) who were admitted as diagnosed as POAG in department of ophthalmology in our hospital were recruited in this study.
Inclusive criteria: Not taking nitrate ester medicine before recruitment; No complication with other ocular or systemic disorder; Peak intraocular pressure higher than 21 mmHg within 24 h before drug application; Opened anterior angle; Presence of unique glaucoma vision field and ocular structural damage. Another 14 normal eyeballs were recruited from healthy donors of cornea transplantation, without glaucoma or other ophthalmology or systemic disease. This study was approved and monitored by the ethical committee of our hospital, and all participants have signed informed consents.
Trabecular meshwork tissues were obtained during the surgery from both control and experimental groups, and were firstly fixed in 1% neutral buffered formalin, followed by paraffin embedding and sectioning into 4 μm slices. HE staining was performed to observe cell morphology.
qRT-PCR
PCR primers were designed based on mRNA sequence of eNOS and CAV1 genes (Genebank access number: NM_003884) for qRT-PCR. Sequences for amplification primers and probes were shown in Table 1. Using β-actin as the endogenous reference, mRNA relative expression of eNOS and CAV1 was measured in all samples. Total RNA was extracted from fresh trabecular meshwork tissues removed during the surgery by RNA rapid extraction kit (QIAGEN). Using total RNA from normal tissues as the control group, RT-PCR was performed using test kit (TianGen). Reverse transcription was firstly performed at 37°C for 2 h, followed by qRT-PCR using cDNA as the template under following conditions: 95°C for 5min, followed by 30 cycles each containing 95°C 1min, 58°C 30 s and 72°C 1 min. Relative expression of eNOS and CAV1 mRNA was calculated.
| Primer | Sequence | |
|---|---|---|
| β-actin | F | 5'-TGTCCCGATGGCGAGTGTTT-3' |
| R | 5'-CCTGTTGGCCATAGTACTGC-3' | |
| β-actin-probe | 5'-FAM-ACCTGACTGAAGTTAGATATGCGCT-TAMRA-3 | |
| eNOS | F | 5'- CCTCACCGCTACAACATCCT -3' |
| R | 5'- TTCACTCGCTTCGCCATCA -3' | |
| eNOS -Probe | 5'-FAM- GCCTTCTGCTCATTCTCCAG-TAMRA-3 | |
| CAV1 | F | 5'-GCATCAGCCGTGTCTATTCC-3' |
| R | 5'-GCAGTTGAGGTTGTTGGTTCT-3' | |
| CAV1-probe | 5'-FAM-TCAGCAATGTCCGCATCAAC-TAMRA-3 |
Table 1. qRT-PCR primer and probe sequences.
Western blot
Trabecular meshwork tissues were homogenized and digested in 100 μL lysis buffer. Protein electrophoresis and Western Blot analysis was performed on supernatant extracted by 13000 g centrifugation for 10min. SDS-PAGE was performed to separate proteins using 15% separating gel and 5% condensing gel. Targeted proteins were transferred to PVDF membrane, which was blocked in 5% defatted milk powder at 37°C for 1h. After TBST rinsing, primary antibody working solution was added (mouse anti-human eNOS at 1:1000, mouse anti-human CAV1 specific antibody at 1:1000, mouse anti-human β-actin at 1:1000, all from Boster Bio, China) for 4°C overnight incubation. Excess primary antibody was removed by TBST, and horseradish peroxidase conjugated rabbit anti-mouse IgG antibody (1:200, Cayman chemical, US) was added for 1h room temperature incubation. After TBST rinsing, freshly prepared DAB substrate was added for 10min development, which was quenched by distilled water. Western Blot results were input into the computer, and integrated gray value was measured by gel imaging system. Using β-actin as the endogenous reference, relative expression level of eNOS and CAV1 was corrected in samples.
Cell culture and CAV1 over-expression
CAV1 gene sequence was amplified by specific primers (CAV1-F: 5’-AGAGG CCTTT GAAAT TGT-3’; CAV1-R: 5’- AAATA CTTTC AGGTC ACGTC-3’). Gene sequence was inserted in pBudCE4.1 mammalian cell over-expression plasmid. Using empty plasmid as the negative control, liposome INTERFERin transfection reagent was applied for cell transfection. Trabecular meshwork cell cline HTMC was purchased from Cell Bank, Chinese Academy of Sciences. Cells were kept in DMEM medium containing 20% fetal bovine serum (FBS), 1 × 106 U/mL, 100 μg/mL streptomycin at 37°C with 5% CO2. Cells were resuscitated and cultured till log-growth phase, digested in trypsin, enumerated and diluted in freshly prepared medium for seeding into 96 well plate. After 24 h incubation, transfection was performed. 1 μL Lipofectamine 2000 (Invitrogen) was diluted into 50 μL antibiotic-/serum- free DMEM medium. After 5 min incubation, empty plasmid pBudCE4.1 or pBudCE4.1 vector with successful CAV1 recombination was added. 100 μL transfection mixture was added into each well, and the plate was incubated at 37°C chamber with 5% CO2. After 6 h, antibiotic-/serum-free medium was then switched for DMEM medium with 10% FBS and gentamycin for 48-72 h incubation and further assays.
eNOS over-expression
eNOS gene amplification primers (eNOS-F: 5’-AGAGG CCVTAT GGGCA ACTTG AAGAG CG-3’; eNOS-R: 5’- AAATA CTTCA GGGGC TGTTG GTGTC TGAG-3’). Gene sequence was inserted into pBudCE4.1 mammalian expression vector, which was then used to transfect HTMC cells. Cell culture and transfection protocols followed those mentioned above.
MTT assay for cell proliferation potency
Those HTMC cells with successful transfection were incubated until log-growth phase, and were digested in trypsin. After centrifugation, the supernatant was discarded, and cells were re-suspended in culture medium. 6 μL Trypan blue solution was mixed with equal volume of cell suspension. After complete mix, 10 μL solution was applied onto cell counting chamber with coverslips, and were enumerated under microscope. Cell suspension was placed into 24-well plate, which was incubated at 37°C chamber with 5% CO2 for 12 h. 70 μL MTT solution was added into each well, and cells were cultured at 37°C chamber with 5% CO2 for 3 h. After mixture, the supernatant was saved and added with DMSO. A570 nm value was measured under a microplate reader.
Assay for cellular oxidative injury
Chemiluminescence assay was applied for measuring free radical level within cells to evaluate oxidative injury. After culture of trabecular cells, cells were collected by centrifugation, and were rinsed in PBS, re-suspended and ruptured. 100 μL cell debris was added with 0.001 M luminol and 600 μL purified water, and were incubated at 37°C for 30 min. 100 μL AAPH was then added. The mixture was placed into chemiluminescent apparatus, and CP6s was measured from reaction tubes using T-2 algorism under 39°C.
Statistical analysis
All statistical analysis was performed on SPSS20.0 software. Data were presented as mean ± standard deviation. Comparison among multiple groups was performed using oneway ANOVA. Difference between two groups was analyzed by student t-test. A statistical analysis was identified when p<0.05.
Experimental Results
HE staining of trabecular meshwork tissues of POAG patients
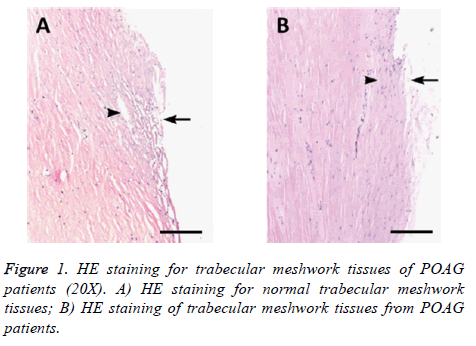
Trabecular meshwork tissues collected from surgery of POAG patients and normal eyeballs were embedded, sectioned and stained under HE approach. Difference of tissue structure and cell morphology was compared between POAG patients and control group (Figure 1). We found normal trabecular meshwork tissues had intact structure under microscope, with evenly distributed endothelial cells, less pigment sedimentation within cells, regular Schlemm tube cavity, clear and even tubular cleft. Trabecular meshwork tissues from POAG patients, however, showed stenosis of tissue cleft, decreased number of endothelial cells, more pigment sedimentation granules, irregular Schlemm tubules, and sometime tubular cavity collapse.
eNOS and CAV1 expression
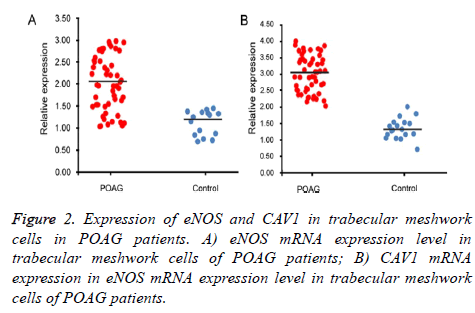
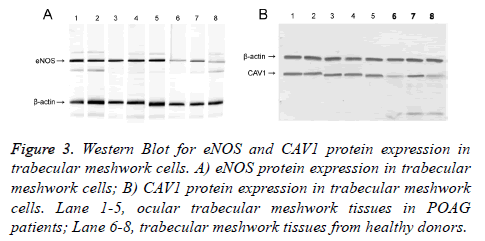
qRT-PCR and Western Blot were used to test expression level of eNOS and CAV1 genes at mRNA and protein levels. Test results were analyzed in gel imaging system to calculate relative expression level of eNOS and CAV1 using β-actin as the reference (Figures 2 and 3). Results showed that relative expression level of eNOS in trabecular meshwork tissues of POAG patients was about 2.01, with significant difference compared to normal group (1.21, p<0.05), indicating significantly higher eNOS expression in POAG trabecular meshwork tissues than normal tissues. As similar results, we found that relative CAV1 expression level was 3.01 in trabecular meshwork tissues of POAG patients, significantly higher than that in normal trabecular meshwork tissues (1.31, p<0.05).
Figure 3: Western Blot for eNOS and CAV1 protein expression in trabecular meshwork cells. A) eNOS protein expression in trabecular meshwork cells; B) CAV1 protein expression in trabecular meshwork cells. Lane 1-5, ocular trabecular meshwork tissues in POAG patients; Lane 6-8, trabecular meshwork tissues from healthy donors.
Over-expression of eNOS and CAV1 in HTMC cells
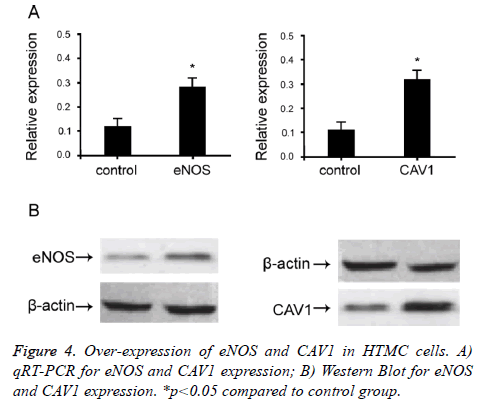
To study the effect of eNOS and CAV1 expression on POAG trabecular meshwork tissues, we over-expressed eNOS and CAV1 in HTMC cells. Mammalian over-expression vector containing eNOS and CAV1 was used to transfect HTMC cells. qRT-PCR and Western Blot were used to confirm cellular expression level of eNOS and CAV1 (Figure 4). When cells were transfected with eNOS or CAV1 over-expression plasmid, relative expression levels of eNOS and CAV1 were significantly up-regulated compared to empty or negative controls (p<0.05). These results indicated successful overexpression, which effectively up-regulate eNOS and CAV1.
Effects of eNOS and CAV1 expression on cell survival
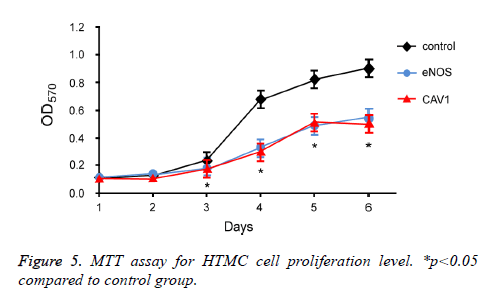
To further investigate the effect of eNOS and CAV1 overexpression on survival of trabecular meshwork cells, MTT assay was performed to measure the proliferation and survival of HTMC cells after eNOS or CAV1 over-expression (Figure 5). 72 h incubation of HTMC cells with eNOS or CAV1 overexpression significantly decreased cell proliferation rate compared to control HTMC cells.
Effects of eNOS and CAV1 expression on clearance of free radicals in cells
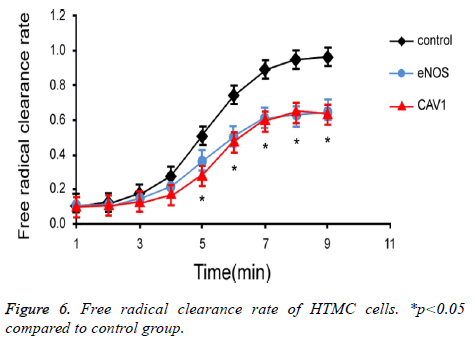
To investigate the effect of eNOS and CAV1 expression on production and clearance of cellular free radicals in HTMC cells, chemiluminescence assay was used to measure the free radical clearance rate of HTMC cells (Figure 6). Results showed significantly lower clearance rate of free radicals in HTMC cells with over-expression of eNOS or CAV1 (p<0.05).
Discussion
Current clinical and pathological study showed that glaucoma was mainly caused by structural or functional change of ocular trabecular meshwork tissues, leading to decreased number of trabecular cells, more pigment deposition, shrinkage of trabecular cleft, further causing drainage blockade of ocular aqueous humor and hyper tension, eventually leading to glaucoma pathogenesis [4]. Investigation on trabecular tissue or cell damage or structural change are thus of critical importance for glaucoma prevention and treatment, and protection of vision. This study separated trabecular meshwork tissues from glaucoma patients, to analyze their structural change, along with expression level of eNOS and CAV1 genes. Consequently molecular study and cell culture demonstrated the effect of eNOS and CAV1 on proliferation and free radical clearance on trabecular meshwork cells.
Glaucoma pathogenesis is correlated with aging. Norma people develop a series of structural and functional change of trabecular meshwork tissues such as decreased cell number of pigment deposition with aging [14]. Transmission electron microscope examined trabecular meshwork tissues of POAG patients and found more potent effects in trabecular tissues of POAG patients [15]. This study revealed up-regulation of eNOS and CAV1 in POAG trabecular tissues, suggesting possible correlation between those expression levels and dysfunction of glaucoma meshwork tissues.
NO is widely distributed in human eyes, and is synthesized under NOS catalyze. NO production within trabecular meshwork can dilate meshwork tissues, relax ciliary muscle, thus draining out aqueous humors. POAG patient’s trabecular meshwork tissues had significantly higher eNOS expression than normal people, and large amounts of eNOS expression leads to over-production of NO inside cells, whilst excess NO synthesis further leads to nuclear injury and other pathological effects [16]. Study showed that NO could quickly react with super oxide anion to produce ONOO-, which is one critical step in NO-induced cell damage [8]. ONOO- is one potent oxidant, and can oxidize sulfur group of cellular proteins, leading to structural damage or inactivation of proteins. ONOO- may also nitrify tyrosine residue of proteins, eventually affecting protein function and cell metabolism [17,18]. By over-expressing eNOS in cells, we also found significantly decreased cell survival rate or free radical clearance rate in cells, as consistent with other groups [17,18].
Caveolin-1 (CAV1) is the major component of caveolae structure of endothelial cell membrane [10]. Previous study showed that CAV1 participated in regulating cell proliferation and growth. This is mainly due to binding of N terminal amino acid sequence of CAV1 protein with multiple critical signal transduction molecules including G protein, EFG receptor, eNOS and PKC, thus modulating activation status of those signal molecules [19]. Currently few studies have been performed on the role of CAV1 in glaucoma pathogenesis. Some groups performed whole genome sequencing and showed the existence of SNP loci within Cav1 gene sequence of certain glaucoma patients. This SNP loci may affect normal function of CAV1 via affecting downstream CAV1transcription [20]. However, this hypothesis still requires further elaboration of experimental data. This study for the first time showed upregulation of CAV1 in trabecular meshwork of POAG patients, although further studies are required to elucidate the correlation between CAV1 up-regulation and glaucoma pathogenesis.
Glaucoma is one of three major blinding disorders in humans, and severely threatens patient’s vision and normal life with aging population [21]. This study revealed expressional change of eNOS and CAV1 in trabecular meshwork in glaucoma patients, and provided clues for further research on glaucoma pathogenesis. This study, however, did not provide sufficient analysis for the reason behind abnormal expression of eNOS and CAV1 genes, thus cannot illustrate expressional regulatory mechanism of eNOS and CAV1 in trabecular meshwork cells. Further studies can be performed via whole genome or transcriptome sequencing to screen out more candidate genes or regulatory factors with abnormal expression patterns in trabecular meshwork tissues from glaucoma patients, thus further elucidating pathogenesis of glaucoma.
Conclusion
Trabecular meshwork cells showed up-regulation of eNOS and CAV1 expression compared to normal trabecular meshwork tissues. Up-regulation of eNOS may suppress cell survival rate, impair cellular potency of free radical clearance, eventually leading to cell and tissue damage.
References
- Kim CS, Seong GJ, Lee NH, Song KC; Namil Study Group, Korean Glaucoma Society. Prevalence of primary open-angle glaucoma in central South Korea the Namil study. Ophthalmology 2011; 118: 1024-1030.
- Weinreb RN, Aung T, Medeiros FA. The pathophysiology and treatment of glaucoma: a review. JAMA 2014; 311: 1901-1911.
- Krupin T, Liebmann JM, Greenfield DS, Ritch R, and Gardiner S, A randomized trial of brimonidine versus timolol in preserving visual function: results from the Low-Pressure Glaucoma Treatment Study. Am J Ophthalmol 2011; 151: 671-681.
- Ren R, Jonas JB, Tian G, Zhen Y, Ma K, Li S, Wang H, Li B, Zhang X, and Wang N, Cerebrospinal fluid pressure in glaucoma: a prospective study. Ophthalmology 2010; 117: 259-66.
- Izzotti A, Saccà SC, Longobardi M, Cartiglia C. Mitochondrial damage in the trabecular meshwork of patients with glaucoma. Arch Ophthalmol 2010; 128: 724-730.
- Last JA, Pan T, Ding Y, Reilly CM, Keller K, Acott TS, Fautsch MP, Murphy CJ, and Russell P, Elastic modulus determination of normal and glaucomatous human trabecular meshwork. Invest Ophthalmol Vis Sci 2011; 52: 2147-2152.
- Li G, Luna C, Qiu J, Epstein DL, and Gonzalez P, Modulation of inflammatory markers by miR-146a during replicative senescence in trabecular meshwork cells. Invest Ophthalmol Vis Sci 2010; 51: 2976-2985.
- Pall ML. Is Open-angle Glaucoma Caused by the NO/ONOO(-) Cycle Acting at Two Locations in the Eye? Med Hypothesis Discov Innov Ophthalmol 2014; 3: 1-2.
- Wu RY, Ma N. Expression of nitric oxide synthase and guanylate cyclase in the human ciliary body and trabecular meshwork. Chin Med J (Engl) 2012; 125: 129-133.
- Pavlides S, Gutierrez-Pajares JL, Danilo C, Lisanti MP, Frank PG. Atherosclerosis, caveolae and caveolin-1. Adv Exp Med Biol 2012; 729: 127-144.
- Syeed N, Hussain F, Husain SA, and Siddiqi MA, 5'-CpG island promoter hypermethylation of the CAV-1 gene in breast cancer patients of Kashmir. Asian Pac J Cancer Prev 2012; 13: 371-375.
- Thompson TC, Tahir SA, Li L, Watanabe M, Naruishi K, Yang G, Kadmon D, Logothetis CJ, Troncoso P, Ren C, Goltsov A, and Park S, The role of caveolin-1 in prostate cancer: clinical implications. Prostate Cancer Prostatic Dis 2010; 13: 6-11.
- Thorleifsson G, Walters GB, Hewitt AW, Masson G, Helgason A, et al.. Common variants near CAV1 and CAV2 are associated with primary open-angle glaucoma. Nat Genet 2010; 42: 906-909.
- Izzotti A, Longobardi M, Cartiglia C, and Sacca SC, Mitochondrial damage in the trabecular meshwork occurs only in primary open-angle glaucoma and in pseudoexfoliative glaucoma. PLoS One 2011; 6: e14567.
- Sihota R, Goyal A, Kaur J, Gupta V, and Nag TC, Scanning electron microscopy of the trabecular meshwork: understanding the pathogenesis of primary angle closure glaucoma. Indian J Ophthalmol 2012; 60: 183-188.
- Li S, Wang W, Wang C, and Tang YY, Possible involvement of NO/NOS signaling in hippocampal amyloid-beta production induced by transient focal cerebral ischemia in aged rats. Neurosci Lett 2010; 470: 106-110.
- Konieczka K, Fränkl S, Todorova MG, Henrich PB. Unstable oxygen supply and glaucoma. Klin Monbl Augenheilkd 2014; 231: 121-126.
- Ayub H, Khan MI, Micheal S, Akhtar F, Ajmal M. Association of eNOS and HSP70 gene polymorphisms with glaucoma in Pakistani cohorts. Mol Vis 2010; 16: 18-25.
- Baig SM, Koschak A, Lieb A, Gebhart M, Dafinger C, Nurnberg G, Ali A, Ahmad I, Sinnegger-Brauns MJ, Brandt N, Engel J, Mangoni ME, Farooq M, Khan HU, Nurnberg P, Striessnig J, and Bolz HJ, Loss of Ca(v)1.3 (CACNA1D) function in a human channelopathy with bradycardia and congenital deafness. Nat Neurosci 2011; 14: 77-84.
- Loomis SJ, Kang JH, Weinreb RN, Yaspan BL, Cooke Bailey JN. Association of CAV1/CAV2 genomic variants with primary open-angle glaucoma overall and by gender and pattern of visual field loss. Ophthalmology 2014; 121: 508-516.
- Caprioli J, Coleman AL; Blood Flow in Glaucoma Discussion. Blood pressure, perfusion pressure, and glaucoma. Am J Ophthalmol 2010; 149: 704-712.