Review Article - Current Trends in Cardiology (2021) Volume 5, Issue 1
Applications of CRISPR/Cas9 genome-editing technology in cardiac research
Haoran Liu1, Yao Fu2, Gui-Lan Chen3*
1Graduate School of Peking Union Medical College, Chinese Academy of Medical Sciences, Beijing, China
2Department of Rehabilitation, West China Second Hospital, Sichuan University, Chengdu, Sichuan, China
3Key Laboratory of Medical Electrophysiology, Ministry of Education & Medical Electrophysiological Key Laboratory of Sichuan Province, Institute of Cardiovascular Research, Southwest Medical University, Luzhou, Sichuan, China
- Corresponding Author:
- Gui-Lan Chen
Institute of Cardiac Research
Southwest Medical University Luzhou
Sichuan
China
E-mail: chenguilan@swmu.edu.cn
Accepted date: December 23, 2020
Citation: Liu H, Fu Y, Chen GL. Applications of CRISPR/Cas9 genome-editing technology in cardiac research. Curr Trend Cardiol 2021;5(1):5-10.
Abstract
Cardiovascular diseases, especially those affect the heart, are main causes of the mortality of humans. Genetic variations play a crucial role in the pathogenesis of cardiac diseases, although acquired factors such as lifestyle habits and environmental factors are also involved. CRISPR/Cas9 system is a novel genome-editing tool which is more efficient and cost-effective than other gene-editing technologies. In recent years, this technology has been adopted in cardiac research such as generation of disease models, identification of novel disease-causing genes and exploration of gene therapies. There still remain obstacles in further application of CRISPR/Cas9 system, like the potential risk of off-target effect and ethical issues. In this review, we summarized the progress and prospect of the application of CRISPR/Cas9 system in experimental and preclinical studies of cardiac diseases.
Keywords
CRISPR, Cas9, Genome-editing, Cardiac disease, Gene therapy.
Introduction
Cardiovascular disease are the leading causes of mortality for patients across the world. According to data from WHO, approximately 17.5 million people died from per year, which is responsible for almost 35% overall deaths worldwide [1]. In America, treatment for costs more than 273 billion dollars each year; however, the incidence of is still rising [2]. Exploration of the pathogenic mechanisms and novel treatments of by scientists have never been stopped. It has been established that genetic susceptibility is one of the most important factors in the pathogenesis of in particular cardiac diseases. Apart from those genes with well-known functions in cardiac system, identification of new genes related to cardiac diseases may offer novel therapeutic method and hence improve the prognosis [3-5].
Gene-editing technology is a promising method to investigate mechanisms of diseases. By utilizing this technology, researchers can introduce or remove mutations in specific sites of the genome to make disease models or avoid the occurrence of diseases. Since 1996, zinc-finger nucleases (ZFNs) and transcription activator-like effector nucleases (TALENs) emerged separately and served as the first and second generation of gene-editing technology, respectively. However, the high cost, low efficiency and limited accessibility have hindered their applications [6]. In 2013, CRISPR/Cas9 system emerged as the third generation of gene-editing technology and has been rapidly adopted by scientists for its advantages such as high efficiency and cost-effectiveness [7]. By far, this novel technology has been applied in several fields such as agriculture, drug discovery as well as medical research [8,9]. In this review, we retrospect the discovery and characteristics of the CRISPR/Cas9 system, and summarize the progress in the application of CRISPR/Cas9 genome-editing technology in cardiac research.
The discovery and characteristics of CRISPR/ Cas9 system
CRISPR/Cas system is the adaptive immune system of bacteria and archaea which offers resistance to invaders such as viruses [10]. More than half of bacteria and 90% of archaea possess this system [11]. The immune mechanism relies on the guidance of RNAs transcribed from sequences named CRISPR that contains special information of invading pathogens and the participation of CRISPR-associated (cas) proteins to generate site-specific DNA cleavage in targeted cells [12].
CRISPR means the clustered regularly interspersed short palindromic repeats, which was initially reported in 1987 by a group of Japanese scientists when they were searching for the nucleotide sequence of isozyme-converting alkaline phosphatase (iap) gene in Escherichia coli [13]. For lacking detailed information about the biological significance of these sequences at that time, scientists denominated this sequence in different ways, which resulted in a complexing nomenclature. Till 2002, there eventually came into an agreement that utilizing CRISPR as the unified acronym [14].
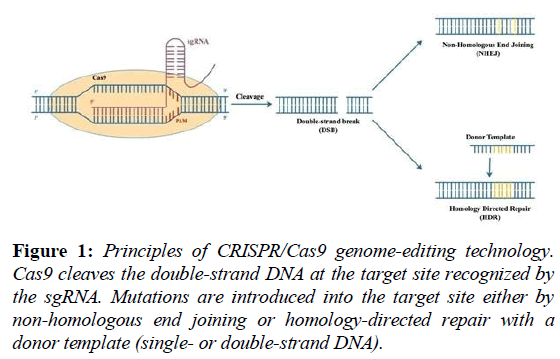
Subsequently, closed to the CRISRP array, researchers identified the cas genes which encoded CRISPR-associated proteins, each protein possesses their own function, for example, the Cas1 protein is used for spacer integration and the Cas3 protein plays a role of helicase [15]. Among all of these CRISPR-associated proteins, the Cas9 protein of type II CRISPR/Cas system from Streptococcus pyogene is the most commonly used nuclease for its function of introducing double-strain breaks (DSBs) at the targeted genomic locus. The cleavage by Cas9 protein is based on the RuvC and HNH endonuclease, and the rejoining of resultant DNA ends can be performed by error-prone non-homologous end joining (NHEJ) or high-fidelity homology directed repair (HDR) (Figure 1) [16].
Figure 1: Principles of CRISPR/Cas9 genome-editing technology. Cas9 cleaves the double-strand DNA at the target site recognized by the sgRNA. Mutations are introduced into the target site either by non-homologous end joining or homology-directed repair with a donor template (single- or double-strand DNA).
In the subsequent studies, the Cas9 protein was guided to cleave specific site of targeted DNA by designed RNA sequence called single guide RNA (sgRNA). Compared with other genome-editing technologies such as ZFNs and TALENs, CRISPR/Cas9 genome-editing technology possesses advantages such as highly-efficient, simplicity, and much lower cost. Although ZFNs and TALENs also enable precise genetic modifications by inducing double-strand DNA breaks (DSBs) at targeted loci in the genome, the molecular mechanisms are different from Cas9. ZFNs or TALENs target site-specific DNA sequences through protein-DNA interactions. The designing and synthesis of proteins with varying DNA binding specificities are difficult for beginners, costly and time- consuming, whereas the RNA guides of Cas9 can be designed easily and synthesized conveniently.
Nowadays, CRISPR/Cas9 system has been widely applied in pathogenic studies as well as therapies of several diseases such as tumor, Duchenne muscular dystrophy (DMD), viral- infection diseases [17]. The CRISPR/Cas9 system has been successfully used in various species such as zebrafish, rats, mice, xenopus, drosophila, worms, dogs, cynomolgus monkeys, pigs, sheep, killifish and salamander [18]. There has never been such a powerful tool for genome-editing in such a wide range of animals.
Applications of CRISPR/Cas9 in cardiac research
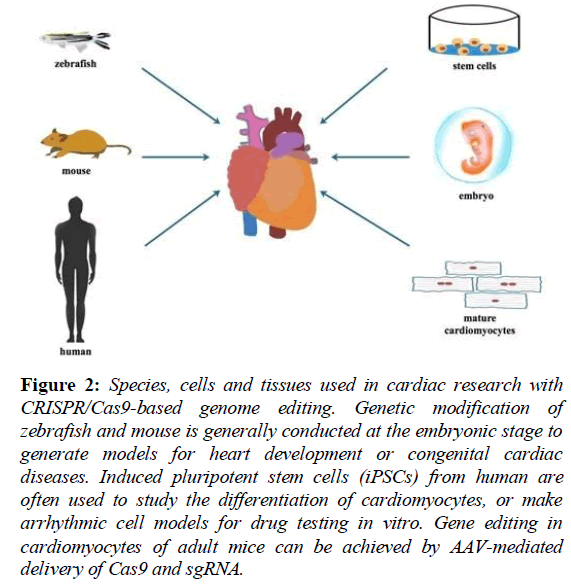
The pathogenesis of cardiac diseases is related to genetic variations. For adult patients who exposed to various factors in the environment, it is hard to assess the accurate role of specific gene mutations in the pathogenesis of cardiac diseases. Nowadays, with the CRISPR/Cas9 genome-editing technology, researchers can easily operate highly-efficient generation of loss-of-function alleles in animals or cells to identify candidate genes for cardiac diseases (Figure 2) [19].
Figure 2: Species, cells and tissues used in cardiac research with CRISPR/Cas9-based genome editing. Genetic modification of zebrafish and mouse is generally conducted at the embryonic stage to generate models for heart development or congenital cardiac diseases. Induced pluripotent stem cells (iPSCs) from human are often used to study the differentiation of cardiomyocytes, or make arrhythmic cell models for drug testing in vitro. Gene editing in cardiomyocytes of adult mice can be achieved by AAV-mediated delivery of Cas9 and sgRNA.
Identification of disease-causing genes
Cardiac diseases involve congenital heart diseases such as Fallot's tetralogy and acquired heart diseases like coronary heart disease. As a low-cost and highly-efficient platform, CRISPR/Cas9 system can be widely used in investigating the genetic regulating mechanisms in cardio-genesis especially seeking for disease-causing genes in congenital heart defects.
Cyanotic congenital heart disease, which involves a series of positional heart defects such as dextrocardia with atrial situs inversus, complete atrioventricular canal defect and pulmonary atresia, is the most common disease in birth defects.Zebrafish is one of the most widely used animal models to study heart development. Its transparent embryo allows direct observation of the larval heart and easy assessment of cardiac parameters. In 2015, researchers conducted mmp21 knock-out in zebrafish via CRISPR/Cas9 gene editing technology. Heart-looping defects were observed in these animals, which validated the hypothesis that mmp21 is crucial for the normal development of cardiac structure [20,21]. In this study, researchers were able to observe the developmental process of the mutant zebrafish heart in real time, and the heart looping defects were detected between 1.5 and 2 hours post fertilization. In the same year, similar practice was operated to target the Dab2 locus in zebrafish embryo to study the function of Dab2 in cardiomyocyte proliferation and differentiation [19]. Furthermore, since the targeting of CRISPR/Cas9 is not to only one single site, researchers successfully performed gene cleavages simultaneously in s1pr2 and spns2, which encode the receptor and transporter of sphingosine-1-phosphate (S1P), respectively. Since both genes are critical for cardiac development, combined cardiac phenotypes were observed in those genetically modified embryos as expected [22-24].
Besides the cyanotic congenital heart disease valvular heart disease is another cardiac disease related to dysplasia of cardiac structure [25]. In America, more than 200,000 people are affected by VHD per year. In view of the etiology of this disease, it has been thought in the past that valvular heart disease is mainly caused by rheumatic fever or ageing of the population. Recently, with CRISPR/Cas9 gene-editing system, researchers discovered the association between valvular heart disease and specific genes. For bicuspid aortic valve (BAV), the most common structural heart disease, medicine therapy can only relieve the symptom and unable to improve the prognosis or outcome of this disease [26]. In 2017, researchers disrupted the expression of GATA4 in induced pluripotent stem cells (iPSCs)-derived myocardial cells by CRISPR/Cas9 gene editing, which obviously obstructed the transition of endothelial cells into normal mesenchymal cells, the crucial step of heart valve development [27].
iPSCs can also be used in the diagnosis of potential arrhythmia. Long QT syndrome (LQTS) is an inherited arrhythmia. Patients who accept the electrocardiogram are characterized by extended QT interphase. Mutations of various genes disrupt the normal activity of ion channels in patients, resulting in the prolongation of action potential in myocardial cells [28]. Traditional examinations such as electrocardiogram cannot diagnose LQTS timely. Nowadays, genetic testing is the most accurate technique for the diagnosis and prediction of LQTS and over 15 subsets of this disease have been genetically defined. However, the diagnosis becomes difficult when facing variants with uncertain significance (VUS). In 2018, researchers utilized CRISPR/Cas9 systems to introduce the mutation T983I into the hERG channel encoding gene KCNH2 in myocardial cells differentiated from iPSCs. The action- potential duration (APD) of myocardial cells which accepted gene-editing was prolonged and the density of hERG K+ current decreased, which explained the mechanism that LQTS onsets. By this way researchers validated that this mutation possessed the potential to cause LQTS [29].
Coronary heart disease is another common cardiac disease, with over 500,000 people affected per year in America, and accounts for more than 60% of total cardiac deaths [30]. The onset of coronary heart disease presents a tendency to familial aggregation, and researchers have identified a large number of loci related to coronary heart disease via genome-wide association studies (GWAS). CRISPR/Cas9 gene editing helps the further identification of candidate genes in these loci. For example, in 2019 researchers utilized CRISPR/Cas9 to generate a deletion of TNF-α sensitive regulatory factor and validated the role AIDA plays in promotion of atherosclerosis [31].
Gene therapy for cardiac diseases
The rapid progression of CRISPR/Cas9 systems provided the potential opportunity for genetic therapy such as correction of the mutant genes or interfering with the expression of disease-causing genes. Targets of gene therapy for cardiac diseases are not restricted to cardiomyocytes, but also embryos, endothelial cells and smooth muscle cells which are involved in the development of cardiac system.
Hypertrophic cardiomyopathy (HCM) is one of the most common causes of sudden cardiac death. It can be caused and inherited by the mutation of MYBPC3 [32]. In 2017, researchers conducted the correction of the mutation in MYBPC3 by CRISPR/Cas9 in human embryos [33]. Embryos were derived from the combination of sperms from males carrying mutated MYBPC3 and ova donated by healthy women. Double-strand breaks (DSBs) were created by CRISPR/Cas9 and the homology directed repair (HDR) was based on the homologous maternal genes. Though mutant genes were predominantly repaired and barely off-target effects were detected, this attempt is currently in the laboratory stage and there are still several uncertainties in editing embryos.
In addition to genetic modification on embryonic level, researchers have explored gene editing in mature cardiomyocytes to change progression of cardiac diseases. Genetic mutations are responsible for a large proportion of arrhythmia. Traditional therapies like anti-arrhythmic drugs or electrophysiological therapy cannot prevent the sudden occurrence of arrhythmia. Now the CRISPR/Cas9 system has been attempted to treat arrhythmia in experimental animals in vivo. The Protein Kinase AMP-Activated Non-Catalytic Subunit Gamma 2 (PRKAG2) cardiac syndrome is an autosomal dominant disease, and H530R in PRKAG2 has been defined the key mutation for the pathogenesis of disease. PRKAG2 cardiac syndrome can progress to ventricular fibrillation, a fatal arrhythmia [34]. In 2016, by injection of CRISPR/Cas9 combined with adeno-associated virus (AAV)- mediated delivery system into mice bearing the H530R PRKAG2 knock-in mutation, researchers successfully corrected the mutation and restored the heart function. This maybe a potential replacement of the traditional therapy, for example, heart transplantation, which is expensive and full of risks in operation [35]. It is worth noting that this genetic rescue was conducted in postnatal mice rather than embryos, which is less technically challenging. As for gene delivery, AAV is the most promising vector in cardiac research, because it possesses the ability of continuing expression in cardiac tissues but causes less inflammatory reactions contrast to adenovirus vector. Nowadays researchers have identified hundreds of serotypes of AAV, among which AAV9 is the most effective carrier to infect cardiomyocytes.
Compared to direct gene editing in cardiomyocytes by CRISPR/Cas9 system, some other cells related to the onset of cardiac diseases such as endothelial cells or smooth muscle cells can also be the candidate targets of genetic therapy. For example, coronary heart disease is one type of metabolic related diseases. Hyperlipidemia is considered the most dangerous risk factor for coronary heart disease [36]. Researchers cannot regenerate necrotic cardiomyocytes; however, the onset of coronary heart disease can be prevented by using gene-editing techniques to reduce lipoprotein levels. Proprotein convertase subtilisin/kexin type 9 (PCSK9) can inhibit the absorption of low-density lipoprotein cholesterol and is a key metabolic regulator in the liver. In 2014, researchers introduced loss-of-function mutation of PCSK9 into the hepatocytes of mice by injection of adenoviral CRISPR/Cas9 vectors, which resulted in reduction of blood cholesterol level for approximately 30% with no off-target effect detected [37]. This approach of disrupting PCSK9 by non-homologous end joining (NHEJ) has been proved in mammals such as mice and monkeys.38 The efficiency of gene editing by CRISPR/Cas9 was as high as 50%, which is promising for further applications. Another group of scientists proposed a novel delivery system based on lipid-like nanoparticles (LLNs) to transduce CRISPR/Cas9 system into hepatocytes [39]. This new tool alleviated the immunogenicity of AAV; however, the level of Pcsk9 protein varies substantially in different gene-edited mice, which indicates that the reliability of this newly vehicle needs to be improved. Lowering cholesterol can also be achieved by suppression of angiopoietin-like 3 (ANGPLT3) gene, which encodes the lipoprotein lipase inhibitor to raise blood cholesterol levels. Chadwick and his colleagues introduced loss-of-function mutation of Angptl3 by injection of adenoviral vectored- CRISPR/Cas9 system into 5-week-old mice, which resulted in a much lower cholesterol level compared to that of Pcsk9- targeted mice [40].
The future of CRISPR/Cas9 genome editing in cardiac research
The development of CRISPR/Cas9 genome-editing technology has accelerated the cardiac research [41]. Cardiac tissue engineering is a promising field for the use of CRISPR/Cas9 in transplantation. Nowadays, xenotransplantation based on animal organs (particularly those of pigs due to the similar size as human’ s) has been considered the solution of releasing shortages of human organs (most of the donated organs at present were obtained from the dying patients). However, the risk of porcine endogenous retrovirus (PERV) in pigs hindered the further clinical application of porcine organs for its potential risk of immune rejection after transplantation [42]. In 2017, Luhan Yang with her colleagues achieved whole-genome PERV-inactivation in pigs by utilization of CRISPR/Cas9 system, which prevented the viral transmission and reduced the safety concern of porcine organs for xenotransplantation [43]. We can expect that the risks of xenotransplantation with porcine organs will be greatly reduced with the CRISPR/Cas9 gene editing technology and more preclinical trials will be conducted in monkeys in the near future.
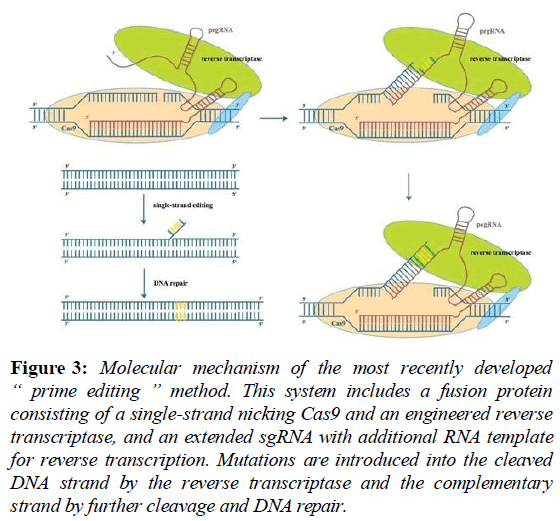
Currently, controversies over genome-editing technology mainly exist in the following aspects.Off-target effect is one of the major technical problems, which means that the designed sgRNA mismatched with the non-targeted DNA sequences and then introduce unanticipated gene mutations [44,45]. This phenomenon can induce instabilities of genomes and disrupting normal gene functions [46]. Researchers have tried several methods to deal with this problem, for example, Banasik and his colleagues achieved accurate detection of off- target cleavage by utilizing integrase-defective lentiviral vectors (LDLVs), which is much more effective than in silico prediction or in vitro selections [47,48]. Huang and Skarnes’s groups improved the CRISPR/Cas9 system by using double- nicking strategy, in which paired sgRNAs were designed and the PAM was oriented in a tail-to-tail configuration [49]. The team led by Feng Zhang discovered new Cas9 protein in Staphylococcus aureus instead of Pyogenic coccus, which is able to enhance the specificity to target locus [50]. Most recently, a new method named “prime editing” was developed by David Liu’s group. This method combined an engineered Cas9 and a reverse transcriptase as a fusion protein, and used a prime editing guide RNA containing the sgRNA and an RNA template for reverse transcription on target site [51]. Cas9 and sgRNA enable the precise cleavage of a single strand, and the reverse transcriptase introduces mutations into this strand according to the sequence of the RNA template (Figure 3).
Figure 3: Molecular mechanism of the most recently developed “ prime editing ” method. This system includes a fusion protein consisting of a single-strand nicking Cas9 and an engineered reverse transcriptase, and an extended sgRNA with additional RNA template for reverse transcription. Mutations are introduced into the cleaved DNA strand by the reverse transcriptase and the complementary strand by further cleavage and DNA repair.
The sequence of the complementary strand is then modified by further DNA cleavage and repair. This method can be used to correct theoretically ~89% of disease-causing mutations in humans with higher efficiency and much lower off-target effect compared to the conventional homology-directed repair. Further improvement of this method or development of novel and safer genome-editing tools may bring Cas9-based technologies closer to clinical use in the future.
The ethic of gene editing technology is another focus of attention, especially after the controversial genetic modification in human embryos and the birth of CCR5-edited babies in 2018. Gene editing in germ cells and somatic cells is quite different. Genes and mutations in germ cells could be passed on to the next generation, which is undoubtedly full of risks. Hence there still exists a long way for the clinical application of gene-editing technology in human beings. Laws and regulations are urgently required to strictly control the use of genome-editing technologies in any unethical ways.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31300965 to G.-L.C., and 81974093 to B.Z.)
Conflict of interests
The authors declare that no competing interests exist.
References
- Thomas H, Diamond J, Vieco A, et al. Global Atlas of Cardiovascular Disease 2000-2016: The Path to Prevention and Control. Glob Heart. 2018; 13(3):143-163.
- Heidenreich PA, Trogdon JG, Khavjou OA, et al. Forecasting the future of cardiovascular disease in the United States: a policy statement from the American Heart Association. Circulation. 2011; 123(8):933-944.
- Corella D, Asensio EM, Coltell O, et al. CLOCK gene variation is associated with incidence of type-2 diabetes and cardiovascular diseases in type-2 diabetic subjects: dietary modulation in the PREDIMED randomized trial. Cardiovasc Diabetol. 2016; 15:4.
- Mormile R. Multiple sclerosis and susceptibility to cardiovascular diseases: Implications of ethnicity-related interleukin-17A gene polymorphism? Med Hypotheses. 2015; 85(3):365-366.
- Pasipoularides A. Linking Genes to Cardiovascular Diseases: Gene Action and Gene-Environment Interactions. J Cardiovasc Transl Res. 2015; 8(9):506-527.
- Kim YG, Cha J, Chandrasegaran S. Hybrid restriction enzymes: zinc finger fusions to Fok I cleavage domain. Proc Natl Acad Sci U S A. 1996; 93(3):1156-1160.
- Jiang W, Bikard D, Cox D, et al. RNA-guided editing of bacterial genomes using CRISPR-Cas systems. Nat Biotechnol. 2013; 31(3):233-239.
- Gao C. The future of CRISPR technologies in agriculture. Nat Rev Mol Cell Biol. 2018; 19(5):275-276.
- Fellmann C, Gowen BG, Lin PC, et al. Cornerstones of CRISPR-Cas in drug discovery and therapy. Nat Rev Drug Discov. 2017;16(2):89-100.
- Sorek R, Kunin V, Hugenholtz P. CRISPR--a widespread system that provides acquired resistance against phages in bacteria and archaea. Nat Rev Microbiol. 2008; 6(3):181-186.
- Maxwell KL. The Anti-CRISPR Story: A Battle for Survival. Mol Cell. 2017; 68(1):8-14.
- Jinek M, Chylinski K, Fonfara I, et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 2012; 337(6096):816-21.
- Ishino Y, Shinagawa H, Makino K, et al. Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. J Bacteriol. 1987; 169 (12):5429-5433.
- Godde JS, Bickerton A. The repetitive DNA elements called CRISPRs and their associated genes: evidence of horizontal transfer among prokaryotes. J Mol Evol 2006; 62 (6):718-729.
- Nunez JK, Kranzusch PJ, Noeske J, et al. Cas1-Cas2 complex formation mediates spacer acquisition during CRISPR-Cas adaptive immunity. Nature Structural & Molecular Biology. 2014; 21:528-534.
- Sternberg SH, Redding Sy, Jinek M, Greene EC, Doudna JA. DNA interrogation by the CRISPR RNA-guided endonuclease Cas9. Nature. 2014; 507:62-67.
- Kessler M, Rottbauer W, Just S. Recent progress in the use of zebrafish for novel cardiac drug discovery. Expert Opin Drug Discov. 2015; 10(11):1231-1241.
- Zou Q, Wang X, Liu Y, et al. Generation of gene-target dogs using CRISPR/Cas9 system. J Mol Cell Biol 2015; 7(6):580-583.
- Hofsteen P, Robitaille AM, Chapman DP, et al. Quantitative proteomics identify DAB2 as a cardiac developmental regulator that inhibits WNT/beta-catenin signaling. Proc Natl Acad Sci U S A. 2016; 113(4):1002-1007.
- Perles Z, Moon S, Ta-Shma A, et al. A human laterality disorder caused by a homozygous deleterious mutation in MMP21. J Med Genet 2015; 52(12):840-847.
- Sutherland MJ, Ware SM. Disorders of left-right asymmetry: heterotaxy and situs inversus. Am J Med Genet C Semin Med Genet. 2009; 151C(4):307-317.
- Ota S, Hisano Y, Ikawa Y, et al. Multiple genome modifications by the CRISPR/Cas9 system in zebrafish. Genes Cells. 2014; 19(7):555-564.
- Kupperman E, An S, Osborne N, et al. A sphingosine-1-phosphate receptor regulates cell migration during vertebrate heart development. Nature. 2000; 406(6792):192-195.
- Kawahara A, Nishi T, Hisano Y, et al. The sphingolipid transporter spns2 functions in migration of zebrafish myocardial precursors. Science. 2009; 323(5913):524-527.
- Iung B, Vahanian A. Epidemiology of acquired valvular heart disease. Can J Cardiol. 2014; 30 (9):962-970.
- LaHaye S, Lincoln J, Garg V. Genetics of valvular heart disease. Curr Cardiol Rep. 2014; 16(6):487.
- Yang B, Zhou W, Jiao J, et al. Protein-altering and regulatory genetic variants near GATA4 implicated in bicuspid aortic valve. Nat Commun. 2017; 8:15481.
- Niekerk CV, Deventer BSV, du Toit-Prinsloo L. Long QT syndrome and sudden unexpected infant death. J Clin Pathol. 2017; 70(9):808-813.
- Garg P, Oikonomopoulos A, Chen H, et al. Genome Editing of Induced Pluripotent Stem Cells to Decipher Cardiac Channelopathy Variant. J Am Coll Cardiol. 2018; 72(1):62-75.
- Ford ES, Roger VL, Dunlay SM, et al. Challenges of ascertaining national trends in the incidence of coronary heart disease in the United States. J Am Heart Assoc. 2014; 3 6):e001097.
- Lalonde S, Codina-Fauteux VA, de Bellefon SM, et al. Integrative analysis of vascular endothelial cell genomic features identifies AIDA as a coronary artery disease candidate gene. Genome Biol. 2019; 20(1):133.
- Prondzynski M, Mearini G, Carrier L. Gene therapy strategies in the treatment of hypertrophic cardiomyopathy. Pflugers Archiv. 2019; 471(5):807-815.
- Ma H, Marti-Gutierrez N, Park SW, et al. Correction of a pathogenic gene mutation in human embryos. Nature. 2017; 548(7668):413-419.
- Banankhah P, Fishbein GA, Dota A, et al. Cardiac manifestations of PRKAG2 mutation. BMC medical genetics. 2018; 19:1.
- Xie C, Zhang YP, Song L, et al. Genome editing with CRISPR/Cas9 in postnatal mice corrects PRKAG2 cardiac syndrome. Cell Res. 2016;26(10):1099-1111.
- Nordestgaard BG, Chapman MJ, Ray K, et al. Lipoprotein(a) as a cardiovascular risk factor: current status. Eur Heart J. 2010; 31(23):2844-2853.
- Ding Q, Strong A, Patel KM, et al. Permanent alteration of PCSK9 with in vivo CRISPR-Cas9 genome editing. Circ Res. 2014; 115(5):488-492.
- Chadwick AC, Wang X, Musunuru K. In Vivo Base Editing of PCSK9 (Proprotein Convertase Subtilisin/Kexin Type 9) as a Therapeutic Alternative to Genome Editing. Arterioscler Thromb Vasc Biol. 2017; 37(9):1741-1747.
- Jiang C, Mei M, Li B, et al. A non-viral CRISPR/Cas9 delivery system for therapeutically targeting HBV DNA and pcsk9 in vivo. Cell Res. 2017;27(3):440-443.
- Chadwick AC, Evitt NH, Lv W, Musunuru K. Reduced Blood Lipid Levels With In Vivo CRISPR-Cas9 Base Editing of ANGPTL3. Circulation. 2018; 137(9):975-977.
- Rezaei H, Khadempar S, Farahani N, et al. Harnessing CRISPR/Cas9 technology in cardiovascular disease. Trends Cardiovasc Med. 2020;30(2):93-101.
- Reardon S. New life for pig-to-human transplants. Nature. 2015; 527(7577):152-154.
- Niu D, Wei HJ, Lin L, et al. Inactivation of porcine endogenous retrovirus in pigs using CRISPR-Cas9. Science. 2017; 357(6357):1303-1307.
- Doench JG, Fusi N, Sullender M, et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat Biotechnol. 2016; 34(2):184-191.
- Zhang XH, Tee LY, Wang XG, et al. Off-target Effects in CRISPR/Cas9-mediated Genome Engineering. Mol Ther Nucleic Acids. 2015; 4(11):e264.
- Xiao A, Wang Z, Hu Y, et al. Chromosomal deletions and inversions mediated by TALENs and CRISPR/Cas in zebrafish. Nucleic Acids Res. 2013; 41(14):e141.
- Wang X, Wang Y, Wu X, et al. Unbiased detection of off-target cleavage by CRISPR-Cas9 and TALENs using integrase-defective lentiviral vectors. Nat Biotechnol. 2015; 33(2):175-178.
- Fu Y, Foden JA, Khayter C, et al. High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat Biotechnol. 2013; 31(9):822-826.
- Shen B, Zhang W, Zhang J, et al. Efficient genome modification by CRISPR-Cas9 nickase with minimal off-target effects. Nature methods. 2014; 11(4):399-402.
- Ran FA, Cong L, Yan WX, et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature. 2015; 520(7546):186-191.
- Anzalone AV, Randolph PB, Davis JR, et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature. 2019:576(7785):149-157.