Research Article - Biomedical Research (2018) Volume 29, Issue 6
Association of single nucleotide polymorphisms in estrogen receptor 1 gene with the risk of idiopathic short stature
Yu Yang1*, Hui Huang1, Yi Yuan1, Wei Wang2, Li Yang1, Liling Xie1, Ying Wang31Department of Endocrinology, Genetics, and Metabolism, Jiangxi Provincial Children’s Hospital, Nanchang 330006, Jiangxi, PR China
2Department of Pediatrics, Ruijin Hospital, Shanghai Jiaotong University School of Medicine, Shanghai 201209, PR China
3Key Laboratory of Health and Disease Genomics, Chinese National Human Genome Center at Shanghai, Shanghai 201209, PR China
- *Corresponding Author:
- Yu Yang
Department of Endocrinology
Genetics and Metabolism
Jiangxi Provincial Children’s Hospital, PR China
Accepted date: December 20, 2017
DOI: 10.4066/biomedicalresearch.29-17-2526
Visit for more related articles at Biomedical ResearchAbstract
Estrogen is a hormone in hypothalamic-pituitary-gonadal (HPG) axis that plays an important role in the regulation of bone maturation and closure of the growth plate. The aim of this study was to investigate the association between single nucleotide polymorphisms (SNPs) in estrogen receptor 1 (ESR1) gene and susceptibility to idiopathic short stature (ISS) in the Chinese population. Six SNPs of the ESR1 gene (rs6557177, rs1884049, rs3020429, rs3798575, rs3778090, rs3798757) were genotyped and we found a significant positive association between rs6557177 genotype and susceptibility to ISS compare to control group in preliminary study (ISS=200, control=220, χ2=6.262, P =0.044). Validation of rs6557177 was conducted by further expanding the sample size to 400 ISS patients and 380 control subjects genotyped by SNaPshot Multiplex System. The frequencies of the rs6557177 CC genotype and C allele were observed to increased in ISS group compared with the control group (χ2=9.913, P =0.007; χ2=5.57, P = 0.018 OR: 1.502; 95% CI: 1.07-2.108 respectively). However, no significant correlation was observed between the rs6557177 and clinical characteristics of the study participants. We demonstrate for the first time that the presence of a C allele or CC genotype at rs6557177 of ESR1 constitutes risk factor for developing ISS in Chinese children.
Keywords
Estrogen receptor 1, Idiopathic short stature, Single nucleotide polymorphisms.
Introduction
Short stature is one of the most popular endocrine disease presenting to pediatric endocrinologists. The prevalence of the short stature around the world is estimated to be 3-5% [1]. The characterization of idiopathic short stature (ISS) is in which the height of a patient is less than -2SDS below the equivalent mean height for the same age, sex, and population group without knowing diseases, and almost 60-80% of short stature fitting the classification of ISS [2]. A typical human linear growth has four major stages: fetal, infancy, childhood, and pubertal growth. Previous reports have shown that hypothalamic-pituitary-gonadal (HPG) axis and growth hormone-Insulin-like growth factor (IGF) axis is the most vital hormonal axis governing growth [3]. Any level of abnormality of GH-IGF axis may cause short stature [4], such as GHR [5], IGF-1R [6,7], IGFALS [8] were associated with ISS, but fewer studies have investigated the association of HPG axis with the risk of ISS.
Estrogens are important endocrine regulators both involved in GH-IGF axis, and gonadotropic axis [9]. Many studies had implied that ESR1 plays a central role of estrogen actions in bone, such as skeletal growth in the growth spurt, epiphyseal fusion during puberty [10-12]. Genome-wide linkage analysis of large samples has shown strong evidence of the association of ESR1 gene with stature [13]. A mutation of the ESR1 gene was previously described as a woman who showed growth retardation with no pubertal growth spurt and breast development [14]. However, there is no study has investigated the correlation of the ESR1 gene with the risk of ISS.
Therefore, we designed a case-control study consisting of 400 children with ISS and 380 normal children and analyzed the SNPs of ESR1 gene to determine the correlation of gene alleles and frequencies, and analyzed the clinical characteristics of these patients. Our study findings would pave the way for future research on the genes responsible for increased susceptibility to ISS.
Materials and Methods
Subjects
The case-control study design was adopted in this experiment. A total of 780 samples were recruited from unrelated individuals of the southern Chinese Han population. This study was approved by the Ethics Committee of Jiangxi Children’s Hospital and written informed consent was obtained from all participants’ guardians.
ISS Group
A total of four hundred children who were clinically diagnosed with ISS between 2008 and 2016 at Jiangxi Provincial Children’s Hospital agreed to participate in this study. The mean age of case group was 9.52 ± 2.20 years. None of the children had yet entered puberty, and the heights of their parents were all within the normal range. Familial short stature and physical, developmental delay of adolescence were excluded for all subjects.
The inclusion criteria [15] were as follows: i) height less than 2 SD of the mean height of children of the same age and sex [16], ii) at least one GH peak value >10 ng/mL in the GH provocation test, iii) normal routine blood, thyroid function, liver, and kidney function, iv) birth weight and length within the normal range, and v) no other inherited metabolic diseases, chromosomal abnormalities, congenital skeletal abnormalities, or chronic diseases.
Control group
Three hundred and eighty normal children who came for regular physical examinations in our hospital during the same period were selected as control subjects. The mean age of control was 9.3 ± 2.04 years. The inclusion criteria were as follows: (i) height within ± 2 SDs of the mean height of the normal children of the same sex and (ii) BMI within the normal range. Children whose heights were between ± 2 SD of the height of other children of the same sex, age and those whose body mass index (BMI) was within the range of 17-23 were included in this group.
Genomic DNA extraction
Two milliliters fasting venous blood was drawn from the ISS and control group subjects, EDTA-2K anti-coagulated, and genomic DNA was extracted according to the routine phenolchloroform- isoamyl alcohol method, which was stored at -20°C until use.
SNP selection and genotyping
The gene covers 295.7 kb of genomic sequence on chromosome 6. The practical SNP data was from the International HapMap Project (www.hapmap.org) and Tagger program to select a subset of tag SNPs based on linkage disequilibrium (LD) patterns (r2>0.8) across gene region, and used the HapMap CHB subset for our tag SNPs selection (the minor allele frequency>0.05) [17]. Six tag SNPs (rs6557177, rs1884049, rs3020429, rs3798575, rs3778090, rs3798757) were dressing by screening and ordered for genetic variations in the . Genotyping of gene was performed by using the SNaPshot Multiplex System (ABI 3730 sequencer, Applied Biosystems, Forster City, CA, USA). After designing primers of different lengths, the SNaPshot reaction was performed targeting different mutational sites.
The SNaPshot polymerase chain reaction (PCR) cycle conditions were: 96°C for 10 s, followed by 25 cycles of 96°C for 10 s, 50°C (53°C) for 5 s, and 60°C for 30 s, and finally 60°C for 30 s and reducing to 4°C. The total volume of PCR amplification was 5 μL containing 1.3 μL dH20, 1.2 μL Pooled Primers, 2 μL Pooled PCR Products and 0.5 μL Reaction Mix. A pair of primers were designed (forward: 5’- actcagatattcgtgtcctt-3’ and reverse: 5’-ttggatacaaccttgaatgc-3’) to amplify a 251 bp fragment from ESR1 that contained rs6557177. The sequencing result of rs6557177 locus was analyzed using GeneMapper Software Version 4.0 (Applied Biosystems, Forster City, CA, USA).
Statistical analysis
Height, IGF-1, IGF-binding protein 3 (BP3) [18] and E2 were measured, and genetic height was used as the reference unit to calculate the SDS. The SDS of IGF-1=(actual IGF-1-the means IGF-1 at the same age)/standard deviation of IGF-1 at the same age.
The differences in alleles, genotypes and dominant and recessive modes were assessed by the chi-square test between ISS patients and controls with the odds ratio (OR) and the 95% confidence interval (95%). The Student t-test or the rank sum test was adopted to analyze the association between genotype and clinical parameters. Statistically, significant differences were indicated when P<0.05. All analyses were performed with SPSS17.0.
Results
Hardy-Weinberg equilibrium and chi-squared test in Preliminary experiment
Genotype distributions of these six ESR1 polymorphisms were in Hardy-Weinberg equilibrium in both patients (n=200) and controls (n=220) (P>0.05), Analysis at the screening stage of the 200 ISS patients and 220 control subjects revealed that rs6557177 was significantly correlated to ISS (χ2=6.262, P=0.044) showed in Table 1. Targeting this locus, validation was conducted by further expanding the sample size to 400 ISS patients and 380 control subjects. The validation revealed no statistical significance in the differences in the distribution frequencies of the alleles, the distribution frequencies of genotypes, or case-control associations of genotype in the dominant and recessive modes.
| SNPs ID | ISS (n=200) | Control (n=220) | OR (95%CI) | P value | Genotype | ISS (n=200) | Control (n=220) | P value |
|---|---|---|---|---|---|---|---|---|
| rs6557177 | CC | 8 | 2 | 0.044* | ||||
| C | 64 | 46 | 1 | 0.017* | CT | 48 | 42 | |
| T | 336 | 394 | 1.631 (1.087-2.448) | TT | 144 | 176 | ||
| rs1884049 | CC | 5 | 5 | 0.195 | ||||
| C | 65 | 55 | 1 | 0.121 | CT | 55 | 45 | |
| T | 335 | 385 | 1.358 (0.922-2.002) | TT | 140 | 170 | ||
| rs3020429 | CC | 4 | 1 | 0.388 | ||||
| C | 40 | 38 | 1 | 0.496 | CT | 32 | 36 | |
| T | 360 | 402 | 1.175 (0.737-1.874) | TT | 164 | 183 | ||
| rs3798575 | GG | 6 | 12 | 0.194 | ||||
| G | 78 | 109 | 1 | 0.067 | GA | 66 | 85 | |
| A | 322 | 331 | 0.736 (0.53-1.022) | AA | 128 | 123 | ||
| rs3798757 | GG | 3 | 6 | 0.198 | ||||
| G | 64 | 91 | 1 | 0.081 | GA | 58 | 79 | |
| A | 336 | 351 | 0.731 (0.513-1.040) | AA | 139 | 136 | ||
| rs3778090 | CC | 31 | 33 | 0.812 | ||||
| C | 158 | 166 | 1 | 0.598 | CT | 96 | 100 | |
| T | 242 | 274 | 1.078 (0.816-1.423) | TT | 73 | 87 |
Table 1: Genotypic and allelic association analyses of ESR1 SNPs in Preliminary experiment.
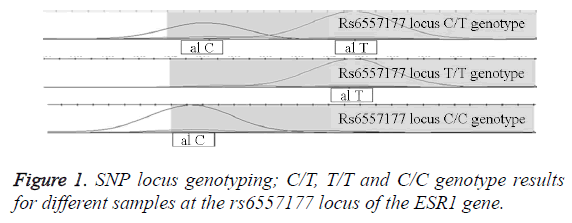
SNP locus genotyping
The SNP sequencing results at the rs6557177 locus of the ESR1 gene were analyzed using GeneMapper4.0 and are shown in Figure 1.
Association study of rs6557177 with ISS
The results of the rs6557177 analysis showed a statistically significant difference between ISS and control groups in the C/T allele distribution frequencies (χ2=5.570, P=0.018, OR=1.502, 95% CI=1.070-2.108) and the genotype distribution frequencies also showed significantly difference between ISS and control groups (χ2=9.913, P=0.007), whereas the C recessive mode did show statistical significance (χ2=9.888, P=0.002, OR=1.585, 95% CI=1.189-2.114). After Bonferroni correction for multiple tests, the difference in C/T alleles, genotype and recessive mode in ISS patients remained statistically significant, which suggested that the C allele, genotype and the C recessive mode of rs6557177 were relevant to susceptibility to ISS (Table 2).
| SNP | ISS (n=400) | Control (n=380) | χ2 | P | OR | 95% CI | Boferroni | ||
|---|---|---|---|---|---|---|---|---|---|
| N | % | N | % | ||||||
| CC | 260 | 65 | 205 | 54 | 9.913 | 0.007* | 0.013* | ||
| CT | 124 | 31 | 156 | 41 | |||||
| TT | 16 | 4 | 19 | 5 | |||||
| Allele | 648 | 81 | 562 | 74 | 5.570 | 0.018* | 1.502 | 1.00 | 0.023* |
| C | 12 | 19 | 198 | 26 | (1.070-2.108) | ||||
| T | |||||||||
| Dominant | 384 | 96 | 361 | 95 | 0.730 | 0.393 | 1.351 | 1.00 | 0.497 |
| CC+CT | 16 | 4 | 19 | 5 | 9.888 | 0.002* | 1.585 | (0.676-2.699) | 0.012* |
| TT | 260 | 65 | 205 | 54 | 1.00 | ||||
| Recessive | 140 | 35 | 175 | 46 | (1.189-2.114) | ||||
| CC | |||||||||
| CT+TT | |||||||||
Table 2: Chi-square test for rs6557177 locus of the ESR1 gene.
Comparison of clinical characteristics under rs6557177 C Recessive mode
The clinical characteristics of the participants, including height, body weight, BMI, and levels of IGF-1, IGFBP3 were summarized (Table 3). The differences in HtSDS, BMI, IGF-1 SDS, IGF-BP3 SDS, and TH SDS of the CC genotype compared to (CT+TT) genotype at loci rs1976667 of ISS were not statistically significant (P>0.05) (Table 4).
| Clinical parameter | ISS | Control |
|---|---|---|
| gender | ||
| male | 208 | 200 |
| female | 192 | 180 |
| Age (years) | 9.52 ± 2.20 | 9.3 ± 2.04 |
| HtSDS | -3.15 ± 1.01 | 1.20 ± 1.03 |
| weightSDS | -1.62 ± 1.33 | 1.33 ± 1.47 |
| BMI (kg/m2) | 15.77 ± 4.24 | 20.73 ± 4.24 |
| IGF-1SDS | -1.48 ± 1.65 | 1.22 ± 1.70 |
| IGFBP3SDS | -1.08 ± 1.60 | 1.25 ± 1.17 |
| TH (cm) | 163.78 ± 5.77 | 172.86 ± 4.16 |
| THSDS | -1.37 ± 1.03 | 1.82 ± 2.11 |
| E2 (pg/ml) | 15.79 ± 4.26 | 16.79 ± 2.28 |
Table 3: Descriptive clinical and laboratory data of studied cases and controls.
| CC(n=400) | CT+TT(n=380) | F/t-value | P | |
|---|---|---|---|---|
| HtSDS | -3.22 ± 1.182 | -3.11 ± 0.987 | -1.012 | 0.312 |
| BMI | 15.300 ± 2.648 | 15.54 ± 2.343 | -0.729 | 0.466 |
| IGF-1 SDS | -1.58 ± 1.808 | -1.53 ± 1.520 | 1.148 | 0.252 |
| IGF-BP3 SDS | -1.12 ± 1.591 | -0.98 ± 1.532 | -1.264 | 0.207 |
| TH SDS | -2.27 ± 4.414 | -1.42 ± 0.938 | -1.839 | 0.067 |
| E2(pg/ml) | 20.38 ± 16.3 | 23.20 ± 17.491 | -1.181 | 0.248 |
Table 4: Correlation analysis of Clinical Characteristics Under rs6557177 C Recessive mode (CC/CT+TT).
Discussion
The heritability of height is estimated to be 80% [13,19], althrough 180 loci were confirmed by recent GWAS study, only explain for approximately 10% of the stature variation [20]. According to large longitudinal cohorts of height studies results, SNPs plays a critical role in childhood growth [21] which implies the complicated genetics of short stature. ISS is thought to be a polygenic disease, both genetic and environmental factors contribute to its pathogenesis, but genetic factors play a more significant part in the development of ISS [22]. Previous reports have shown that mutations in the genes along the GH-IGF axis and genes that regulate the development of the cartilage growth plate are involved in the development of ISS [4,23,24]. At present, studies are focused on the GH-IGF-1 axis genes, but from GWAS results, most genes and biological pathways associated with height were located in growth plate [21], such as BMP2, SOX8, WNT4, et al. To reduce the expensive burden of growth hormone therapy, it’s profitable to reveal the etiology of ISS.
The ESR1 gene is located at position 6q25.1, which includes eight exons, seven introns. Obviously, Estrogen take a vital part in the HPG axis in the regulation of bone maturation and closure of the growth plate [11], and Estrogen signaling participated in longitudinal bone growth and for bone remodeling via ERα. Genome-wide linkage analysis of large samples has shown strong evidence of the association of ESR1 gene with stature [13]. Schuit et al. [25] studied the polymorphisms in the ESR1 gene and reported that ERα affected adult height in normal individuals. Dahlgren et al. [26] reported that rs2179922 which was in intron 4 of ESR1 showed an association with male stature in two Swedish population cohorts. Previous reports have also revealed that SNPs of ESR1 could influence the onset of menstruation in girls, and body weight, fat accumulation, and BMI in children and adolescents [27,28]. Furthermore, ESR1 polymorphism is associated with osteoporosis [29]. Kang et al. [30] showed that in boys, ESR1 engaged in constitutional delay of growth and puberty. And Hero et al. [31] observed that a two-year long treatment that used a potent aromatase in boys with ISS has significantly improved the predicted adult height.
The absence of ESR1 in mice caused insulin resistance and glucose intolerance in both gender. Multicystic ovaries without corpora lutea, hemorrhagic and hypoplastic uteri were presented in a female ESR1 knockout mice; however, the male ESR1 knockout mice showed obesity and reduced fertility [32]. Until now, there were only two individuals were reported ESR1 mutations. The first was a woman who showed growth retardation, with no pubertal growth spurt, no breast development, and no insulin resistance [14]. In contrast, the other case, who was a man with ESR1 mutation, showed average height, with continued linear growth until adulthood, osteoporosis, delayed epiphyseal closure, and insulin resistance [33].
The frequencies of the rs6557177 CC genotype and C allele were observed to increased in ISS group compared with the control group (χ2=9.913, P =0.007; χ2=5.57, P = 0.018 OR: 1.502; 95% CI: 1.07-2.108 respectively). Carrying the CC genotype significantly influenced the genetic susceptibility to ISS (P=0.018, OR=1.502, 95% CI 1.070-2.108), which was most likely a moderate risk factor (1<OR<2), and presented a C recessive genetic mode of inheritance. These results indicated that ISS carrying the CC genotype at the rs6557177 locus were probably more likely to develop ISS. However, no significant correlation was observed between the rs6557177 and clinical features of the study participants, other clinical features related to ESR1 gene maybe need to confirm.
The rs6557177 locus is in the intron of the ESR1 gene. There is no study has investigated the structure and function of the encoded protein at this locus to date. Our study found that the rs6557177 locus of the ESR1 gene might be relevant to the genetic susceptibility to ISS. Currently, there are no studies on the encoded proteins at the rs6557177, and thus further study on the relationship between structure and function is required to clarify the association of the rs6557177 with genetic susceptibility to ISS.
In conclusion, our study revealed that polymorphism of rs6557177 in ESR1 gene might be linked with ISS risk and provided new insights into the mechanism underlying the pathogenesis of ISS. Moreover, our data would serve as a theoretical basis for developing novel methods for treatment of ISS based on gene mutations by regulating the transcription level of ESR1 gene.
Acknowledgement
Project supported by National Natural Science Foundation of China (Grant No. 81460501).
References
- Argente J. Challenges in the management of short stature. Horm Res Paediatr 2016; 85: 2-10.
- Cohen P, Rogol AD, Deal CL, Saenger P, Reiter EO, Ross JL, Chernausek SD, Savage MO, Wit JM. Consensus statement on the diagnosis and treatment of children with idiopathic short stature: a summary of the Growth Hormone Research Society, the Lawson Wilkins Pediatric Endocrine Society, and the European Society for Paediatric Endocrinology Workshop. J Clin Endocrinol Metab 2008; 93: 4210-4217.
- Murray PG, Clayton PE. Endocrine control of growth. Am J Med Genet C Semin Med Genet 2013; 163: 76-85.
- Waldman LA, Chia DJ. Towards identification of molecular mechanisms of short stature. Int J Pediatr Endocrinol 2013; 2013: 19.
- Pagani S, Petkovic V, Messini B, Meazza C, Bozzola E, Mullis PE, Bozzola M. Heterozygous GHR gene mutation in a child with idiopathic short stature. J Pediatr Endocrinol Metab 2014; 27: 329-334.
- de Graaff LC, Clark AJ, Tauber M, Ranke MB, Johnston LB, Caliebe J, Molinas C, Amin N, van Duijn C, Wollmann H, Wallaschofski H, Savage MO, Hokken-Koelega AC. Association analysis of ten candidate genes in a large multinational cohort of small for gestational age children and children with idiopathic short stature (NESTEGG study). Horm Res Paediatr 2013; 80: 466-476.
- Yang Y, Huang H, Wang W, Yang L, Xie LL, Huang W. Association of insulin growth factor-1 receptor gene polymorphisms with genetic susceptibility to idiopathic short stature. Genet Mol Res 2013; 12: 4768-4779.
- Domené HM, Scaglia PA, Martínez AS, Keselman AC, Karabatas LM, Pipman VR, Bengolea SV, Guida MC, Ropelato MG, Ballerini MG, Lescano EM, Blanco MA, Heinrich JJ, Rey RA, Jasper HG. Heterozygous IGFALS gene variants in idiopathic short stature and normal children: impact on height and the IGF system. Horm Res Paediatr 2013; 80: 413-423.
- Liu Z, Mohan S, Yakar S. Does the GH/IGF-1 axis contribute to skeletal sexual dimorphism? Evidence from mouse studies. Growth Horm IGF Res 2016; 27: 7-17.
- Chagin AS, Sävendahl L. Oestrogen receptors and linear bone growth. Acta Paediatr 2007; 96: 1275-1279.
- Börjesson AE, Lagerquist MK, Windahl SH, Ohlsson C. The role of estrogen receptor alpha in the regulation of bone and growth plate cartilage. Cell Mol Life Sci 2013; 70: 4023-4037.
- Shim KS. Pubertal growth and epiphyseal fusion. Ann Pediatr Endocrinol Metab 2015; 20: 8-12.
- Hirschhorn JN, Lindgren CM, Daly MJ, Kirby A, Schaffner SF, Burtt NP, Altshuler D, Parker A, Rioux JD, Platko J, Gaudet D, Hudson TJ, Groop LC, Lander ES. Genomewide linkage analysis of stature in multiple populations reveals several regions with evidence of linkage to adult height. Am J Hum Genet 2001; 69: 106-116.
- Quaynor SD, Stradtman EW Jr, Kim HG, Shen Y, Chorich LP, Schreihofer DA, Layman LC. Delayed puberty and estrogen resistance in a woman with estrogen receptor alpha variant. N Engl J Med 2013; 369: 164-171.
- Wit JM, Clayton PE, Rogol AD, Savage MO, Saenger PH, Cohen P. Idiopathic short stature: definition, epidemiology, and diagnostic evaluation. Growth Horm IGF Res 2008; 18: 89-110.
- Li H, Ji CY, Zong XN, Zhang YQ. Height and weight standardized growth charts for Chinese children and adolescents aged 0 to 18 years. Zhonghua Er Ke Za Zhi 2009; 47: 487-492.
- Gabriel SB, Schaffner SF, Nguyen H, Moore JM, Roy J, Blumenstiel B, Higgins J, DeFelice M, Lochner A, Faggart M, Liu-Cordero SN, Rotimi C, Adeyemo A, Cooper R, Ward R, Lander ES, Daly MJ, Altshuler D. The structure of haplotype blocks in the human genome. Science 2002; 296: 2225-2229.
- Xu S, Gu X, Pan H, Zhu H, Gong F, Li Y, Xing Y. Reference ranges for serum IGF-1 and IGFBP-3 levels in Chinese children during childhood and adolescence. Endocr J 2010; 57: 221-228.
- Silventoinen K, Sammalisto S, Perola M, Boomsma DI, Cornes BK, Davis C, Dunkel L, De Lange M, Harris JR, Hjelmborg JV, Luciano M, Martin NG, Mortensen J, Nisticò L, Pedersen NL, Skytthe A, Spector TD, Stazi MA, Willemsen G, Kaprio J. Heritability of adult body height: a comparative study of twin cohorts in eight countries. Twin Res 2003; 6: 399-408.
- Lango Allen H, Estrada K, Lettre G, Berndt S., Weedon MN, Rivadeneira F, Willer CJ, Jackson AU, Vedantam S, Raychaudhuri S. Hundreds of variants clustered in genomic loci and biological pathways affect human height. Nature 2010; 467: 832-838.
- Wood AR, Esko T, Yang J, Vedantam S, Pers TH, Gustafsson S, Chu AY, Estrada K, Luan J, Kutalik Z. Defining the role of common variation in the genomic and biological architecture of adult human height. Nat Genet 2014; 46: 1173-1186.
- Wit JM. Definition and subcategorization of idiopathic short stature: between consensus and controversy. Horm Res Paediatr 2011; 76: 3-6.
- Lettre G. Using height association studies to gain insights into human idiopathic short and syndromic stature phenotypes. Pediatr Nephrol 2013; 28: 557-562.
- Wit JM, Oostdijk W, Losekoot M, van Duyvenvoorde HA, Ruivenkamp CA, Kant SG. Mechanisms in endocrinology: Novel genetic causes of short stature. Eur J Endocrinol 2016; 174: R145-173.
- Schuit SC, van Meurs JB, Bergink AP, van der Klift M, Fang Y, Leusink G, Hofman A, van Leeuwen JP, Uitterlinden AG, Pols HA. Height in pre- and postmenopausal women is influenced by estrogen receptor alpha gene polymorphisms. J Clin Endocrinol Metab 2004; 89: 303-309.
- Dahlgren A, Lundmark P, Axelsson T, Lind L, Syvänen AC. Association of the estrogen receptor 1 (ESR1 ) gene with body height in adult males from two Swedish population cohorts. PLoS One 2008; 3: e1807.
- Kulik-Rechberger B, Skorupski P, Bogusiewicz M, Miotła P, Rechberger T. Height at menarche is influenced by estrogen receptor alpha gene polymorphisms. J Endocrinol Invest 2010; 33: 332-338.
- Voorhoeve PG, van Mechelen W, Uitterlinden AG, Delemarre-van de Waal HA, Lamberts SW. Estrogen receptor-alpha gene polymorphisms and body composition in children and adolescents. Horm Res Paediatr 2011; 76: 86-92.
- Wajanavisit W, Suppachokmongkorn S, Woratanarat P, Ongphiphadhanakul B, Tawonsawatruk T. The Association of Bone Mineral Density and G2014A Polymorphism in the Estrogen Receptor Alpha Gene in Osteoporotic Hip Fracture in Thai Population. J Med Assoc Thai 2015; 98: S82-S87.
- Kang BH, Kim SY, Park MS, Yoon KL, Shim KS. Estrogen receptor alpha polymorphism in boys with constitutional delay of growth and puberty. Ann Pediatr Endocrinol Metab 2013; 18: 71-75.
- Hero M, Norjavaara E, Dunkel L. Inhibition of estrogen biosynthesis with a potent aromatase inhibitor increases predicted adult height in boys with idiopathic short stature: a randomized controlled trial. J Clin Endocrinol Metab 2005; 90: 6396-6402.
- Heine PA, Taylor JA, Iwamoto GA, Lubahn DB, Cooke PS. Increased adipose tissue in male and female estrogen receptor-alpha knockout mice. Proc Natl Acad Sci U S A 2000; 97: 12729-12734.
- Bulun SE. Aromatase and estrogen receptor alpha deficiency. Fertil Steril 2014; 101: 323-329.